Team:Frankfurt/Notebook
From 2012.igem.org
| Home | Team | Project | Organisms | New Yeast RFC | Notebook | Registered Parts | Modeling | Safety | Attributions | Official Team Profile |
|---|
Contents |
Labwork
May and June 2012
- Arrangements for labwork
- preparation of competent cells (E.coli, S.cerevisiae), agarose plates (LB, YEPD, SCD-ura,…), medium for E.coli and S.cerevisiae
- Purchasing of the equipment (reaction tubes, glass bottles, pipette tips,..)
- Primer design
July 2012
- Plasmid isolation of p426, p423, pUD8e, pUD22e from E.coli
- Isolation of chromosomal DNA of CEN.PK2-1C
- Trials to get the genes, promoters and terminators via PCR
August 2012
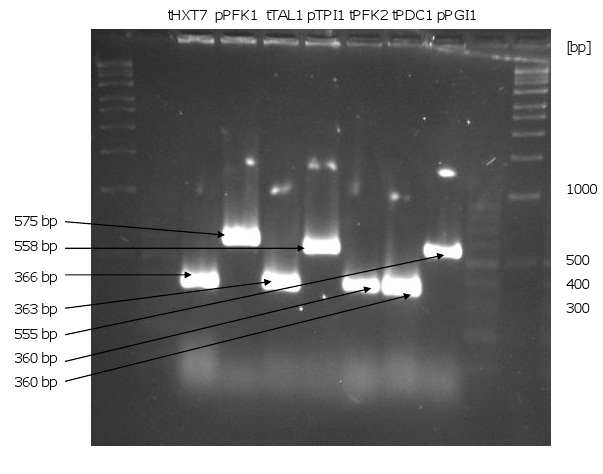
- PCR of the genes, promoters and terminators
- all genes (without KO and KAH), promoters and terminators could be amplified
- Linearization of p426 and p423 with SpeI and XhoI
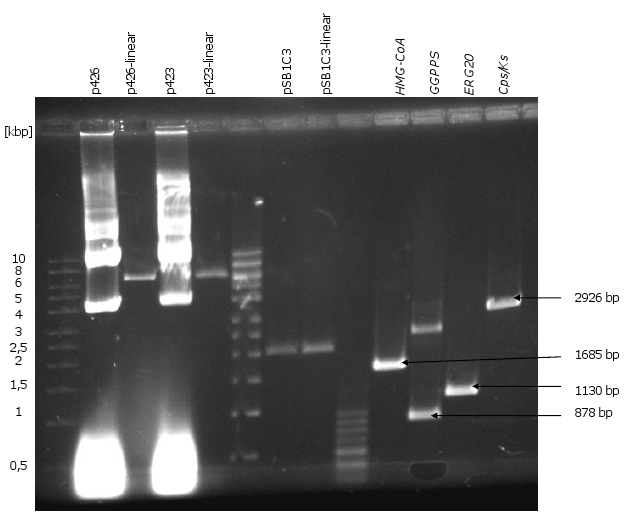
| Templates | Amplified DNA Fragments |
|---|---|
| synthesized sequence of HMG-CoA | HMG-CoA |
| synthesized sequence of GGPPS | GGPPS |
| synthesized sequence of Cps/Ks | CPS/KS |
| chromosomal DNA of CEN.PK2-1C | ERG20 |
- Biobrick production of the genes HMG-CoA, ERG20, CPS/KS
- restriction of 3 µg of the genes with EcoRI and PstI
- ligation of biobrick genes with linear pSB1C3
- transformation of the ligation in E.coli
- plasmid isolation of E.coli clones
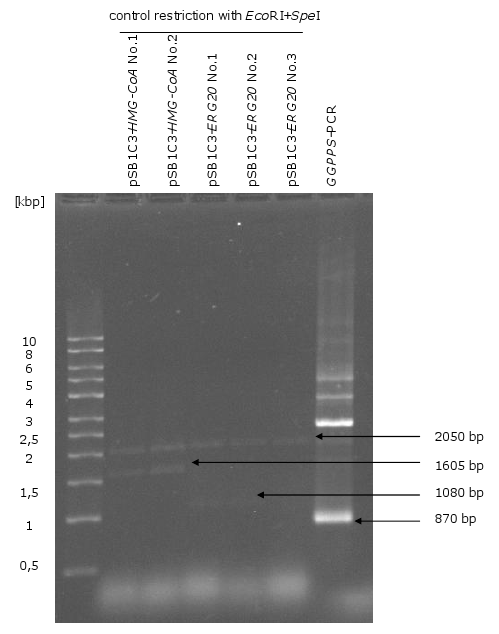
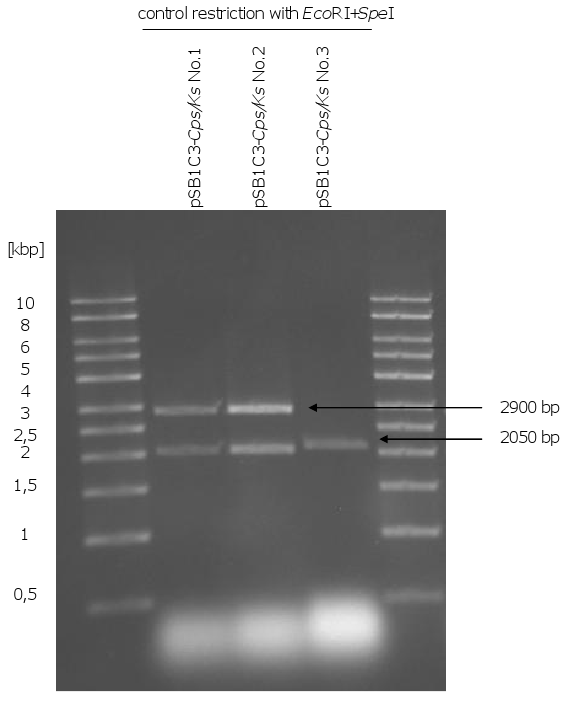
- control restriction of biobrick plasmids with EcoRI and SpeI
- restriction of 3 µg of the genes with EcoRI and PstI
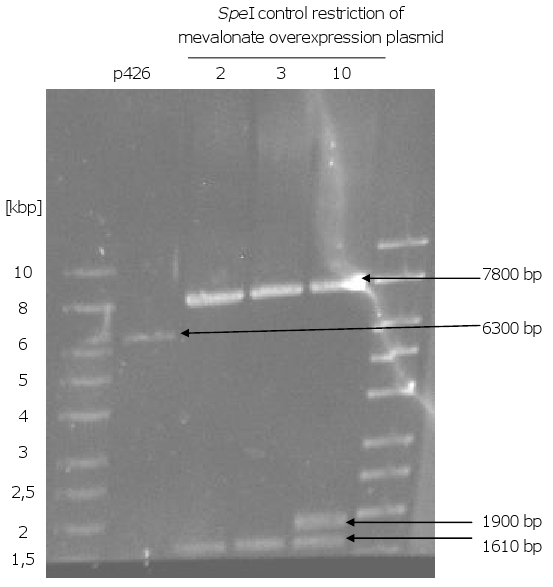
- Formation of the mevalonate overexpression plasmid via gap repair
- first and second yeast transformation with equimolar quantities of DNA fragments for mevalonate overexpression (p426 with 7 inserts): only very small colonies could grow after the first and second transformation
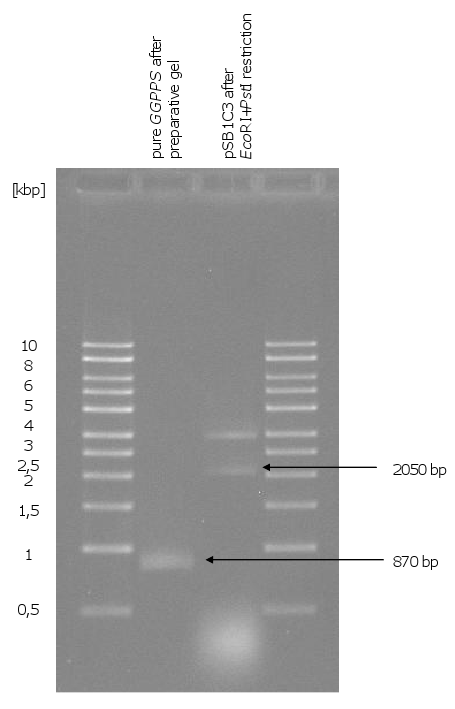
- using pure GGPPS (purification of a preparative gel) for the third yeast transformation: normal size of the colonies
- inoculation of several clones of the third yeast transformation
- plasmid preparation of the clones
- transformation of the plasmids in E.coli
- Amplifying pSB1C3 for biobrick production
- trials to amplify pSB1C3, whose blunt ends were ligated and transformed in E.coli
- pSB1C3 should be linearized by EcoRI and PstI : did not work (two fragments instead of one)
- preparative gel of the linear fragment: very low concentration of linear pSB1C3 (was not sufficient for ligation)
- GC analysis
- GC analysis of the wild type CEN.PK2-1C (standard GGOH): as expected no GGOH could be observed
- Assembly of the KO and the KAH fragments
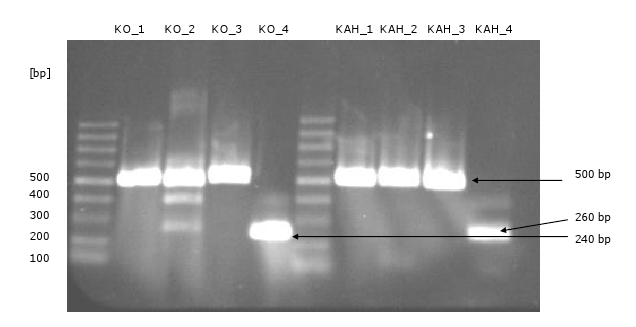
- amplification of the fragments via PCR (there are four fragments of each gene with an overhang to the fragment beside of 30 bp)
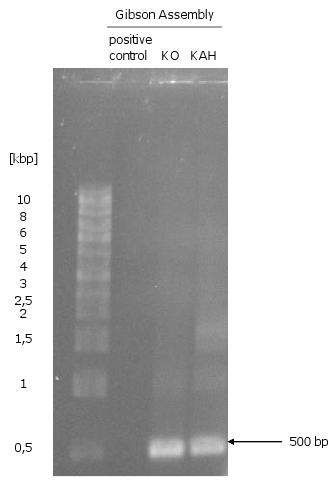
- Gibson assembly of the fragments of KO and KAH: did not work
September 2012
- Formation of the mevalonate overexpression plasmid via gap repair
- plasmid isolation (p426 with 7 inserts) from E.coli
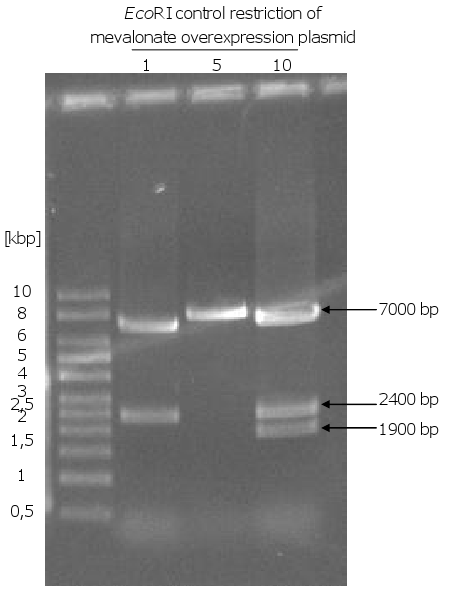
- control restriction of the plasmids with EcoRI and SpeI: one clone out of 10 got the right sizes
- Ergosterol experiment
- idea: maybe the clones of the first and the second yeast transformation grow better after ergosterol addition (0,02 g/l)): wildtype with and without ergosterol and one of the clones with and without ergosterol (no significant difference in growth could be observed)
- GGPPS-PCR
- PCR of GGPPS with shorter synthesis time in order to get only the correct fragment
- Amplification of pSB1C3 for biobrick production
- transformation of pSB1C3-RFP in E.coli
- isolation of the plasmid from E.coli
- linearization with EcoRI and PstI: it worked (only the correct fragment was observed)
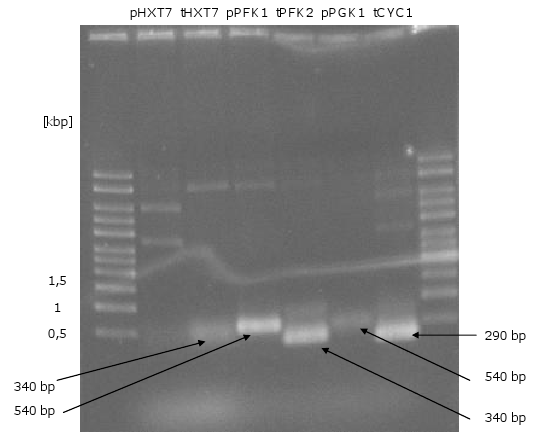
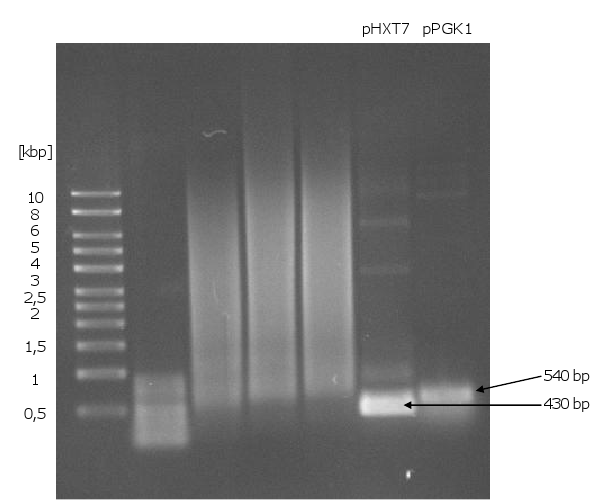
- PCR of biobrick-promoters and -terminators
- PCR of biobrick promoters and terminators that were used to build the mevalonate overexpression plasmid
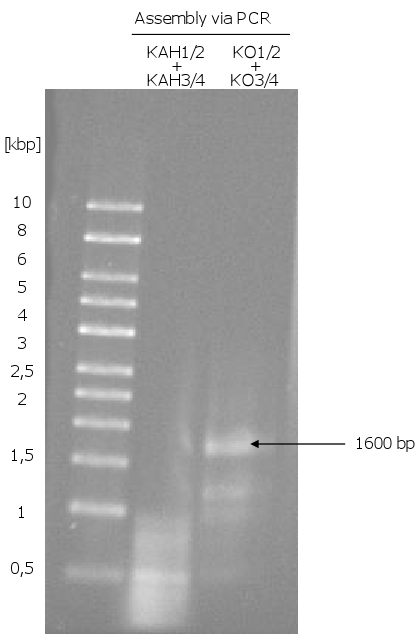
- Assembly of the KO and the KAH fragments
- trials to assembly the KO and the KAH fragments via PCR (the overhangs were used as primer): KO worked, KAH not
- amplification of KO
- Biobrick production of the genes GGPPS, KO and the promoters and terminators
- restriction of 3 µg of the DNA fragments with EcoRI and PstI
- ligation of the promoters, terminators, KO and the GGPPS with pSB1C3
- transformation of the ligation in E.coli
- plasmid isolation of E.coli clones
- control restriction of biobrick plasmids with EcoRI and SpeI
- Midi-preparation of plasmids for sequencing
- biobrick plasmids
- mevalonate overexpression plasmid
- GC
- GC analysis of the wild type CEN.PK2-1C with p426 and the wild type with the mevalonate overexpression plasmid in SCD-ura
- standard GGOH
- Formation of the plasmid for steviol synthesis
- yeast transformation with equimolar quantities of DNA fragments for steviol synthesis (p423 with 7 inserts)
- problem: could not assemble KAH, assembled KAH1/2 and KAH3/4 were used (KAH1/2 and KAH3/4 only got a 30 bp overhang instead of a 45 bp overhang): gap repair did not work
Methods and Protocols
Plasmid Preparation
Plasmid Preparation of E.coli (Mini Preparation)
- transfer 1,5 ml of an overnight culture in a reaction tube
- centrifuge at 8000 rpm at RT for 5 min
- resuspend the pellet in 100 µl solution 1
- addition of 200 µl solution 2
- mix till the solution is clear
- incubate at RT for 5 min
- addition of 150 µl cold solution 3, mix till protein clumps are build
- incubate 10 min on ice, centrifuge 15 min with 10000 rpm
- transfer the supernatant in a clean reaction tube
- fill the reaction tube with 96 % EtOH, mix and let it precipitate at -20 °C for 10 min
- centrifuge 10 min at 10000 rpm
- wash the pellet with 70 % EtOH
- dry the pellet and resuspend it with 30 µl water or TE buffer
Solution 1
- 50 mM glucose
- 10 mM EDTA
- 25 mM Tris-HCl pH8
Solution 2
- 0,2 M NaOH
- 1 % SDS
Solution 3
- 3 M KaAc pH 5,5
Plasmid Preparation of Saccharomyces cerevisia
- overnight culture in 5 ml
- centrifuge 2 ml of the cells for 1-2 min
- wash the cells with water
- resuspend the pellet in 400 µl buffer 1 with RNase
- addition of 400 µl buffer 2 and mix carefully
- addition of 2/3 volume of glass beads
- cell destruction: vibrax the cellsin a 2 ml reaction tube for 5 min at 4°C
- transfer 500 µl of the supernatant in a clean reaction tube
- addition of 250 µl buffer 3, mix, incubate it for 10 min on ice
- centrifuge at 10000 rpm, 15 min
- transfer the supernatant in a clean reaction tube, fill with isopropanol and mix
- centrifuge at 13000 rpm, 30 min
- wash the pellet with cold 70 % EtOH and let it dry
- resuspend the DNA in 30 µl water or TE buffer
P1
- 50 mM Tris/HCl pH 8
- 10 mM EDTA
- 100 µg/ml RNase A
P2
- 0,2 M NaOH
- 1 % SDS
P3
- 3 M KAc
Transformation
Yeast Transformation
- Inoculate the synthetic complete medium (SC) with the strain
- Incubate with shaking overnight at 30°C
- Harvest the cells at a OD600 0,5-0,6 by centrifugation (3000x g, 5 min, RT)
- Wash the cells with 0,5 vol of sterile water (resuspend by shaking, centrifugate with 3000x g, 5 min, RT)
- Resuspend the cells in 0,01 vol of sterile water, transfer the suspension to a reaction tube and pellet the cells (3000x g, 5 min, RT)
- Resuspend the pellet in 0,01 vol of sterile filtrated FCC (frozen competent cell) solution
- Aliquot 50 µl of the solution into the reaction tubes
- Store the cells at -80°C for at least one night (up to one year)
- Mastermix for the FCC transformation mixture:
| Substance | Volume [µl] |
| PEG 3350 (50% (w/v)) | 260 |
| LiAcetat 1.0 M | 36 |
| Single-stranded carrier DNA (10 mg/ml) | 10 |
| Total volume | 306 |
- Prepare DNA aliquots: Solute enough DNA (e.g. 100 ng plasmid) in 54 µl of water
- Unfreeze the cells in a 37°C block for 15-30 sec
- Centrifuge the solution at 13000x g for 2 minutes
- Remove the supernatant
- Add 306 µl of FCC transformation mixture to the cells
- Add 54 µl of the DNA to the solution and vortex shortly
- When all the reaction tubes are prepared, vortex the samples well until all pellets are completely resoluted
- Incubate the samples for 40 minutes at 42°C in a heating block
- Centrifuge the cells at 13000x g for 30 sec and pour off the supernatant
- Resuspend the cells in sterile water by vortexing
- Spread onto the appropriate medium
- Let the cells grow at 30°C
Yeast transformation and gap repair (example)
| DNA Fragment | Size[bp] | Concentration[ng/µl] | Equimolar Quantities°[ng] | Used Volume[µl] |
|---|---|---|---|---|
| HMG-CoA | 1600 | 224 | 222 | 1 |
| tHXT7 | 360 | 51,6 | 50 | 1 |
| pPFK1 | 600 | 54 | 83 | 1,5 |
| ERG20 | 1100 | 139 | 152 | 1 |
| tPFK2 | 360 | 47,4 | 50 | 1 |
| pPGK1 | 600 | 49 | 83 | 1,7 |
| GGPPS | 900 | 14,5 | 125 | 8,6 |
| 5520 | 765 | + 19,8 µl linear p426 + 18,4 µl water |
° determination of 50 ng for 360 bp
use of a vector-insert relation of 1:1
samples: linear p426 with 7 inserts
- positive control (p426)
- negative control (linear p426)
plating the samples on SCD-ura and incubation at 30° C
General procedure of a transformation via gap repair
- Amplification of the required DNA fragment with homologue ends of the plasmid (in our case homologue ends of the gene/promoter/terminator beside the DNA fragment) via PCR and an agarose gel to review the correct size
- Linearization of the plasmid and an agarose gel for review
- Yeast transformation with the PCR products and the linear plasmid (homologue recombination)
- Selection on an agar plate without the metabolite, which is on the plasmid for yeast selection (in our case it was uracil on p426 and histidin on p423)
- Inoculation of clones and isolation of the plasmids from yeast
- Transformation of a plasmid in E.coli and selection on LB medium with ampicillin (the plasmids have an ampicillin resistance)
- Isolation of the plasmids from E.coli
- Diagnostic restriction of the plasmid in order to find the correct plasmid with all inserts
- Transformation of the correct plasmid in yeast to do further experiments
E.coli Transformation
a) Production of competent cells
1. Inoculate 400 ml LB medium (pre-warmed to 37°C) in a 1 l flask with 100 µl of a fresh over night culture
2. Harvest the cells at a OD600 0,60-0,65 (in a room with 37°C, sterile)
3. Aliquot the solution in 8 x 50 ml falcons (sterile, pre-cooled on ice)
4. Incubate for 30 min on ice
5. Centrifuge for 12,5 min with 4000x g, 4°C
6. Resuspend the pellets in each 10 ml sterile water (4°C)
7. Centrifuge for 10 min with 4000x g, 4°C
8. Repeat steps 6 and 7 two times
9. Resuspend the cells in 10 ml 10% glycerine (+millipor, sterile, 4°C)
10. Centrifuge for 15 min with 4000x g, 4°C
11. Add 800 µl of 10% glycerine (sterile, 4°C) to the cells
12. Aliquot 40 µl of the solution in each reaction tube (pre-cooled) and freeze them in liquid nitrogen
13. Store them up to 6 months at -70°C
b) Transformation (on ice)
1. Add DNA (e.g. 1 µl of extracted yeast plasmids) to the frozen cells
2. Unfreeze the cells on ice
3. Fill the cells into cuvettes for electroporation
4. Electroporation (for 2 mm cuvettes: voltage I = 2,5 kV; resistance R = 2000 ohm; electrical current A = 25 µF; for 1 mm cuvettes: I = 2,2 kV; resistance R = 2000 ohm; electrical current A = 25 µF)
5. Fill the cells with 1 ml of SOC medium back in a reaction tube
6. Shake them for 1 hour at 37°C
7. Centrifuge for 1 min 8000x g
8. Decant the supernatant
9. Spread onto the appropriate medium
PCR
Culture Media
| Full Medium (YEPD) for Yeast | |||
|---|---|---|---|
| Yeast Extract | 1 % (weight/volume) | ||
| Pepton | 2 % (w/v) | ||
| Glucose | 2 % (w/v) | ||
| Synthetic Complete Medium (SC) for Yeast | |||
|---|---|---|---|
| Yeast Nitrogen Base | 0.17 % (w/v) | ||
| Ammoniumsulfate | 0.5 % (w/v) | ||
| Glucose | 2 % (w/v) | ||
| Amino Acid Mix° | 50 ml/l | ||
| Histidin** | 0.25 mM | ||
| Tryptophan** | 0.19 mM | ||
| Leucin** | 0.35 mM | ||
| Uracil** | 0.44 mM | ||
pH has to be regulated with KOH to pH=6.3
° contains no His, Leu, Trp and Uracil
** addition of this components depents on the respective selection medium
| SOC-Medium for Regeneration of transformed Escherichia coli`s after Electroporation | |||
|---|---|---|---|
| Trypton | 2 % (w/v) | ||
| Yeast Extract | 0.5 % (w/v) | ||
| NaCl | 10 mM | ||
| KCl | 2,5 mM | ||
| MgCl2 | 10 mM | ||
| MgSO4 | 10 mM | ||
| Glucose | 20 mM | ||
pH has to be regulated to pH=6.8-7.0
| Full Medium (LB) for E.coli | |||
|---|---|---|---|
| Yeast Extract | 0.5 % (w/v) | ||
| Trypton | 1 % (w/v) | ||
| NaCl | 0.5 % (w/v) | ||
pH has to be regulated with NaOH to pH=7.5
Every cluture medium has to be autoclaved to be sterile.
Agar Plate
LBampicilline-Agar
Add 2 % agar to LB-medium. After autoclaving and cooling-down to 60 °C steril ampicillin is added. Plates were poured.
LBchloramphenicol-Agar
Add 2 % agar to LB-medium. After autoclaving and cooling-down to 60 °C ethanolic chlorampenicol solvation is added.
SCD-Agar
Add 2 % agar to SCD-medium. After autoclaving and cooling-down steril amino acid solution is added. Dependent on the respective selective medium Histidin (0.25 mM), Trypthophan (0.19 mM), Uracil (0.44 mM) or Leucin (0.35 mM) are added. Plates were poured.
YEPDG418-Agar
Add 2 % agar to YEPD-medium. After autoclaving and cooling-down sterile G418 (final concentration 2g/l) is added. Plates were poured.
Gel Electrophoresis
| Agarose Gel (1x) | |||
|---|---|---|---|
| TAE puffer | 1x | ||
| Agarose | 1 % (w/v) | ||
Solve agarose in TAE by boiling it. After cooling-down to 55-60 °C gel is poured.
| TAE Puffer (50x) for Gel Electrophoresis | |||
|---|---|---|---|
| EDTA | 18,6 g | ||
| Tris | 242g | ||
| Glacial Acetic Acid | 57,2 ml | ||
| Purified Water | 1000ml | ||
pH has to be regulated with glacial acetic acid to pH=8.
Gels were run with a tension of 80-140 V.
Kits
PCR Purification Kit from Qiagen
Gel Extraction Kit from Qiagen
Midi Plasmid Preparation Kit from Qiagen
 "
"