Team:Chalmers-Gothenburg/Biodetection of hCG
From 2012.igem.org
Contents |
Biodetection of the hCG Hormone
Our project idea is to develop a biosensor for human chorionic gonadotropin (hCG), which is a hormone that is produced in the body during pregnancy. Consequently, by using this new technique for detecting the hCG hormone, we would like to create a biodegradable pregnancy test kit.
To construct the biosensor, we will use Saccharomyces cerevisiae and replace its endogenous G protein-coupled receptor, Ste2, with the human luteinizing hormone receptor (LH/CG), which is the receptor that binds to hCG with high affinity. For an output signal, we will also attempt to enable Saccharomyces cerevisiae to produce indigo in response to detecting hCG. This will be done by coupling our receptor and genes needed for indigo production, with the yeast pheromone pathway.
Novel and interesting aspects of this project
There are several novel aspects of this project. The receptor that we will utilize in our biosensor has never been expressed in yeast before. In addition, production of indigo has never successfully been carried out in yeast earlier and we hope that indigo can serve as a new and efficient reporter. We would also like to illustrate the usefulness of exploiting the pheromone pathway in yeast when constructing a biosensor. Naturally, we also hope that our new technique for detecting hCG could contribute to the development of a cheap, biodegradable and reusable pregnancy test.
The design of the biosensor
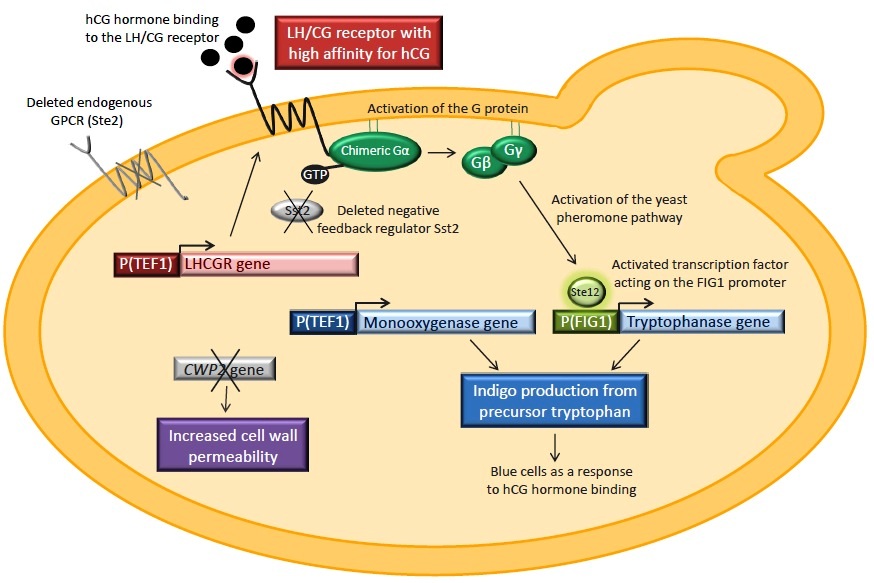
The yeast strain that we used when constructing our biosensor has kindly been provided to us by the Kondo group at Kobe University in Japan. This strain, called IMFD-73, has been optimized for expression of heterologous GPCRs and for coupling of these to the yeast pheromone pathway. The strain has the endogenous G protein-coupled receptor, Ste2, deleted in order to prevent it from isolating G proteins in inactive complexes. Sst2, a negative feedback regulator of the pheromone pathway, is also knocked out to increase the sensitivity of the signaling system. The strain also contains a yeast/human chimeric Gα subunit which should ensure efficient coupling of the human GPCR with the yeast pheromone pathway.
In this strain, we will express the LH/CG receptor. The receptor will be fused to either of two signal peptides: the native human one or the signal peptide from a yeast membrane protein. When expressing the receptor, both signal peptides will be tested. The genes tnaA and fmo, encoding tryptophanase and flavin-containing monooxygenase respectively, will also be introduced into the yeast strain. These enzymes catalyze the conversion of tryptophan to indigo. tnaA will be coupled with the pheromone pathway by setting it under the control of the pheromone-induced FIG1 promoter whereas the fmo will be expressed constitutively. Finally, we will also delete the CWP2 gene. This gene encodes a cell wall mannoprotein and by deleting the gene we hope to increase the cell wall permeability. This is done in order to improve the chances of the hCG hormone to pass the yeast cell wall and thus reach and bind to the receptor.
Hence, this is how the biosensor should work: when hCG binds to the LH/CG receptor this will activate the chimeric G protein, which in turn activates the pheromone pathway. Ultimately this will induce the expression of the tnaA gene and tryptophanase will be produced. Thus, both indigo-synthesizing enzymes are now present and indigo-production is enabled. As an output signal to detecting the hormone, cells should therefore turn blue.
An overview of the biological construct that should serve as a biosensor for hCG. The LH/CG receptor is expressed in yeast. STE2 and SST2 are deleted and the strain expresses a human/yeast chimeric Gα subunit. The tryptophanase gene is under the control of the pheromone-induced FIG1 promoter whereas the monooxygenase is expressed constitutively. CWP2 is removed in order to increase the cell wall permeability.
How to construct the biosensor
Expression of LH/CG receptor
We will express the The LH/CG receptor with either the native or a yeast signal peptide. The map of the plasmid that we will construct and transform yeast with can be seen below.
The map of the plasmid that we will insert into yeast. The LHCGR gene is set under the control of the strong, constitutive promoter PTEF1.
Expression of indigo synthesizing enzymes
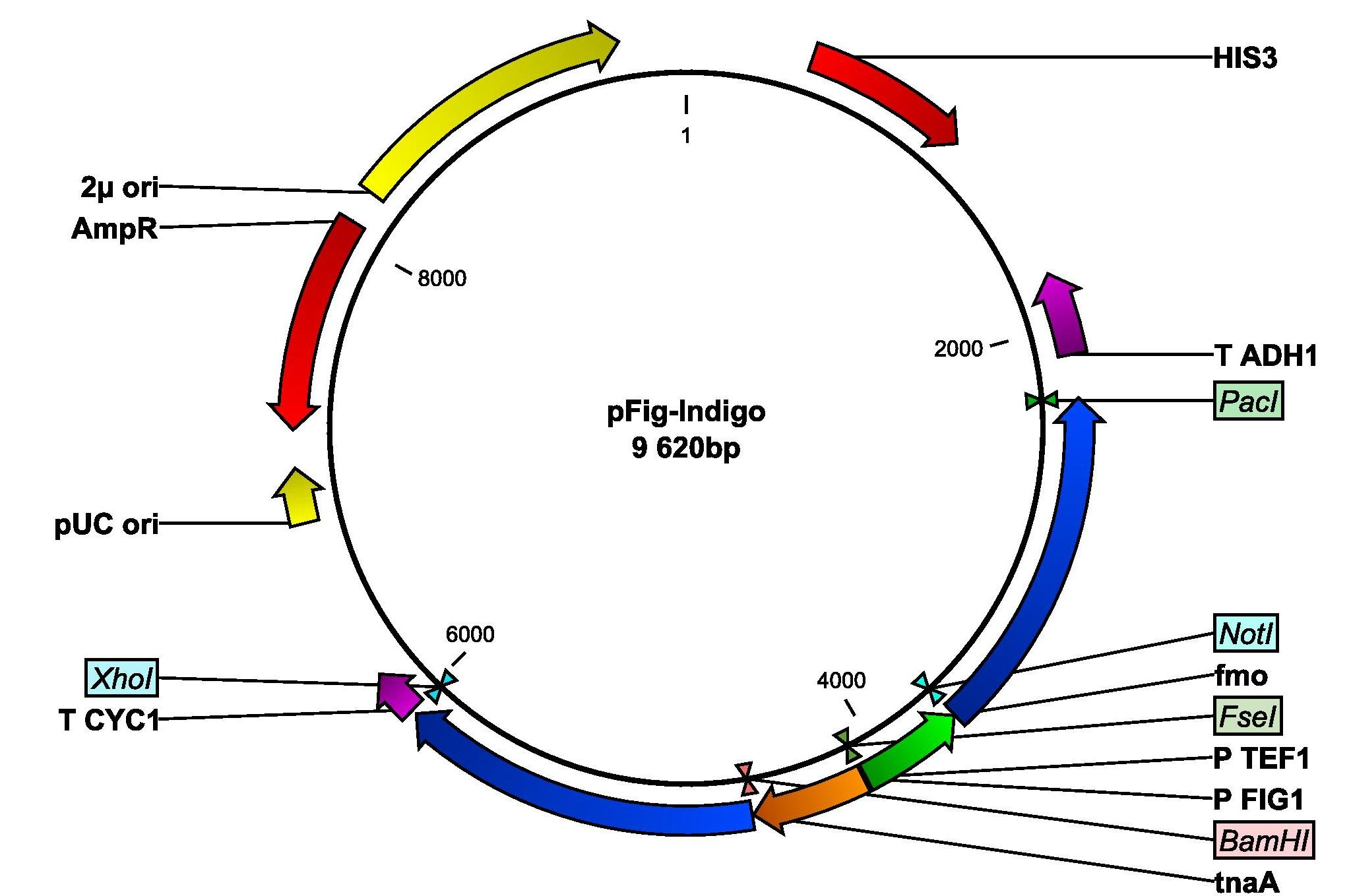
We will insert the two genes fmo and tnaA into yeast with the following plasmid. The tnaA gene is amplified from E.colis genomic DNA and is set unter the control of the pheromone-induced FIG1 promoter. The fmo gene, kindly provided by Si Wouk Kim from Chosun University in South Korea, is set under the control of the constitutive TEF1 promoter.
Plasmid map of the PFigIndigo that will be inserted into yeast. The plasmid contains the genes fmo and tnaA whereof the tnaA genes set under the control of the pheromone-induced FIG1 promoter and thus coupled with the pathway. The fmo gene is set under the control of the TEF1 promoter.
Deletion of the CWP2 gene
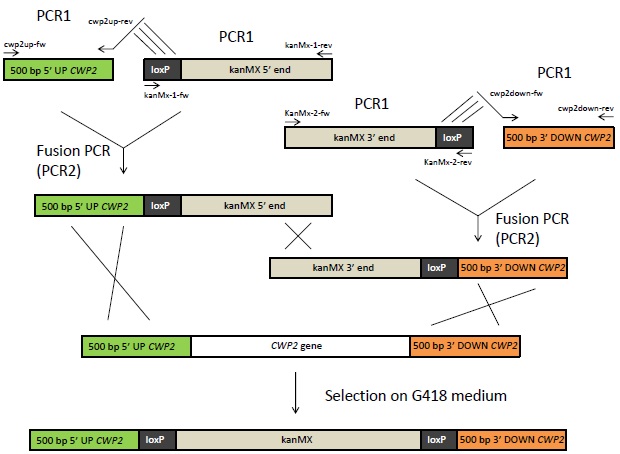
The yeast cell wall limits the permeability and large molecules will not easily pass through it [1]. Hence, if a ligand of a heterologously expressed GPCR is larger than 70-80 amino acid residues, there is risk that it will not be able to activate the receptor [1]. The hCG hormone, the ligand in this project, has two subunits consisting of 92 and 145 amino acids respectively [2] and this could constitute a problem for the hCG biosensor. To solve this problem, we delete the CWP2 gene, which encodes a cell wall mannoprotein. This will probably increase the cell wall permeability and hopefully allow hCG to pass through the cell wall. The gene deletion is attempted by using the Bipartite method, which is outlined below.
Overview of the Bipartite method. 500 bp of the up- and downstream sequence of the CWP2 gene will be amplified from genomic DNA using primers with 5' extensions that are complement to the loxP sites as indicated with lines above. The 5' and 3' end of the kanMX (kanamycin resistance) gene flanked to loxP sites, available in vectors, will also be amplified by PCR. The fragments from these PCR1 reactions will be fused by Fusion PCR. The resulting fragments of this reaction can then be inserted into yeast. Homologous recombination will lead to the exchange of CWP2 with kanMX in the yeast's genome. The cells can be selected on G418 medium.
Results
Spot test analysis and growth measurements in urine were performed in order to test the survival of yeast in urine. These initial experiments showed that yeast cells are able to survive in urinary medium which is an important prerequisite for the biosensor.
The gene CWP2, encoding a mannoprotein in the cell wall, was successfully deleted in the yeast strain IMFD-73. The deletion was confirmed by colony PCR. In addition, a lyticase assay showed that the cell wall in [IMFD-73 Δcwp2::kanMX] was degraded faster than in IMFD-73 and this leads us to the conclusion that the cell wall is somewhat weakened in the strains with the deleted cell wall mannoprotein CWP2.
Two different plasmids both containing the receptor gene LHCGR but one with the human signal peptide and the other one with a yeast signal peptide were created. Both plasmids were successfully cloned into IMFD-73 and [IMFD-73 Δcwp2::kanMX] but in neither strains could the receptor be proved as functional in detection of hCG.
The Indigo group managed to obtain different colour changes in different growth media by introducing a plasmid containing the genes tnaA and fmo (pIndigo). For instance, blue bubbles could be seen in one medium and other media turned into different shades of brown. These changes of colour were strictly dependent on the presence of pIndigo and could not be detected in any of the cultures with the strain containing the control plasmid. This indicates an at least partial activity of the pathway. However, we did not manage to confirm the presence of bio-indigo and to determine exactly which compound or reaction that was responsible for the colourful bubbles or colour changes.
[IMFD-73 Δcwp2::kanMX] was successfully transformed with both receptor genes and genes required for bio-indigo production. However, the system was never functional as a pregnancy test kit since it did not give any significant response in the presence of hCG. Have a look in our result section for more detailed analysis of the results and the discussion and further perspectives of the project.
Submitted parts
[http://partsregistry.org/Part:BBa_K753000 BBa_K753000]: Yeast FIG1 promoter
Promoter of the FIG1 gene in yeast. The transcription factor Ste12, activated in the yeast pheromone pathway, binds to the FIG1 promoter and initiate the expression of the FIG1 gene.
Five pheromone response elements (PRE, consensus sequence ATGAAACA) have been identified in the 5'-upstream regulatory sequence of the FIG1 gene. Phosphorylated Ste12 binds to them and thereby activates transcription. The PREs in the FIG1 promoter differ slightly from the PRE consensus sequence ATGAAACA:
Sequence Location
TGAAACG -405
TGAAACC* -217
TGACACA* -206
TGAAACG -173
TGAAATA -133
Sites marked with an asterisk occur on the opposite strand
The strictly regulated transcriptional feature of the FIG1 promoter can be useful when investigating the pheromone pathway. The promoter can be used to express reporter genes in response to activation of the pheromone pathway.[3]
References
[http://www.cell.com/trends/biotechnology//retrieve/pii/S0167779905001356?cc=y] Ladds G, Goddard A, Davey J. Functional analysis of heterologous GPCR signaling pathways in yeast. Trends in Biotechnology. 2005;23(7):367-373. [http://www.unboundmedicine.com/medline/ebm/record/22027338/abstract/hCG_five_independent_molecules_] Cole LA. hCG. Five independent molecules. Clinica chimica acta; international journal of clinical chemistry. 2012;411(1-2):48-65. [http://www.google.com/url?sa=t&rct=j&q=&esrc=s&source=web&cd=2&ved=0CC8QFjAB&url=http%3A%2F%2Fwww.ncbi.nlm.nih.gov%2Fpmc%2Farticles%2FPMC2140177%2F&ei=D21QUI-wEML80QX7p4DIAg&usg=AFQjCNEkaOP9TOz-IPF5IM0YNrEIWFHBhQ&sig2=SSi3dFbhddcLFi3-0kk-og] Erdman S, Lin L, Malczynski M, Snyder M. Pheromone-regulated Genes Required for Yeast Mating Differentiation. The Journal of cell biology. 1998;140(3):461-483.
 "
"