Team:SJTU-BioX-Shanghai/Project
From 2012.igem.org
AleAlejandro (Talk | contribs) (→Introduction) |
AleAlejandro (Talk | contribs) (→Introduction) |
||
| Line 44: | Line 44: | ||
Previous researchers have focused on building protein, RNA or DNA scaffold as constitutive assemblies carrying enzymes. They have succeeded in increasing product yields. However, the amount of those scaffolds could be limited by its expression or copy level, leading to restriction on further acceleration. With ''Membrane Magic'', we made ''E.coli'' membrane into a huge scaffold accommodating enzymes without limitation of scaffold amount. Moreover, protein assembly on membrane could readily receive extracellular or intracellular signal, so the whole system becomes highly tunable. | Previous researchers have focused on building protein, RNA or DNA scaffold as constitutive assemblies carrying enzymes. They have succeeded in increasing product yields. However, the amount of those scaffolds could be limited by its expression or copy level, leading to restriction on further acceleration. With ''Membrane Magic'', we made ''E.coli'' membrane into a huge scaffold accommodating enzymes without limitation of scaffold amount. Moreover, protein assembly on membrane could readily receive extracellular or intracellular signal, so the whole system becomes highly tunable. | ||
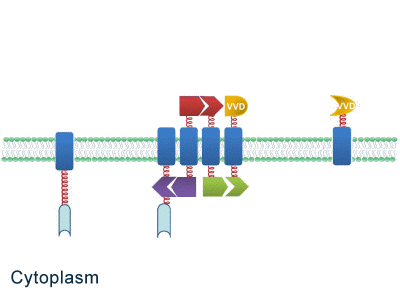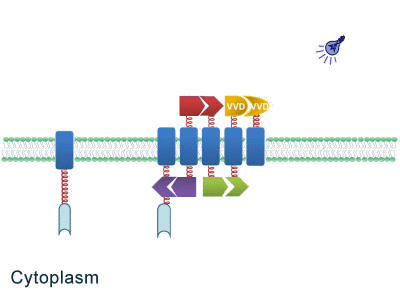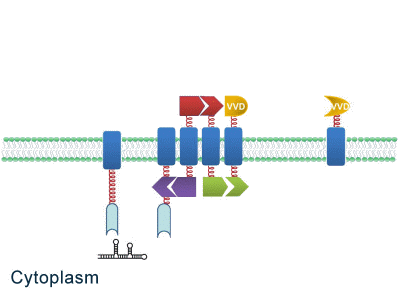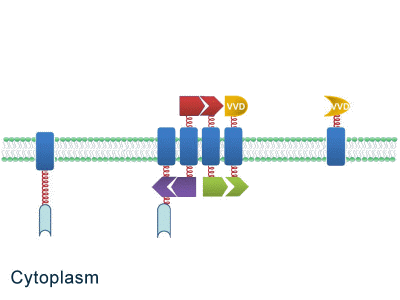
| - | One of our devices, called ''Membrane Accelerator'', functions by localizing and organizing enzymes on membrane surface. ''E.coli'' inner membrane serves as a two-dimensional plane that can accommodate various protein assemblies linked with enzymes. Otherwise diffusing enzymes can form clusters on membrane through interacting protein domains and ligands. Enzyme clusters help substrates flow between enzymes, and thus increase yields of sequential biological reactions. We not only applied the ''Membrane Accelerator'' into biosynthetic pathway but also biodegradation pathway, which is proposed | + | One of our devices, called ''Membrane Accelerator'', functions by localizing and organizing enzymes on membrane surface. ''E.coli'' inner membrane serves as a two-dimensional plane that can accommodate various protein assemblies linked with enzymes. Otherwise diffusing enzymes can form clusters on membrane through interacting protein domains and ligands. Enzyme clusters help substrates flow between enzymes, and thus increase yields of sequential biological reactions. We not only applied the ''Membrane Accelerator'' into biosynthetic pathway but also biodegradation pathway, which is proposed for the first time in synthetic biology. Previous researches on scaffold system all focused on biosynthesis. |
<html> | <html> | ||
| Line 52: | Line 52: | ||
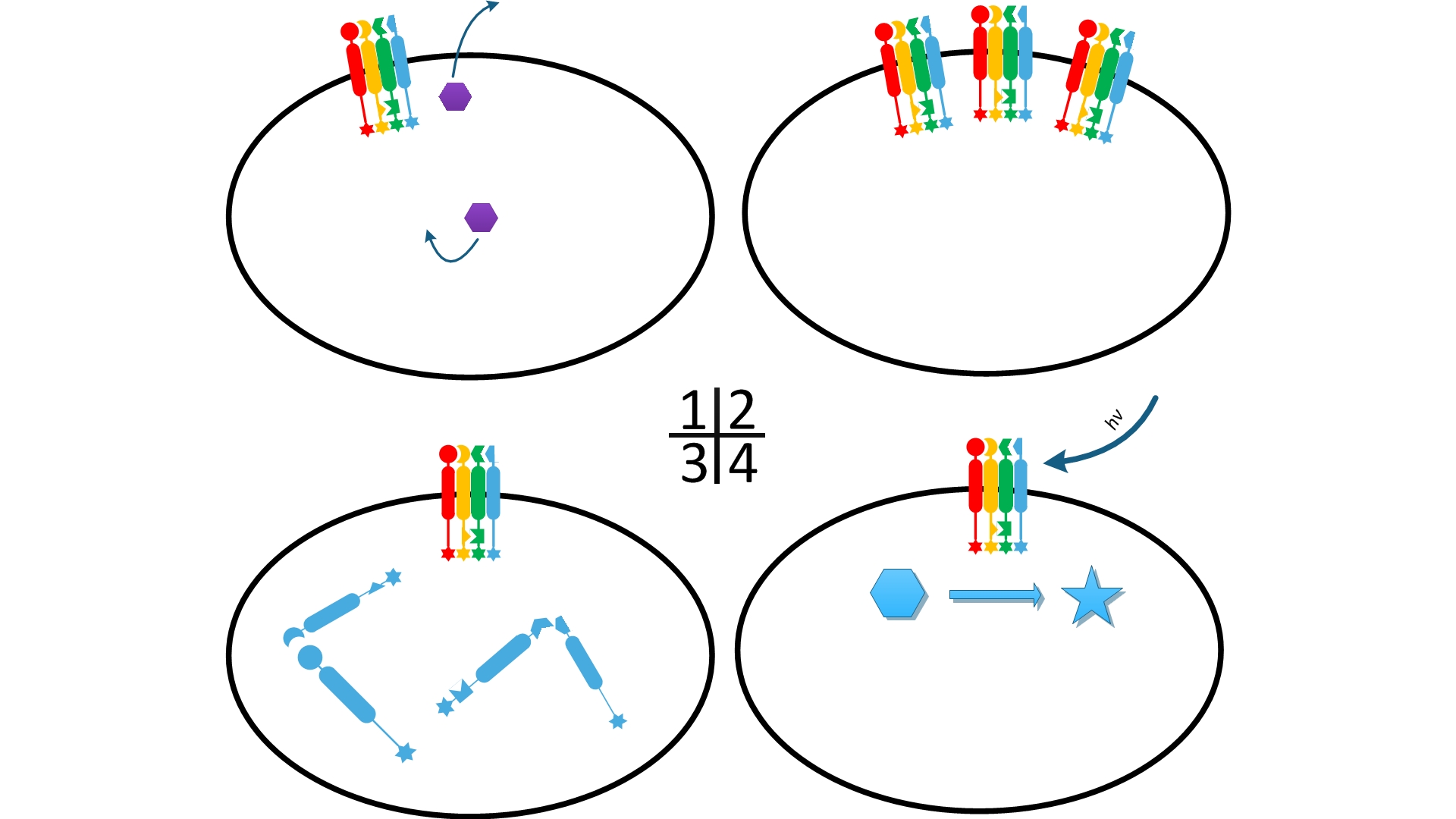
''Fig.1:'' Sketch of ''Membrane Accelerator'' | ''Fig.1:'' Sketch of ''Membrane Accelerator'' | ||
| - | Although some work has been done in reaction acceleration, | + | Although some work has been done in reaction acceleration, it has always been a challenge to artificially and dynamically control those reactions. Our ''Membrane Rudder'' device, however, offers a novel method to control the direction of biochemical reactions through varieties of signals. We further combined the whole post-translational control system with genetic circuits by recruiting RNA aptamer and its corresponding binding protein. Thus RNA signal could also be recruited to dynamically control biochemical reaction. |
<html> | <html> | ||
Revision as of 08:55, 23 October 2012
| ||
|
 "
"