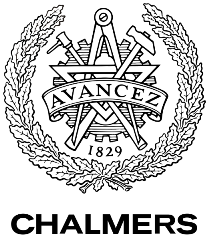Team:Chalmers-Gothenburg/Notebook
From 2012.igem.org
Week 1, 4th - 8th of June
Monday, 4th of June
The laboratory work has begun and after preparing this work for several months we are all very excited to finally get it started. Today we started preparing sterile equipment and we also wrote risk declarations for all of our planned experiments. We also introduced ourselves and held a brief presentation of our project to the very nice people working at the department of System’s biology here at Chalmers University of Technology.Tuesday, 5th of June
Today our dear yeast strains that were kindly provided to us by the Kondo research group of Kobe University, Japan, were transferred to new and fresh YPD agar plates. We also prepared buffers needed for making competent E. coli and we inoculated E. coli DH5α in LB medium.
Wednesday, 6th of June
We prepared a stock of competent E. coli DH5α. Today we also amplified the shuttle plasmids pSP-GM1 and pIYC04 that we will use in our laboratory work, by transforming E. coli DH5α. The cells were then grown on LB-Amp agar plates. We also worked with the team wiki page and created a banner and a logo for the site.
Thursday, 7th of June
Not much laboratory could be done today. Two colonies from each plate containing the E. coli DH5α that were transformed with either pSP-GM1 or pIYC04, were inoculated in ampicillin-containing media and were grown overnight. For the rest of the day, we worked with creating a team brochure and we also established a Facebook page for the team. Click [http://www.facebook.com/ChalmersGothenburgiGEMTeam2012 here] to visit our page and Like our team!
Friday, 8th of June
We purified our pSP-GM1 and pIYC04 plasmids from the transformed E. coli.
Week 2: 11th - 15th of June
This week, the real lab work began. In order to create the biodegradable pregnancy test kit, we have three main tasks to solve. These are 1) the expression of the receptor, 2) the insertion of the indigo producing enzymes and 3) the deletion of a cell wall protein in order to enhance the chances of the ligand to pass the cell wall. Therefore we divided ourselves into three groups, the “receptor group”, the “cell wall group” and the “indigo group” each responsible for its task. Running PCR was a big part of the work this week and we would like to thank Microsynth AG for sponsoring us with the primers.
Receptor group
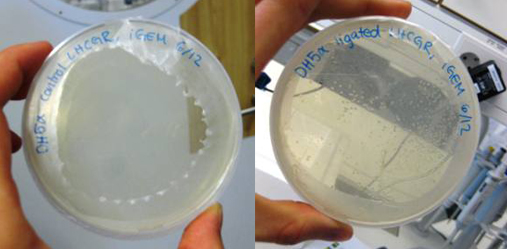
This week, we ran PCR with our lhcgr primers from Microsynth together with our LHCGR construct. This was done in order to obtain the gene for the LH/CG receptor fused with the yeast signal peptide of the LHCGR construct. We religated our generated PCR product and then transformated E. coli DH5α with it. We also did a control where E. coli was transformed with the non-ligated PCR product. We were very pleased to find that our cells that were transformed with the ligated LHCGR construct were growing very nicely on the LB-Amp plate while the control cells did not.
We purified the LHCGR construct from the transformed E. coli and we then continued our work with cutting out the LHCGR gene, that should now be fused with the yeast signal, from the religated construct and ligate it into our shuttle plasmid pSP-GM1. Simultaneously, the original LHCGR construct was also cut to generate the LHCGR gene with its associated endogenous human signal peptide. After cutting we ran gel electrophoresis to verify correct fragment sizes and the DNA from the bands containing the LHCGR gene with the human respectively yeast signal peptide were extracted and purified along with the cut pSP-GM1. Finally, we also attempted to ligate the two LHCGR inserts into the pSP-GM1 plasmids. Will the ligation work? This we will need to find out next week!
Indigo group
PCR reactions were run for the tnaA and the fmo gene. The tnaA gene was taken directly from E. coli with colony-PCR whereas the fmo gene was kindly provided by Si Wouk Kim from Chosun University, South Korea. After two days we succeeded to purify the fmo gene from gel. When the tnaA gene did not show up on the gel after the two first days we got the advice from Rahul Kumar (who works here in the lab at the department of System’s biology at Chalmers University of Technology) to boil the E. coli cells in a basic solution before the colony PCR. This time the colony PCR worked and the tnaA gene could be purified from the gel. Thanks Rahul!
We made a ligation mixture of the fmo gene with pIYC04 and tried to transform the ligation mix into E. coli. Unfortunately nothing had grown on the ligation plate (nor the control plate) and we had to redo the ligation. So the last day of the week we did two ligations in parallel, one with the fmo gene and pIYC04 and another one with the tnaA gene and pIYC04. These will be transformed into E. coli next week and hopefully one of them will work then.
The cell wall group
We started with the construction of the knock-out fragments. First, we ran PCR with genomic yeast DNA in order to get the 500 up- and downstream fragments of the CWP2 gene. The two overlapping kanMX fragments were also amplified by PCR. For these PCR reactions, we used HPLC purified primers that were sponsored by Microsynth AG. After PCR the fragments were checked and purified on gel.
We have learned that everything takes a little bit longer than expected; it took actually the whole week to get all the four fragments, since some PCR reaction had to be repeated several times. However, we are still on schedule. On Friday, we could then start with Fusion PCR in order to fuse the fragments. Of course it did not work the first time and we have the feeling that this will be the more tricky part of the cell wall deletion. But we will come up with some different approaches next week!
 "
"