Team:Cambridge/HumanPractices/FutureDirections
From 2012.igem.org
(Created page with "{{Template:Team:Cambridge/CAM_2012_TEMPLATE_HEADNEW}} <html> <script type="text/javascript"> $(document).keydown(function(e){ //e.which is set by jQuery for those browsers t...")
Newer edit →
Revision as of 06:36, 26 October 2012
Contents |
Judging Form
- Please help the judges by filling out this form. Tell them what medal you think you deserve and why. Tell them which special prizes you should win. Help them find your best parts. Show them how you thought about the safety of your project. Helping the judges will help you too.
- Team: Cambridge
- Region: Europe
- iGEM Year:2012
- Track:Foundational Advance
- Project Name:Parts for a reliable and field ready biosensing platform
- Project Abstract: Implementation of biosensors in real world situations has been made difficult by the unpredictable and non-quantified outputs of existing solutions, as well as a lack of appropriate storage, distribution and utilization systems. This leaves a large gap between a simple, functional sensing mechanism and a fully realised product that can be used in the field.
We aim to bridge this gap at all points by developing a standardised ratiometric luciferase output in a Bacillus chassis. This output can be linked up with prototyped instrumentation and software for obtaining reliable quantified results. Additionally, we have reduced the specialized requirements for the storage and distribution of our bacteria by using Bacillus' sporulation system. To improve the performance of our biosensing platform we have genetically modified Bacillus’ germination speed. Lastly, we demonstrated the robustness of our system by testing it with a new fluoride riboswitch, providing the opportunity to tackle real life problems.
iGEM Medals for non-software teams
- We believe our team deserves the following medal:
- Bronze
- Silver
- √Gold
Because we met the following criteria (check all that apply and provide details where needed)
Requirements for a Bronze Medal
- √Register the team, have a great summer, and plan to have fun at the Regional Jamboree.
- √Successfully complete and submit this iGEM 2012 Judging form.
- √Create and share a Description of the team's project using the iGEM wiki and the team's parts using the Registry of Standard Biological Parts.
- √Plan to present a Poster and Talk at the iGEM Jamboree.
- √Enter information detailing at least one new standard BioBrick Part or Device in the Registry of Standard Biological Parts. Including:
- √Primary nucleaic acid sequence
- √Description of function
- √Authorship
- Safety notes, if relevant.
- √Acknowedgment of sources and references
- √Submit DNA for at least one new BioBrick Part or Device to the Registry.
Additional Requirements for a Silver Medal
- √Demonstrate that at least one new BioBrick Part or Device of your own design and construction works as expected; characterize the operation of your new part/device.
- √Enter this information and other documentation on the part's 'Main Page' section of the Registry
Part Number(s): [http://partsregistry.org/Part:BBa_K911004 BBa_K911004]
Additional Requirements for a Gold Medal: (one OR more)
- Improve an existing BioBrick Part or Device and enter this information back on the Experience Page of the Registry.
Part Number(s): None - √Help another iGEM team by, for example, characterizing a part, debugging a construct, or modeling or simulating their system.
Link to this information on your wiki. Page name: Team:Cambridge/Outreach/Collaboration - √Outline and detail a new approach to an issue of Human Practice in synthetic biology as it relates to your project, such as safety, security, ethics, or ownership, sharing, and innovation.
Link to this information on your wiki.
Page name: Team:Cambridge/HumanPractices/Overview,Team:Cambridge/HumanPractices/MarketResearch,Team:Cambridge/HumanPractices/FutureDirections
iGEM Prizes
All teams are eligible for special prizes at the Jamborees. more... To help the judges, please indicate if you feel you should be evaluated for any of the following special prizes:
- √Best Human Practice Advance
- √Best Experimental Measurement
- Best Model
Please explain briefly why you should receive any of these special prizes:
Best Human Practice Advance:
We feel that we deserve this prize for three reasons:
- We explored the impacts, *both positive and negative*, of synthetic biology as a solution to real world problems, through interviewing professionals working in a relevant field, namely the impact of arsenic water contamination in Bangladesh.
- We recognized existing problems with the way the current direction of synthetic. On going through the registry we found that most of the characterization data for biosensing parts is often neither comparable nor replicable. We have worked to solve this issue, for example with our ratiometric dual channel output.
- *Our project doesn’t stop here*, in Chanel number 6 (Team:Cambridge/HumanPractices/FutureDirections) we considered the future implications and technological applications of our project, as well as the means by which it could be improved by subsequent users. We feel that the end to an iGEM project should not be the conclusion of an idea, but the start of it.
Best BioBrick Measurement Approach:
It is absolutely vital that a quantitative, numerical, robust, and flexible measurement approach exists to relay information to a user that is an accurate representation of the input processed by a biological device. Working from these principles, the following was done:
- We designed and built Biologger, a *cheap, arduino-based, fully functional automatic rotary device* that has an incorporated ratiolumnometer
- Our project is entirely open-sourced and open-platform. We have published source code for the two applications which serve to operate the device, one for PCs and the other for Android devices, as well as the open source circuit design that provides this ratiometric reading. Furthermore, the Android app is able to receive its data wirelessly, which we feel is a great advance in BioBrick measurement.
- Our dual-channel luciferase reporter was successfully tested with a dilution series of E.coli transformed with the Lux Operon (under pBAD) biobrick (Part BBa_K325909) of the Cambridge iGEM 2010 team. It can detect, with good accuracy, both different light intensities, as well as the percentages of blue or orange frequencies in a sample.
- Our device was successfully tested using artificial light to detect different frequencies (colours) as well.
Having done all the above, we believe that this fully open-sourced instrumentation kit (mechanical) chassis, electronics, software code), estimated at *$35.00* (or $85.00 if a Bluetooth modem is required), is a complete BioBrick measurement solution for any and all BioBricks with a light output.
Team_Parts
To help the judges evaluate your parts, please identify 3 of your parts that you feel are best documented and are of the highest quality.
- Best new BioBrick part (natural)
- [http://partsregistry.org/Part:BBa_K911003 BBa_K911003]
- Best new BioBrick part (engineered)
- [http://partsregistry.org/Part:BBa_K911004 BBa_K911004]
- Best improved part(s): None
List any other parts you would like the judges to examine:[http://partsregistry.org/Part:BBa_K911001 BBa_K911001], [http://partsregistry.org/Part:BBa_K911008 BBa_K911009], [http://partsregistry.org/Part:BBa_K911008 BBa_K911008]
Please explain briefly why the judges should examine these other parts:
- Magnesium Sensitive Riboswitch [http://partsregistry.org/Part:BBa_K911001 BBa_K911001]
As a riboswitch sensing construct, this part is an entirely new type of biosensor (along with the fluoride construct) that could potentially change the way we think about designing input genetic circuits. Unlike the fluoride riboswitch, it is a derepression system and therefore serves to demonstrate the principle that riboswitches can be used regardless of whether they turn on or off their reporter. - Fluorescent ratiometric construct for standardizing promoter output [http://partsregistry.org/Part:BBa_K911009 BBa_K911009]
Fluorescence is a major cornerstone for biosensors in the registry, however, most parts do not involve the use of a ratiometric output, which has been shown in the literature to provide much more reliable and meaningful data. This part not only furthers the development of ratiometric measurements in molecular biology but due to the choice of promoters and terminators it can be used to characterize the difference in activity between E. coli and B. Subtilis - Fast Germination (B. subtilis) [http://partsregistry.org/Part:BBa_K911008 BBa_K911008]
This part is entirely novel for the registry and fully utilizes the recombination machinery inherent in the Bacillus chassis. Have spores that can germinate at a faster rate is certainly a worthy achievement and could help with experiments with B. Subtilis that any future iGEM teams may wish to perform.
iGEM Safety
For iGEM 2012 teams are asked to detail how they approached any issues of biological safety associated with their projects.
The iGEM judges expect that you have answered the four safety questions Safety page on your iGEM 2012 wiki.
Please provide the link to that page: Page name: Team:Cambridge/Safety
Attribution and Contributions
For iGEM 2012 the description of each project must clearly attribute work done by the team and distinguish it from work done by others, including the host labs, advisors, and instructors.
Please provide the link to that page, or comments in the box below: Page name: Team:Cambridge/Attributions
Comments
If there is any other information about your project you would like to highlight for the judges, please provide a link to your wiki page here: Team:Cambridge/Overview/DesignProcess
Chanel number six
One of the big features of our project is the use of a second fluorescence/ luminescence channel to give a reliable estimate of cell activity, against which reporter activity can be adjusted. However, there is no need to limit ourselves to only two channels - the more channels that are used, the more data that can be squeezed out of a single biosensor strain.
Design considerations
An issue with this proposal is that with additional channels, there comes a greater risk of cross talk between the channels. This is part of the reason we have chosen to isolate our different biosensor strains in separate containers. In essence, each container is a different channel, separated in space rather than in wavelength. However, with the progression of our knowledge of the field of sythetic biology and the development of improved predictive software, we should be able to avoid this problem in the future.
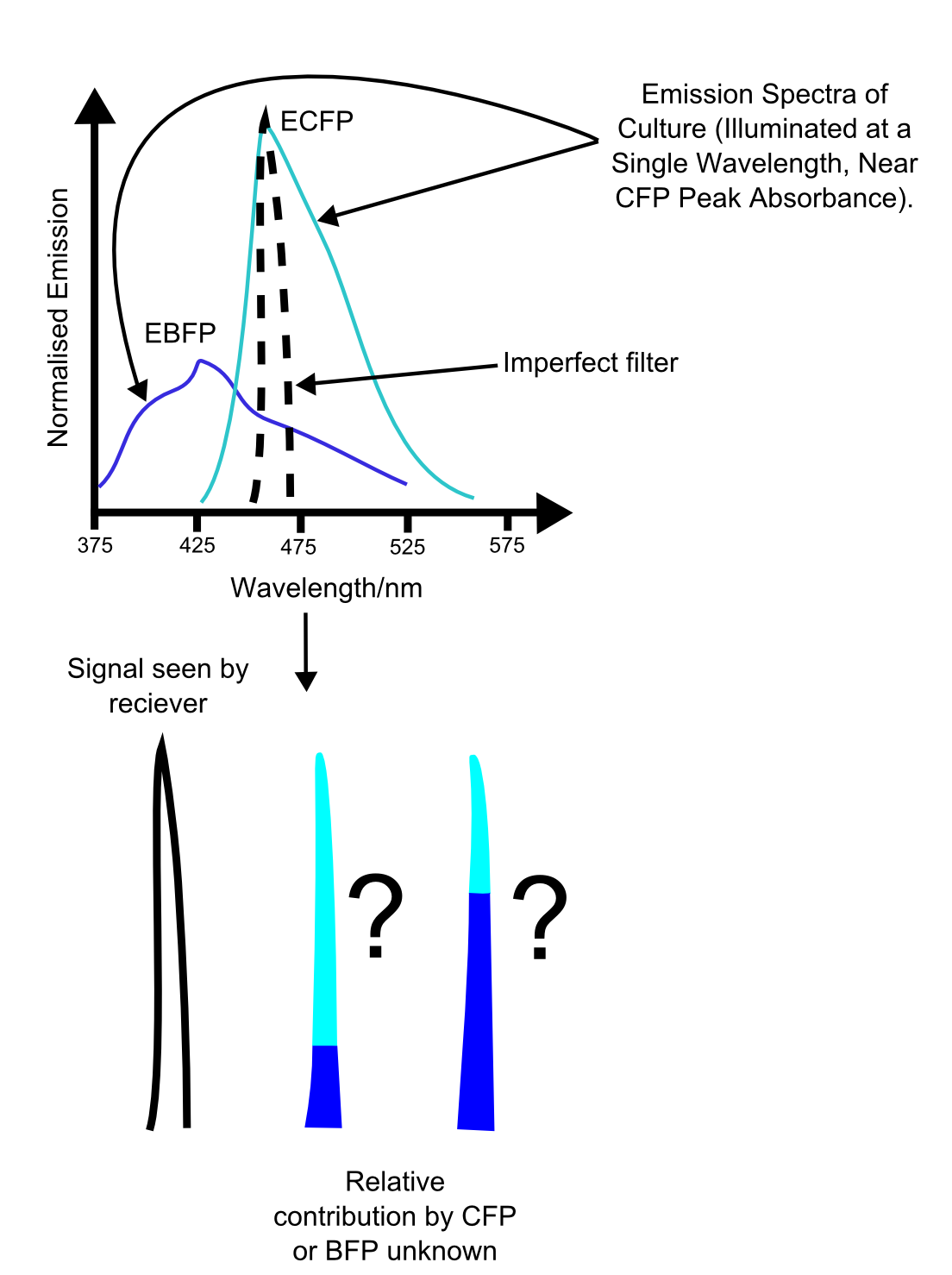
More fundamental is the problem of reliably distinguishing the different channels. While a two channel system is relatively simple to use, use of three or more fluorescent proteins may cause ambiguities in the data, as shown in the figure. These problems are similar to those seen in vision, where the overlap in the spectral sensitivities of the three cone types leads to impairment of our ability to accurately distinguish different hues. In this case though, the main difficulty is with the growth of unreliability of the measurements with increasing numbers of overlapping emission curves.
To get around this, we propose using additional forms of channel. For example, it may be possible to use the frequency of oscillation of a genetic circuit as an output. By using both amplitude and frequency of such an oscillatory system, it may be possible to encode two variables with a single fluorescent protein, effectively doubling the possible number of outputs. It is possible to concieve of other possible output channels - given that light is the most simple thing to detect in microbiological contexts (hence why fluorescent proteins and luminescent proteins are the most popular reporter choices for various assays of microscopic samples), it would be interesting to investigate the possibility of using other properties of the emitted light. Would it be possible to control the polorisation of the light, for example? Such a system would most likely need advances in cell fixation techniques, so as to ensure that the cells are aligned in the same direction.
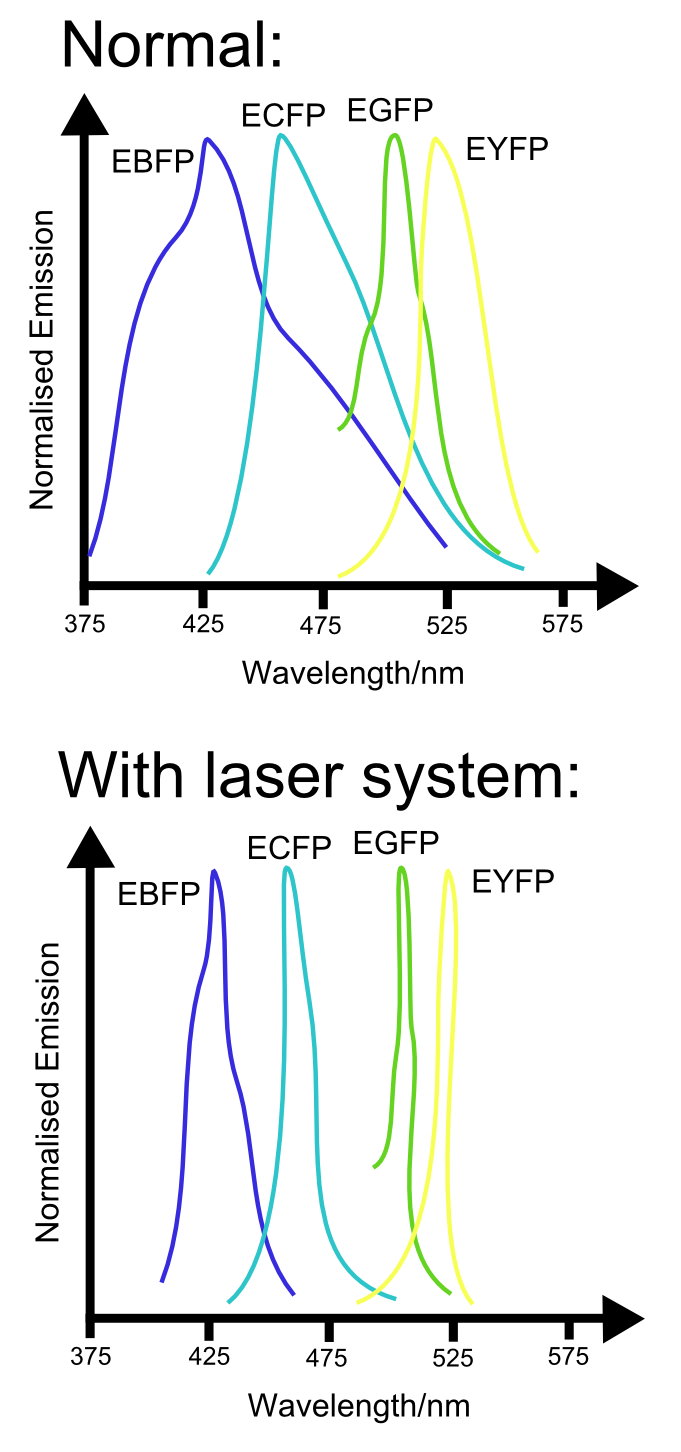
Alternatively, the development of techniques that enable GFP (and, in the future, most likely other fluorescent proteins) to act as lasers may sharpen the emission spectra of these proteins. In doing so, the number of different fluorescent proteins that could be sqeezed into the available spectral range would be greatly increaced. Again, this would enable us to use even more channels.
Unfortuneately, such biological lasers have only been demonstrated in animal cells, and the cell walls of bacteria may interfere with the ability to generate effective lasing. Nevertheless, if a sensing standard within eukaryotic cells is ever developed, this may become a useful tool for increasing the range of such a system.
However, either of these techniques will require considerable leaps in our ability to create complex constructs. Gibson may be suitable, but from our experience it will need to be made more reliable before we use it to make the large constructions needed.
Potential applications
Should the prospect of reliable construction of plasmids with three or more channels become a reality, the oppertunities will be vast. One of the most simple (yet effective) may be the development of improved control systems for bioreactors. A three channel system should be all that is needed for the accurate and speedy detection of changes in the metabolism of cultured cells within the bioreactor. As discussed in [http://schaechter.asmblog.org/schaechter/2009/11/the-limitations-of-lb-medium.html this article], magnesium may often becomes a limiting nutrient within bacterial cultures, causing a distinct shift in the metabolic properties of the cells. One channel would therefore be devoted to the measurement of magnesium in the culture with the magnesium riboswitch we have characterized, causing automatic release of magnesium chloride by the control system into the medium. A second channel would monitor the levels of sigma factors usually only present during the plateau phase of growth, causing the release of fresh media if it detects that growth is about to slow down, and so prevent changes in the metabolic properties of the cells. The third channel would be made from a constitutive channel which would monitor the overall state of the cells, ensuring that their overall metabolism remained within acceptable bounds.
Such a strain would only need to be introduced into the bioreactor in small quantities, allowing the main, economically useful cells to dominate. Small samples would be continually taken, and the fluorescence channels quickly analysed to provide data for the control system.
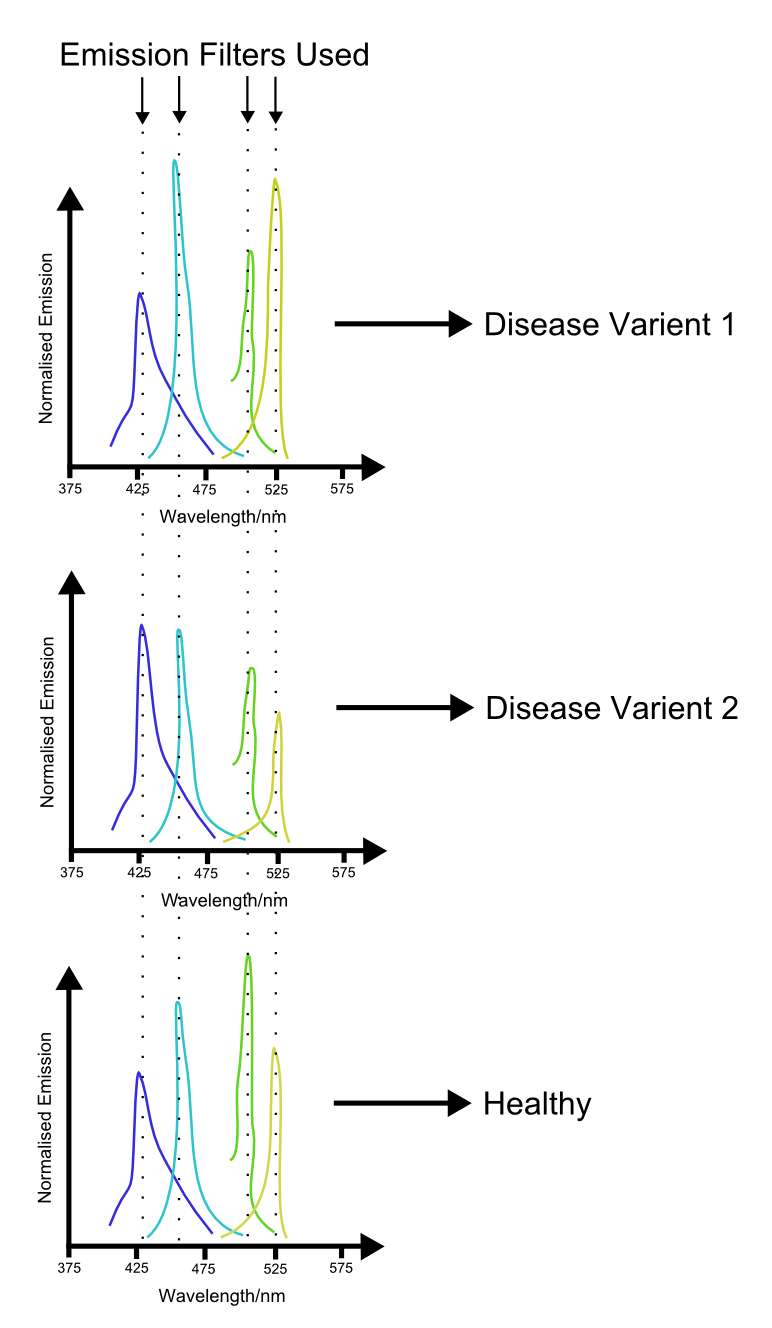
With increacing complexity of our ability to use multiple channels comes increacing complexity of the applications. Further into the future, applications of such a system may include medical science. We propose that multiple indices of health be hooked up to different channels in such a system. Because the biosensing proteins involved in such a proposal are likely to only work in a eukaryotic environment, a eukaryotic chassis will need to be used. A blood sample from the patient is then innoculated with the sensory cells. An assay is performed by monitoring the output of the different channels, potentially using the laser system proposed above. The profile generated by this procedure may then produce a characteristic profile which can then be used to 'fingerprint' different diseases.
For example, a potent indicator of cardiovascular disease is the ratio of total cholesterol and HDL levels. [http://www.sciencemag.org/content/271/5248/518 HDL] and [http://www.sciencedirect.com/science/article/pii/0092867484901880 LDL] receptors exist which, while they do not directly couple to sensory pathways, could be adapted in the future to cause metabolic changes in a reporter cell. A three channel system, with total cholesterol, HDL levels and cell activity as its outputs would give a quick and single reaction assay for cardiovascular health.
Such a system, when combined with methods that are capable of analysing large quantities of data quickly, may also yield useful associations between many previously unchecked variables. With a construct with a series of steroid hormone detectors, for example, we would be able to discover the association between different disease types and different hormone levels. This data could then be put back into the diagnostic side of the system, improving the diagnostic power of the tool.
Extending this 'fingerprinting' idea into the far future, when GMOs have likely become a feature of everyday life, many interesting ideas can be thought of. For example, bacterial 'paints' could be developed which secreted different pigments according to the chemical profile of the underlying surface. After leaving to dry/develop for several hours, the specific chemical properties of the surface could be determined. This could be useful for geochemical surveys before well drilling, circumventing the problems that have been faced by such measures currently and, potentially, rendering our field testing kit obsolete.
Lastly, as the name of this section implies, we think an innovative product would be a perfume which responds to the wearers moods. Previous iGEM teams have already demonstrated the possibility of making e.coli more pleasingly fragrant, with the aromas of wintergreen and banana. Hooking up this output (a multichanel output in chemical space) to an output responsive to compounds in human sweat could, for example, cause the release of soothing eucalyptus oils in response to stress related compounds. How fragrant.
References
- Gather M. and Yun S., Single-cell biological lasers, Nature Photonics (2011) 5, 406–410
- Prospective Studies Collaboration, Blood cholesterol and vascular mortality by age, sex, and blood pressure: a meta-analysis of individual data from 61 prospective studies with 55 000 vascular deaths, The Lancet (2007), 370, Issue 9602, 1829–1839
 "
"