Team:Amsterdam/data/experimental
From 2012.igem.org
(Difference between revisions)
| Line 80: | Line 80: | ||
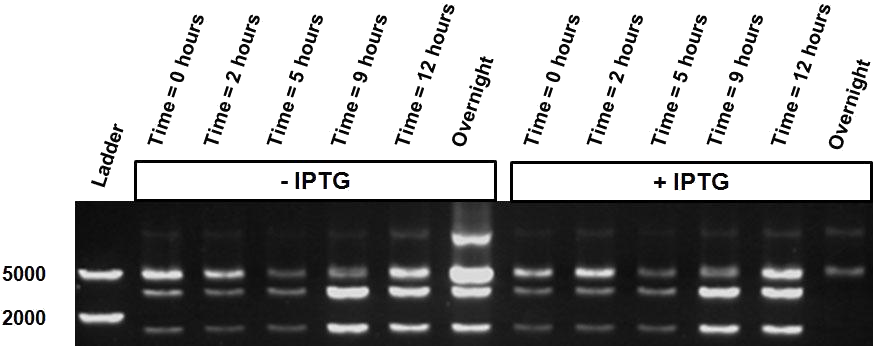
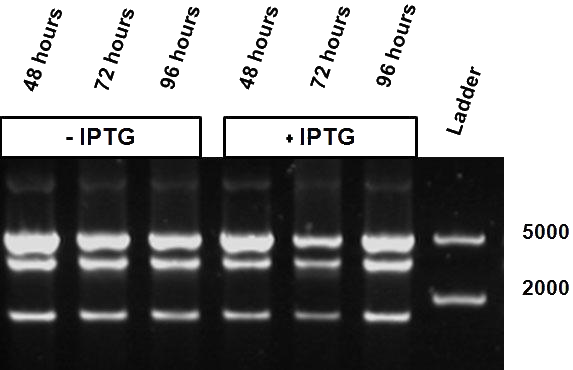
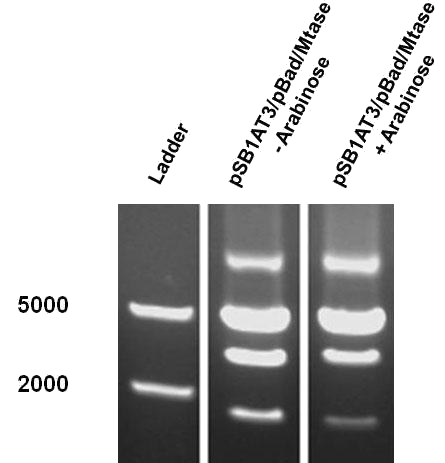
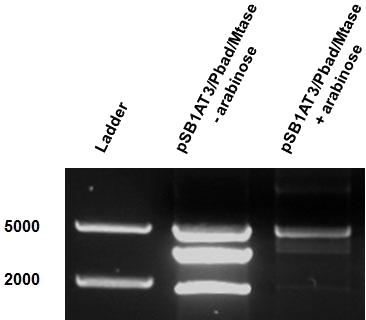
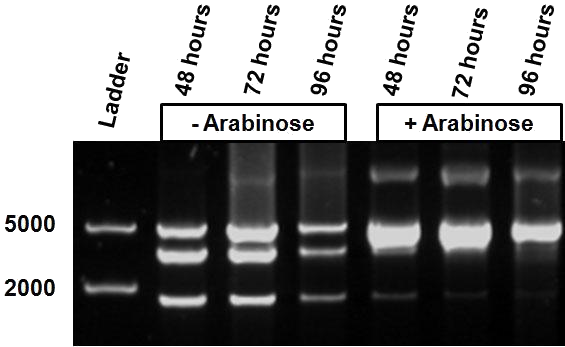
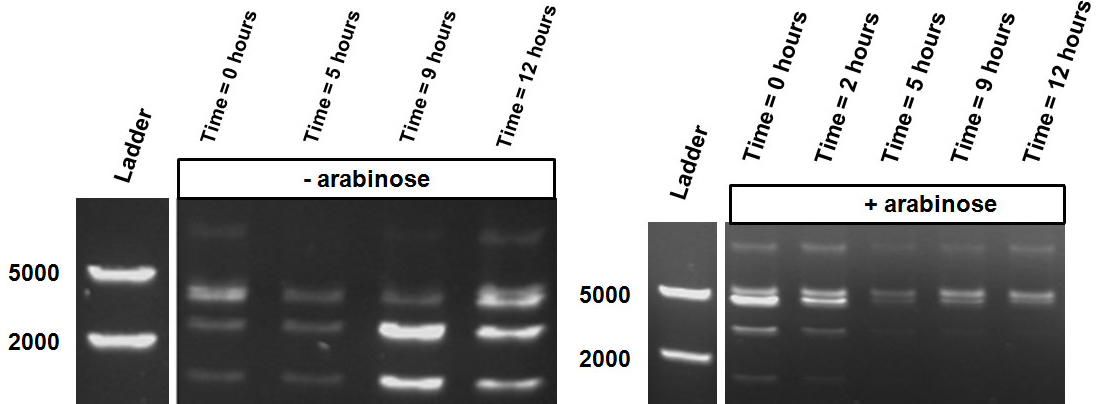
| - | Over the course of time the methylation-dependent restriction profile observed in the gel showed a shift towards the ‘on’ digestion profile in the presence of arabinose. This result shows that our Cellular Logbook is able to sense and write an arabinose signal present in the medium in a shorter period of time.<br\><br\><br\> | + | Over the course of time the <br\> methylation-dependent restriction profile observed in the gel showed a shift towards the ‘on’ digestion profile in the presence of arabinose. This result shows that our Cellular Logbook is able to sense and write an arabinose signal present in the medium in a shorter period of time.<br\><br\><br\><br\><br\> |
<h2>Towards the Cellular Logbook</h2> | <h2>Towards the Cellular Logbook</h2> | ||
[[File:Amsterdam_exp_fig_11.png|300px|right|thumb|Figure 12]] | [[File:Amsterdam_exp_fig_11.png|300px|right|thumb|Figure 12]] | ||
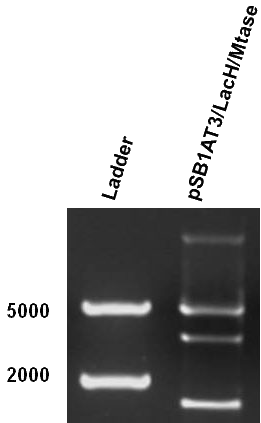
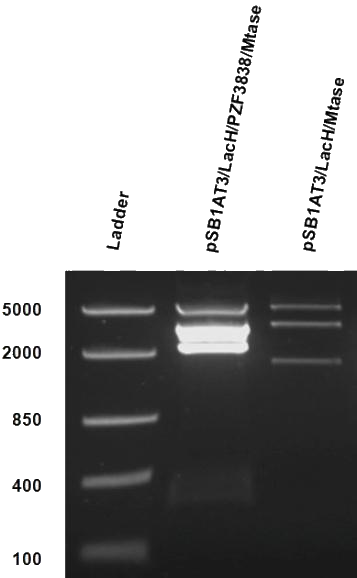
[[File:Amsterdam_exp_fig_12.png|300px|right|thumb|Figure 13]] | [[File:Amsterdam_exp_fig_12.png|300px|right|thumb|Figure 13]] | ||
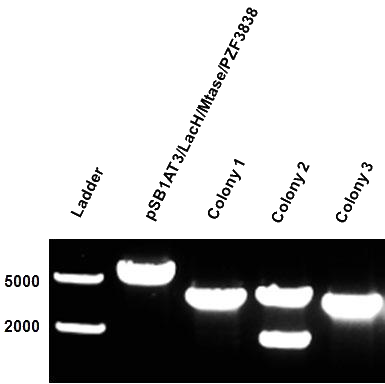
| - | After months of cloning attempts, it appears that we succeeded in obtaining the final version of our Cellular Logbook: the pSB1AT3/LacH/PZF3838/Mtase/Mem2X (BBa_K874300). Digestion of extracted plasmid DNA with the restriction enzyme BamHI, present in the pSB1AT3 backbone vector and in the reader module (BBa_K874040), ensured that both the Polydactyl Zinc Finger PZF3838 (BBa_K874001) and the reader module (BBa_K874040) were successfully cloned in as shown in Figure | + | After months of cloning attempts, it appears that we succeeded in obtaining the final version of our Cellular Logbook: the pSB1AT3/LacH/PZF3838/Mtase/Mem2X (BBa_K874300). Digestion of extracted plasmid DNA with the restriction enzyme BamHI, present in the pSB1AT3 backbone vector and in the reader module (BBa_K874040), ensured that both the Polydactyl Zinc Finger PZF3838 (BBa_K874001) and the reader module (BBa_K874040) were successfully cloned in as shown in Figure 12 (colony 2, 2 bands expected: 3699 and 1769 bp). Preliminary characterization of BBa_K874300 was attempted only once due to time pressure and consisted of a ScaI digestion in the absence of IPTG (figure 13). Compared to the pSB1AT3/LacH/Mtase, BBa_K874300 shows a significant switch to the non-methylated profile, illustrated by the tremendous intensity of the lower bands (cut plasmid) compared to the first one (partially cut plasmid). These results show that the presence of the PZF3838 enhances the specificity of the writer.<br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\><br\> |
<h1>Reference List</h1> | <h1>Reference List</h1> | ||
Revision as of 03:35, 27 September 2012
 "
"