Team:Potsdam Bioware/Project/Part Virus
From 2012.igem.org
Contents |
Selection Module
Abstract Selection Module
The "selection module" comprises a set of methods to select the desired cell clones from the collection of cells each expressing one variant of the antibody construct. Depending on the needs, different selection methods might have specific advantages. We set up three methods, whereby two are well established but discontinuous: i) fluoresecent activated cell sorting (FACS), which yields many data but requires expensive instrumentation and ii) magnetic beads, which are less expensive and easy to automatize; and last but not least, we develop iii) a novel continuously acting viral selection method. The latter method is based on the assumption that antibody presentation and affinity can be used to modulate viral infection. Virus particles displaying the desired antigen can then either deliver a growth promoting or inhibiting signal. We achieve this by manipulating virus-like particles based on the recombinant Adeno-Associated Virus (rAAV) to present the antigen on the viral surface by either genetic fusion to the capsid protein or by a general enzymatic coupling step using sortase ligation technology. So far we have tested cell sorting (FACS) based on antibody expression, we have coupled YFP, which is the antigen for our test antibody system, to magentic beads, and we have produced rAAV fused at the genetic level to the antigens YFP, CFP and the sortase ligation motif. As a growth inhibiting signal our virus particles deliver a thymidine kinase gene, which enables killing by ganciclovir, and as growth promoting signal a neomycin/geneticin resistance cassette. We are in the process to determine which selection methods works the best.
Introduction
The selection module kicks in when the diverse set of antibodies is expressed on the surface of the cells. In our system each CHO cell couples the genotype of an antibody construct to the phenotype presented on the surface. Therefore we can use all tools already available for selection of cell surface display systems. Reading literature on these methods we opted for fluorescent activated cell sorting (FACS) and magentic beads
The CHO-cells express antibodies presenting them on their surface. To check the antibodies being on the surface of the CHO-cells and being right expressed we focused on different selection systems. Therefore we used several methods like Magnetic Beads, preparative FACS and retargeted adeno-associated virus (AAV). We separated the viral selection system in two parts according to the presented antibodies on the surface of the CHO-cells. We designed the recombinant AAV2 to express the required antigens and to present them on the surface. One of the selection systems is based on the GFP-nanobody - YFP-antigen – complex and another selection system is based on the anti-EGFR-antibody - EGFR complex. After a stable antigen-antibody-complex was formed, the virus infects the cells with a necessary gene for detection and selection.
AAV2 Selection system
Selection system of cells with Advanced Antibody Construct
The first selection system is based on the expression of several fluorescence proteins. The CHO-cells present on their surface the GFP-nanobody. This nanobody binds GFP and YFP with a high affinity and high specifity. The recombinant AAV carries YFP on the surface to target specifically the CHO-cells with GFP-nanobody and to infect them.
To examine the effect of the enzyme AID stably trasfected in the CHO-cells to induce the hypersomatic mutation, we designed a recombinant virus with CFP on the surface. GFP-nanobody expressed on CHO-cells binds YFP, but not the closely realted CFP. Our goal was to show the AID mutates the GFP-nanobody to an antibody with a higher affinity to CFP. Formation of a stable antigen-antibody complex enables the virus binding to the cell and infect it with a mVenus-signal. By excitation the infected cells could be visualised by shining with yellow colour.
To select the cells with the desired antigens the Virus gives the cells a survival signal by infecting them. Virus with the same CFP-antigen on the surface couldn't bind to the GFP-nanobody. Altered by the AID GFP-nanobody binds CFP linked to the virus following by the infection with Kanamycinresistance genecassette. The Kanamycin/Neomycin gene codes for the aminoglycoside phosphotransferase enzyme, which inactivates by phosphorylation the aminoglycoside antibiotics such as geneticin (G418). Cultured in the correct Geneticin concentration, which was determined by titration against nontransfected controls, selection for stable transfectants is possible.
Selection system of cells with Smaller Antibody Construct
The second selection system consists of the EGFR-antigen binding to the anti-EGFR-antibody (scFv425). The scFv425 has a high affinity for EGFR-antigen. After a stable binding the cell would be transduced with transgene encoding for cyan fluorescence protein CFP. The fluorescence was detected at the CFP channel by an excitation at 405 nm and an emission at 425–475 nm. A higher transduction rate would mean the modification of anti-EGFR-antibody to a higher affinity to EGFR which was driven by Activation-induced cytidine deaminase (AID). The selection of cells with improved antibody also relies on the transduction of cells with Kanamycin resistance gene cassette.
AAV2 Genome
AAV is a DNA virus with single-stranded linear genome of 4.7 kb. The genome consists of two open reading frames rep and cap which are flanked by two inverted terminal repeats (ITR).
The rep gene located on the 5'-end of the genome encodes four overlapping multifunctional regulatory proteins. The transcription of the Rep proteins begins at the promoters p5 (Rep 78/Rep 68) and p19 (Rep 52/Rep 40). The two larger Rep proteins Rep78 and Rep68 are required for genome replication, the regulation of gene expression, and for site specific integration. The smaller Rep proteins Rep52 and Rep40 are involved in the encapsidation of the AAV-Genome.
The open reading frame cap on the 3'-end of the AAV-genome encodes the three viral capsid proteins VP1, VP2, and VP3. The transcription of the mRNA is regulated by the p40 promoter. Alternative splicing results in two mRNA the one encodes VP1 and the other VP2/ VP3. Appropriation (Verwendung) of an alternate splice acceptor removes the first AUG start codon for VP1. Thereby mainly formed mRNA encodes the VP2 and VP3 capsid proteins. The first AUG codon in the transcript is the initiation codon for VP3. The translation of VP2 begins at an upstream ACG non-methionin start codon resulting in an approximately 10-fold lower translation of VP2. The stop codon of the three capsid proteins is identical. The stoichiometric ratio of the three capsid proteins VP1, VP2 and VP3 forming the icosahedral form is 1:1:10.
AAV2 Infection
AAV2 viruses kill cancer cells without harming healthy ones. Viruses cross the membrane by a process known as penetration.
The virus particals present in the cell media come closer to the cells by diffusion. After several attachments to the cell membrane the virus binds to the cell receptors and enter the cell by receptor mediated endocytosis. There are three cell receptors playing a major role in infectivity. Ubiquitously expressed cell surface Heparan sulfate proteoglycan (HSPG) functions as a primary receptor account for the broad host range of AAV . aVß5 integrin and fibroblast growth factor 1 (FGFR-1) have a co-receptor activity for successful viral entry into the host cell. Bound AAV particles enter the cell very rapidly through clathrin mediated endocytosis. Each endosome carries a single AAV particle. After internalization the virus requires acidic environment for sufficient realease into the cytosol. AAv accumulates in the perinuclear and entry through the nuclear pore complex (NPC) into the nucleus.There uncoating and gene expression take place.
AAV2 Lytic and lysogenic pathways
After entry the nucleous, AAV follows either lytic or lysogenic life cycles. In the presence of a helper virus, Adenovirus or Herpesvirus, the AAV is directed in a lytic life cycle. AAV undergoes a productive infection, which includes rapid replication in the nucleous and the afterward release of the new particles into the environment by bursting cells.
If there is no helper virus, AAV establish latency by integration into the host genome. The Rep expression is limited. When AAV infects a human cell, it integrates in a site-specific manner into the chromosome 19, called AAVS1. Essential components are the ITRs and the AAVS1 in cis and Rep proteins 78/68 in trans. Rep78/68 binds to the Rep-binding site (RBS) on ITR and unwinds the ITR at the terminal resolution site (trs).
The productive cycle is initiated by infecting with the helper virus, what affects stronger Rep expression and episomal AAV DNA replication. Rep proteins excise the AAV genes out of the host cell’s chromosome. The excised genes are replicated in the nucleous and the capsid proteins package the replicated genes in the cell. When a large number of these particles have assembled, the helper virus induce cell lysis and release the newly assembled partikles into the environment.
recombinant AAV2
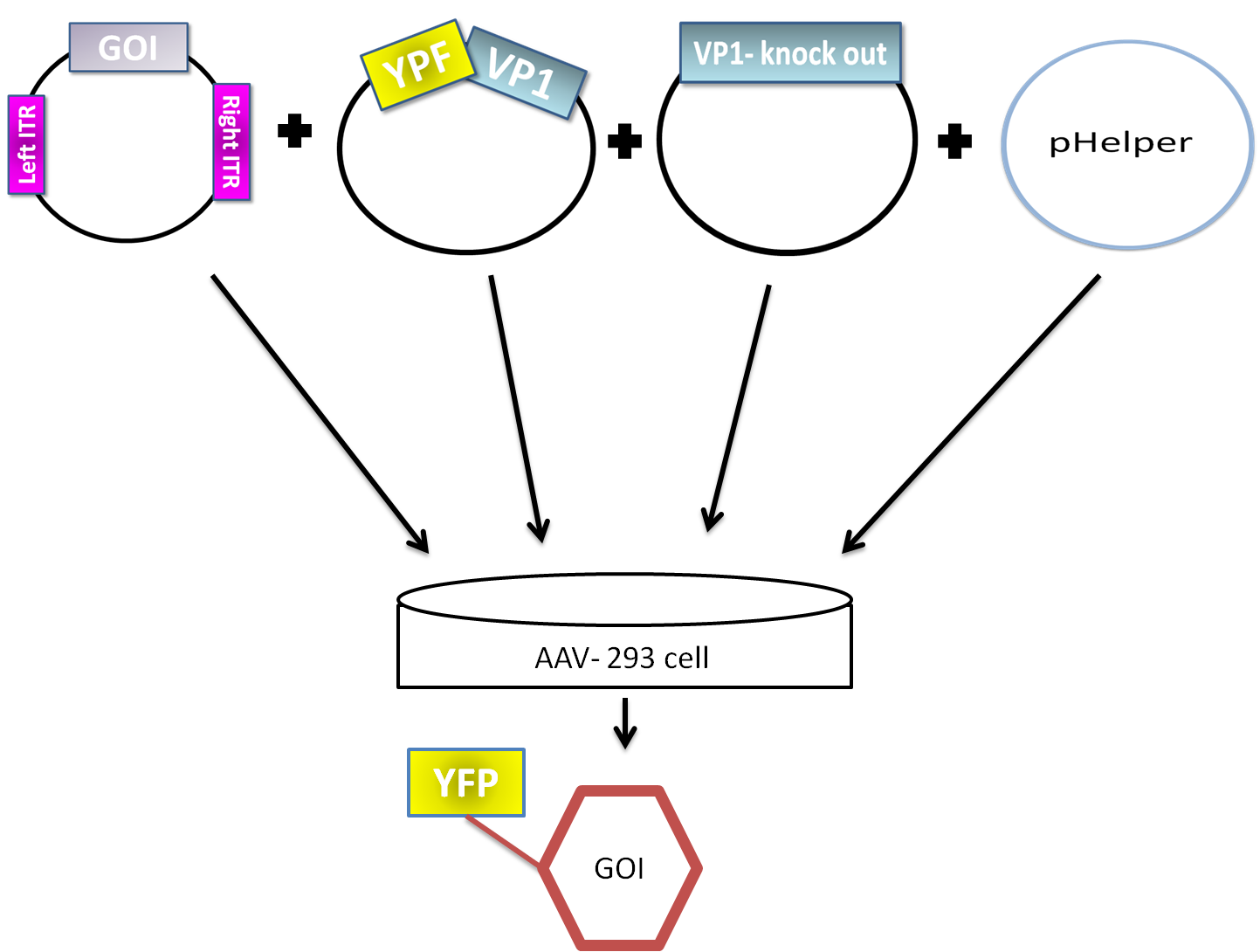
AAV are replication-deficient parvoviruses, although AAV can replicate in the absence of a helper virus, they require a coinfection of adenovirus for an efficient replication. To bypass the co-infection with adenoviruses, which is associated with disadvantages as the contamination of AAV preparation, all of the required genes of adenovirus were encoded on a helperplasmid. The Helperplasmid consiting of adenovirus E2A, VA und E4 genes, is transfected in the AAV-293-cells, which stably express the adenovirus E1 gene. The production of infectious adeno-associated virus particles is able by cotransfecting the AAV-293 cells with four plasmids: as even said the plasmid that carries the helper genes taken from adenovirus, a plasmid coding the protein fused on VP1 or VP2 cap-gene, a vectorplasmid that contains the wildtype AAV2 Rep-Cap genes with VP2 or VP1 knock out and an AAV2 ITR-providing plasmid carrying the gene of interest. The transgenecassette flanked by the ITRs is wrapped as a single strand in the virus capsid. The replication of the Plasmid with transgenecassette is implemented with rep-proteins and enveloped in the viral capsid formed by cap-proteins.
Results
...
targeting the virus against the small construct
To target the virus to the small construct of the CHO-cells we designed plasmids required for viral assembly. EGF-Receptor is a big protein which would disturb the viral assembly, so we focused on the enzyme sortase which covalently attaches proteins to each other. The completely expressed protein EGFR should contain the C-terminal Sortase motif to be recognized by the enzyme Sortase. N-terminal Sortase motif was fused to the N-terminus of VP2/3 gene of adenoassociated Virus in the Biobricks (BBa_K929200) and (BBa_K929201) to allow the ligation by the Sortase.
The first the plasmid with n-terminal sortase motif was designed. The biobrick pCMV_DARPin-E01_Middle-Linker_[AAV2]-VP23 was used to get the fusion parts linked to the VP2/3 cap-gene. The fusion parts Kozak and the N-terminal sortase motif were constructed as a primer close to the VP2/3 – gene, which was used as annealing part. The reversed primer was designed to enclose the whole VP2/3 region. For the following digestion the restriction sites for XbaI and PstI were also presented in the primers.
Amplificate product generated during the pcr was digested with the enzyms XbaI and PstI. Than it was ligated into the biobrick containing the CMV promoter [http://partsregistry.org/Part:BBa_I712004 (Bba_I712004)].
The next the extracellular ligand-binding domain of EGF-Receptor was constructed to be expressed in e. Coli. The coding part for the ligand-binding domain was inserted downstream to the PelB sequence and upstream to the His-tag and c-terminal sortase motif. The constructed plasmid contain the RFC25 accoring suffix and prefix.
To express the EGF-receptor ligand-binding domain in e.coli, the constructed part was ligated in the arabinose containing plasmid. The expression was induced with arabinose and was incubated 4 h by 30 °C. GelBILD
CHO-cells with GFP-nanobody on the surface generated by our group bind YFP. Our task was to produce recombinant virus with YFP on the surface. Such a ligand-receptor complex should provide a targeted and efficient transport of the virus through the cell membrane and also allow the release of the viral DNA in the cell nucleus. To modify the surface exposed proteins of the recombinant virus the desired genes were fused to the N-terminus of VP2 cap-gene. VP2 cap-gene is not essential for viral infectivity, what makes it an ideal candidate for the insertions of sequences to retarget the particle. Therefore to reduce and to define the AAV tropism, the primary receptor motif, heparan sulfat proteoglycan motif, was koncked-out.
The transgene flanked by the ITRs we used in our system was CFP reporter gene. When the cells were infected with the recombinant AAV, the reporter gene was first trascribed in the nucleous and translated to the mature protein in the cytoplasm. The CFP-detection was under UV-light, where the cells should exhibit bright cyan fluorescence. But no cyan fluorescence was detected. The yellow fluorescence was detectible, pointing out that the virus was completly assembled. To check whether the viral assembly worked we infected the AAV-HT1080 cells with the recombinant virus, like it was described in the AAV Helper-Free System Instruction Manual. The next day several HT1080 cells were gleaming cyan by the excitation.
Discussion
Materials and Methods
quantitative real-time PCR
The quantitative real-time polymerase chain reaction (qPCR) is one of the leading methods of the biological research. The significant difference between PCR and qPCR is the detection via fluorescence. The change in the fluorescence intensity can be detected in each cycle. This fluorescence is proportional to the amount of the PCR product. This technique allows the user to follow the PCR reaction in real time. Because of the digital acquisition of the fluorescence, there are no more post-PCR steps (like gel electrophoresis) necessary. (Kubista, M et al. 2006)(Heid, Chr. et al. 1996). Using a standard curve of known sequence copies, the amount of DNA of the samples can be exactly detected. For this detection an intercalating fluorescent dye, like SYBR green, is used. SYBR green is an unspecific florescence dye, which binds within the minor groove of the double stranded DNA. By binding into the DNA, the fluorescent signal increases significantly and can be detected.
We used the qPCR to determine the genomic virus titer. Therefore, the cellular and plasmid DNA left in our virus stocks have to be digested. For that reason 5 µL of the supernatant from pelleted cell lysate was treated with 7.5 µL DNaseI and 5 µL MgCl2 (c= 50 mM) in a final volume of 50 µL. Additionally, the samples were incubated at 37 °C for 30 min with a subsequent inactivation at 65 °C for 10 min. PCR reactions were carried out with 5 µL of our digested samples. (Rohr et al. 2002, Rohr et al. 2005)
References
- Russel and Kay et al. (1999), Blood [http://www.ncbi.nlm.nih.gov/pubmed/10419876pubmed]
- Bartlett et al. (1999), J Virol [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC111768/pubmed]
- Kubista M.; Andrade, JM.; Bengtsson, M.; Forootan, A.; Jonak, J; Lind, K.; Sindelka, R; Sjoback, R.; Sjogreen, B.; Strombom, L.; Stahlberg, A.; Zoric, N.; “The real-time polymerase chain reaction”, Mol Aspects Med. 27(2-3):95-125, 2006.
- Heid, Chr.; Stevens, J.; Livak, K.; Williams, P.; “Real Time Quantitative PCR”, Genome Reserch 6: 986-994, 1996
- Rohr, U.-P. et al., 2005. Quantitative real-time PCR for titration of infectious recombinant AAV-2 particles. Journal of virological methods, 127(1), pp.40-5. [http://www.ncbi.nlm.nih.gov/pubmed/15893564pubmed].
- Rohr, U.-P. et al., 2002. Fast and reliable titration of recombinant adeno-associated virus type-2 using quantitative real-time PCR. Journal of virological methods, 106(1), pp.81-8. [http://www.ncbi.nlm.nih.gov/pubmed/12367732pubmed]
 "
"