Team:NRP-UEA-Norwich/Week3
From 2012.igem.org
(→Week 3) |
(→Week 3) |
||
| Line 7: | Line 7: | ||
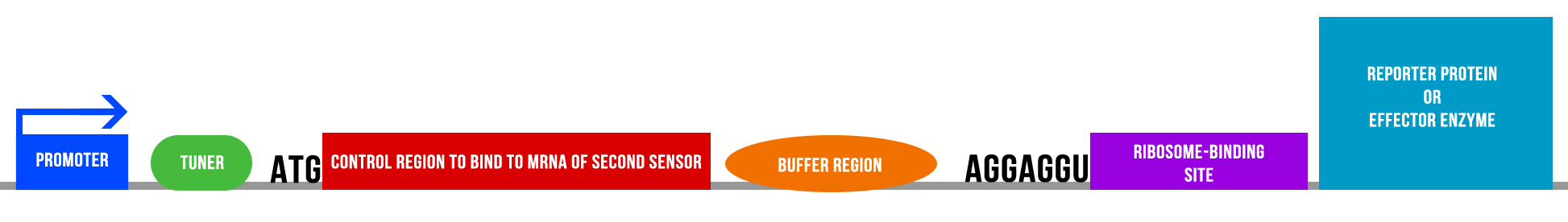
Over the weekend the group met up to discuss the two potential projects we could go forward with using our synthesised gene and the Biobrick it will ultimately produced. We developed two main themes; one was a multi-sensor system sensing oxygen-containing physiologically-relevant molecules and reporting on the levels of each molecule; the other was a comparator circuit sensing levels of molecular species and using novel gene expression to express reporters for specific molecules within the environment to quantitatively measure them. We decided on the latter as we felt it would have the highest scientific impact, a diagram of the gene system for one of the sensors is given below. | Over the weekend the group met up to discuss the two potential projects we could go forward with using our synthesised gene and the Biobrick it will ultimately produced. We developed two main themes; one was a multi-sensor system sensing oxygen-containing physiologically-relevant molecules and reporting on the levels of each molecule; the other was a comparator circuit sensing levels of molecular species and using novel gene expression to express reporters for specific molecules within the environment to quantitatively measure them. We decided on the latter as we felt it would have the highest scientific impact, a diagram of the gene system for one of the sensors is given below. | ||
[[File:NRPDiagram1.png | center | 950px]] | [[File:NRPDiagram1.png | center | 950px]] | ||
| - | The concept behind the project involves the mRNA control region of one sensor binding to the mRNA control region of a second sensor and thus for one sensor to 'switch off' the other to a certain degree, and for the overhang to be used to measure levels of a substrate molecule. For example in our nitric oxide-sensing project a sensor for nitric oxide and nitrites (sensor 1) may be used in conjunction with a sensor for only nitrites (sensor 2); the control region of mRNA produced from sensor 2 would switch off sensor 1 per the levels of nitrite in the environment, and therefore all expression of sensor 1 would be directly related to nitric oxide. | + | The concept behind the project involves the mRNA control region of one sensor binding to the mRNA control region of a second sensor and thus for one sensor to 'switch off' the other to a certain degree, and for the overhang to be used to measure levels of a substrate molecule. For example in our nitric oxide-sensing project a sensor for nitric oxide and nitrites (sensor 1) may be used in conjunction with a sensor for only nitrites (sensor 2); the control region of mRNA produced from sensor 2 would switch off sensor 1 per the levels of nitrite in the environment, and therefore all expression of sensor 1 would be directly related to nitric oxide levels in the atmosphere. |
==Day 1== | ==Day 1== | ||
Revision as of 13:53, 25 July 2012
 "
"