Team:Bielefeld-Germany/Results/vector
From 2012.igem.org

Summary
Contents |
Description of the shuttle vector system
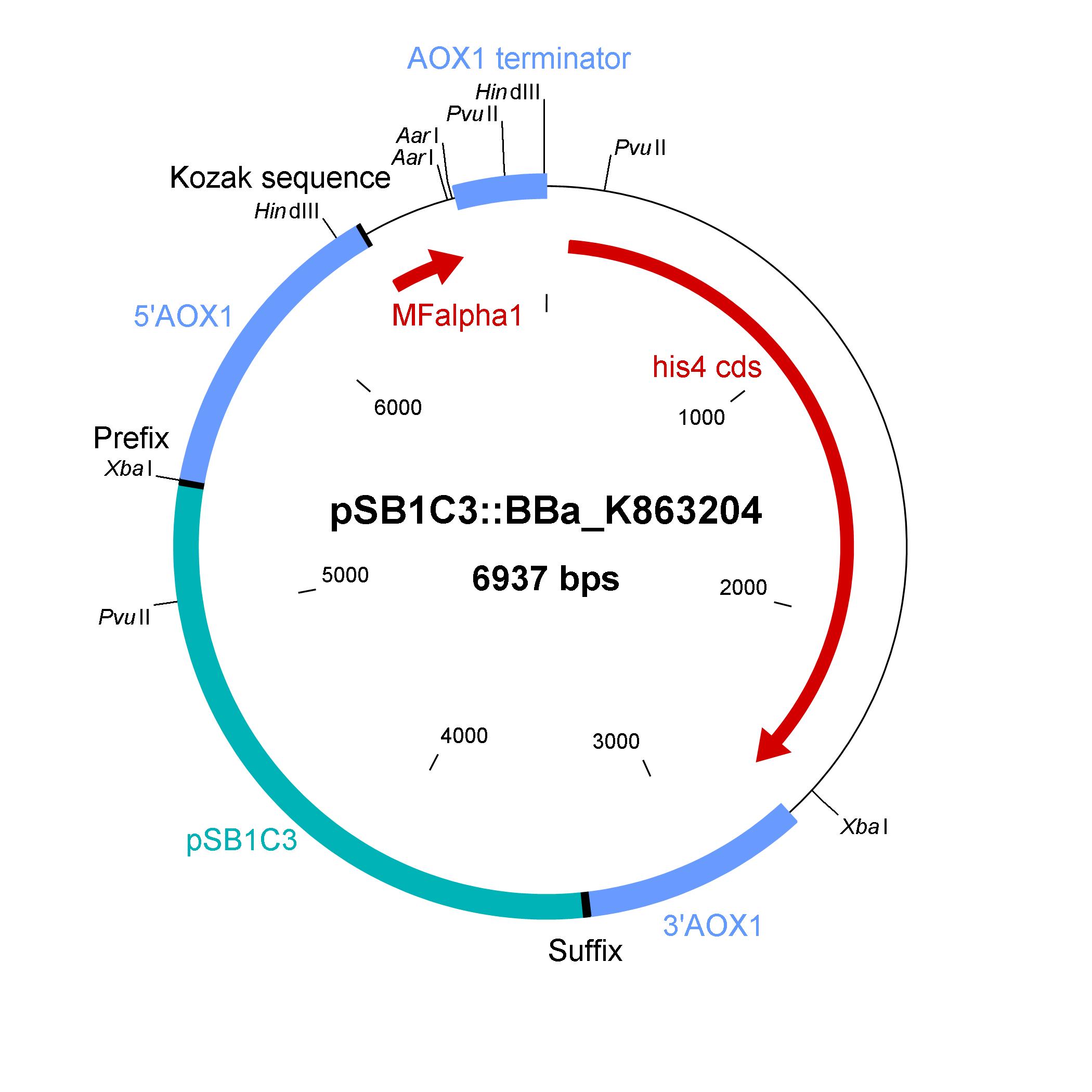
The design of a minimal shuttle vector system with defined regions (or better DNA fragments) for expression and secretion of proteins of interest (POI) like laccase which needed glycolisation is the basic concept. The shuttle vector needs a bacterial part for cloning in bacteria (like E. coli) and an eucaryotic part for genomic integration and selection in yeast (like P. pastoris). For other teams it is able to clone in frame their POI via the AarI restriction site. With only one restriction-ligation-cloning step the shuttle vector will be ready to use and integrate in the eucaryote P. pastoris.
The shuttle vector consists of the plasmid pSB1C3, 5' UTR of alcohol oxidase 1 gene (aox1) containing the aox1 promoter region, Kozak sequence, mating factor alpha 1 (MFalpha1), AarI restriction site, aox1 terminater, his4 gene and 3' UTR of aox1 gene. Cloning and plasmid replication in E. coli is able via the pSB1C3 part. The protein of interest (like laccase) can be included in frame with MFalpha1 via AarI restriction site. With the N-terminal MFalpha1 the POI could be secreted in the media. Genomic intregation of MFalpha1-taged POI is able via the 5' UTR and 3' UTR of the aox1 gene. This allows a double cross over and the genomic integration without any bacterial proportion of DNA which could be a decisive point for industrial application. The complementation of histidine auxotrophie via his4 gene is choosen instead of a zeocine resistance. This selection strategy is choosen because we want to avoid the application of antibiotics as many as possible.
Gibson assembly is used for building the shuttle vector (see the figure below) and the fragments with 5' overlap were amplified via PCR. In addition, die fragments were designed as basic BioBrick parts for use. The origin of the DNA sequence for design of the shuttle vector and the source of DNA for PCR is listed in the table below.
Elements of the shuttle vector and their origin
| Element | Origin of DNA sequence of design | Origin of DNA sequence of PCR |
|---|---|---|
| [http://partsregistry.org/Part:pSB1C3 pSB1C3] | [http://partsregistry.org/Part:pSB1C3 pSB1C3] | [http://partsregistry.org/Part:pSB1C3 pSB1C3] |
| [http://partsregistry.org/Part:BBa_K863200 5'UTR of aox1] | [http://products.invitrogen.com/ivgn/product/V19520 plasmid pPICZalphaA (Invitrogen)] | P. pastoris wild type X-33 |
| Kozak sequence | [http://partsregistry.org/wiki/index.php?title=Part:BBa_J63003 BBa_J63003] | integrated in primer sequence |
| [http://partsregistry.org/Part:BBa_K863203 MFalpha1] | [http://products.invitrogen.com/ivgn/product/V19520 plasmid pPICZalphaA (Invitrogen)] | [http://products.invitrogen.com/ivgn/product/V19520 plasmid pPICZalphaA (Invitrogen)] |
| [http://partsregistry.org/Part:BBa_K863203 aox1 terminater] | plasmid pPIC9K | P. pastoris wild type X-33 |
| [http://partsregistry.org/Part:BBa_K863202 his4] | plasmid pPIC9K | P. pastoris wild type X-33 |
| [http://partsregistry.org/Part:BBa_K863201 3'UTR of aox1] | plasmid pPIC9K | P. pastoris wild type X-33 |
Shuttle vector in E. coli
The developed shuttle vector could transformed in E. coli and analyzed by restriction with the enzymes PvuII and HindIII. The restriction pattern is positiv. A [http://partsregistry.org/cgi/partsdb/dna.cgi?part_name=BBa%20K863204 sequencing] was also done.
Shuttle vector in P. pastoris
After linearization by restriction with EcoRI and SpeI the shuttle vector could be transformed in competent P. pastoris cells. On selective media plates for complementation of histidine auxotrophy some transformants growth (see the picture below). For differentiation between the M+ and Ms genotype a PCR with the Primer 5AOX-Genotyp-FW and TT-Genotyp-RV was done.
GFP integrated in shuttle vector
Cloning of green fluorescent protein (GFP) via the AarI restriction site into the shuttle vector was done for testing the system. A restriction analyze with enzymes ans a sequencing could not be done. The next steps will be (a) linearization with EcoRI and SpeI restriction enzymes, (b) transformation in competent P. pastoris cells, (c) genotype characterization by PCR with 5AOX-Genotype-FW and TT-Genotype-RV primers, (d) cultivation with methanol induction for GFP production and at last (e) fluorescence measure of the cultivation broth.
| 55px | | | | | | | | | | |
 "
"