Team:USP-UNESP-Brazil/Associative Memory/Background
From 2012.igem.org
(→Genetic Construction) |
(→Hopfield Associative Memory Networks) |
||
| Line 2: | Line 2: | ||
===Hopfield Associative Memory Networks=== | ===Hopfield Associative Memory Networks=== | ||
| - | The idea of this project is based on the associative memory network introduced by J.J. Hopfield in the 80’s [http://en.wikipedia.org/wiki/Hopfield_network]. The structure of a Hopfield network is simple, all neurons are interconnected, | + | The idea of this project is based on the associative memory network introduced by J.J. Hopfield in the 80’s [http://en.wikipedia.org/wiki/Hopfield_network]. The structure of a Hopfield network is simple, all neurons are interconnected, and that brings about some interesting memory properties and provides a model for understanding human memory. |
| - | We have chosen to built a Hopfield network because of its simplicity and robustness. The same methodology can be used to the construction of networks with different architectures, such as the called “perceptrons” [http://en.wikipedia.org/wiki/Perceptron]. In contrast to a Hopfield network, a perceptron is commonly used as a classifier | + | We have chosen to built a Hopfield network because of its simplicity and robustness. The same methodology can be used to the construction of networks with different architectures, such as the called “perceptrons” [http://en.wikipedia.org/wiki/Perceptron]. In contrast to a Hopfield network, a perceptron is commonly used as a classifier. |
===Biological Mechanism=== | ===Biological Mechanism=== | ||
Revision as of 01:09, 27 September 2012
 Introduction
Introduction Project Overview
Project Overview Plasmid Plug&Play
Plasmid Plug&Play Associative Memory
Associative MemoryNetwork
 Extras
ExtrasHopfield Associative Memory Networks
The idea of this project is based on the associative memory network introduced by J.J. Hopfield in the 80’s [http://en.wikipedia.org/wiki/Hopfield_network]. The structure of a Hopfield network is simple, all neurons are interconnected, and that brings about some interesting memory properties and provides a model for understanding human memory.
We have chosen to built a Hopfield network because of its simplicity and robustness. The same methodology can be used to the construction of networks with different architectures, such as the called “perceptrons” [http://en.wikipedia.org/wiki/Perceptron]. In contrast to a Hopfield network, a perceptron is commonly used as a classifier.
Biological Mechanism
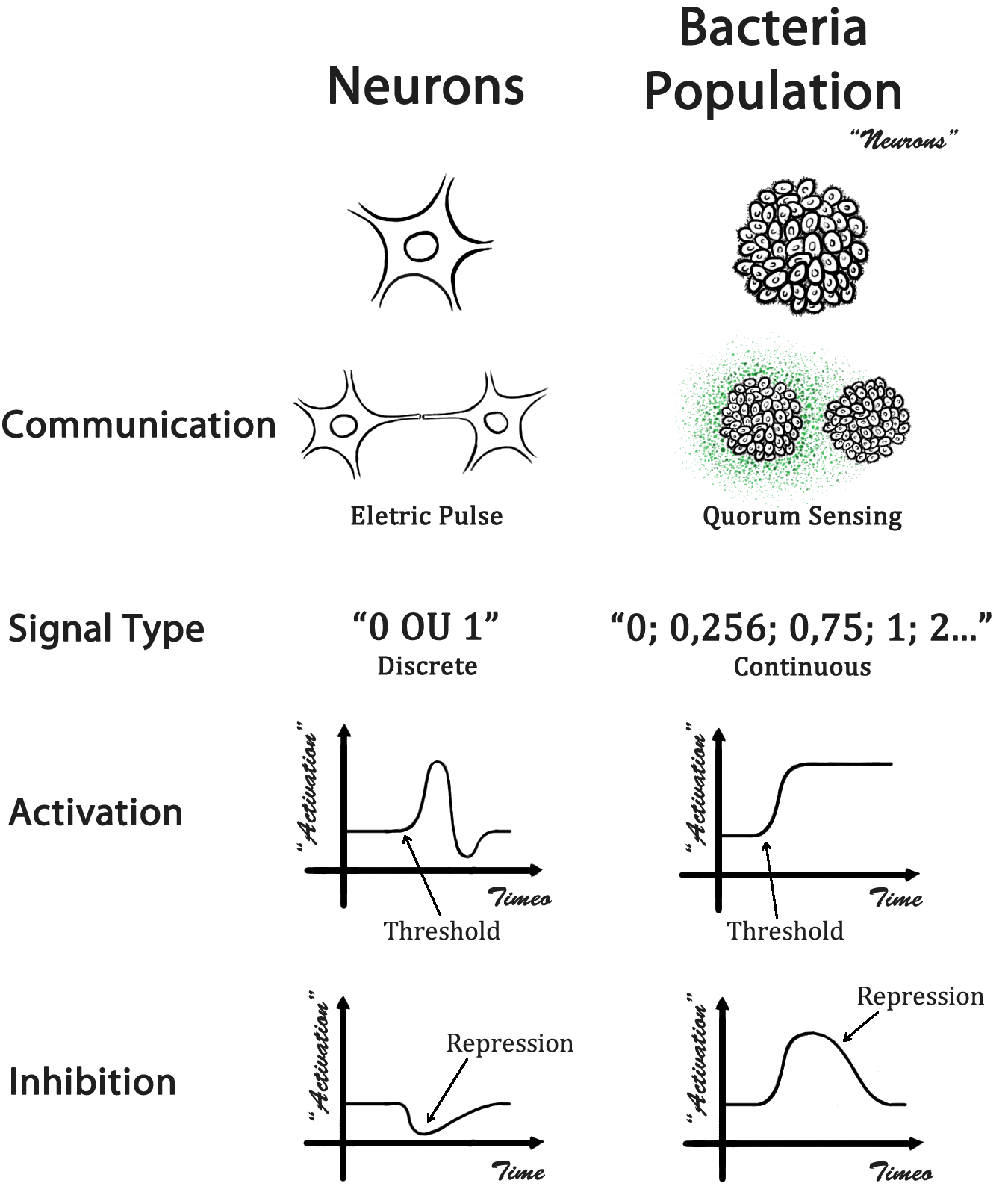
In a biological neural network, the cells occupy a specific location and the information is addressed through a direct physical contact - the neuron axonal projections. In our case, a population of bacteria represents a single neuron and the information is addressed by a quorum sensing molecule (QSM). With different QSM, it is possible to address the information in a specific manner. A comparison between a biological neural network and our design is presented in Fig 1.
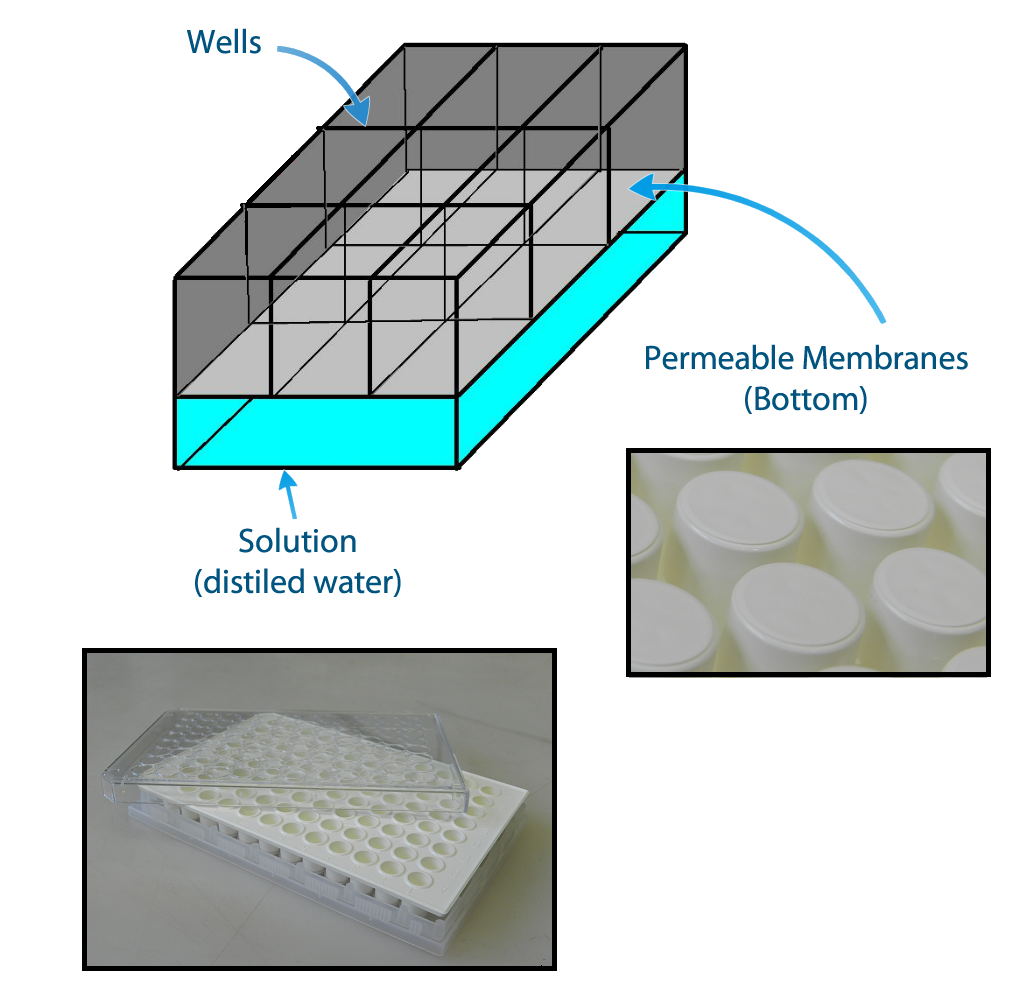
To measure if one population is active, we have planned the construction of a device that keeps each population of bacteria in a fixed position and enable the communication between different populations via QSM, figure 2. The device can be constructed using a plate of 96 wells with membranes attached to the bottom. The membranes allow the diffusion of the quorum sensing substances but prevent the flux of bacterial populations.
Genetic Construction
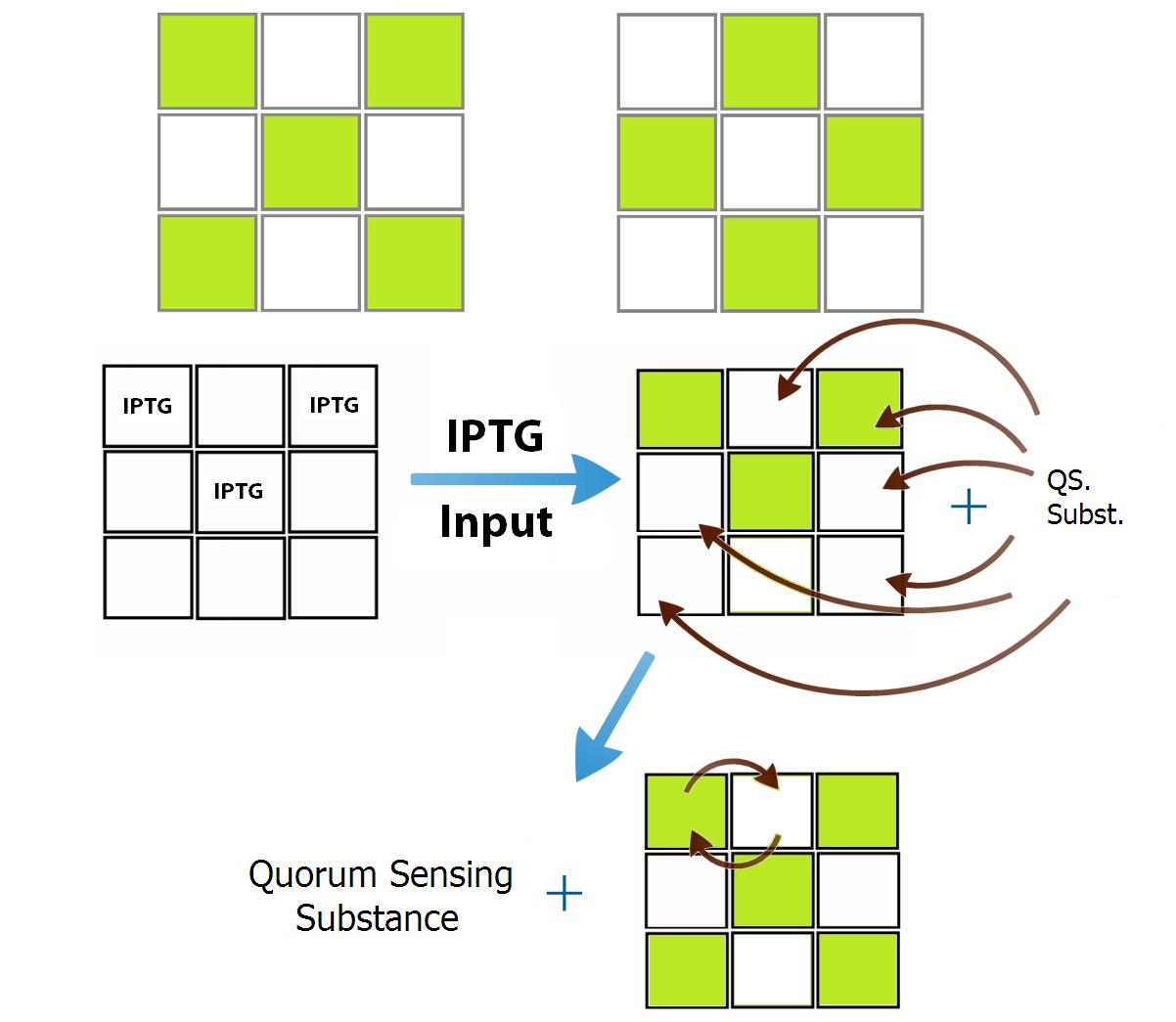
The training of the network is previously defined in silico and then, it is inserted in the bacteria through a genetic construction. Each population of bacteria ("neuron") is defined by his own QSM and, because of that, the number of "neurons" is limitated to the number of QSMs. As a proof of concept we designed two populations of bacteria that comunicate between them in a repressive manner. n order to make your test visually interesting, we used our 3x3 wells device and we trained our network to recognize two antisymmetric pattern, figure 3. Since they are antisymmetric, only two different population of bacteria are needed and we chose to represent the patterns "X" and a "O". In this case each position of the letter “X” inhibits all positions of “O” and activates the positions of its own pattern (and vice-versa).
In the Registry of Parts [http://partsregistry.org/Main_Page] there are 4 quorum sensing systems well characterized. However, there is a strong activation crosstalk between two of them (Las and Rhl). Therefore, we decided to use the system Cin and Rhl.
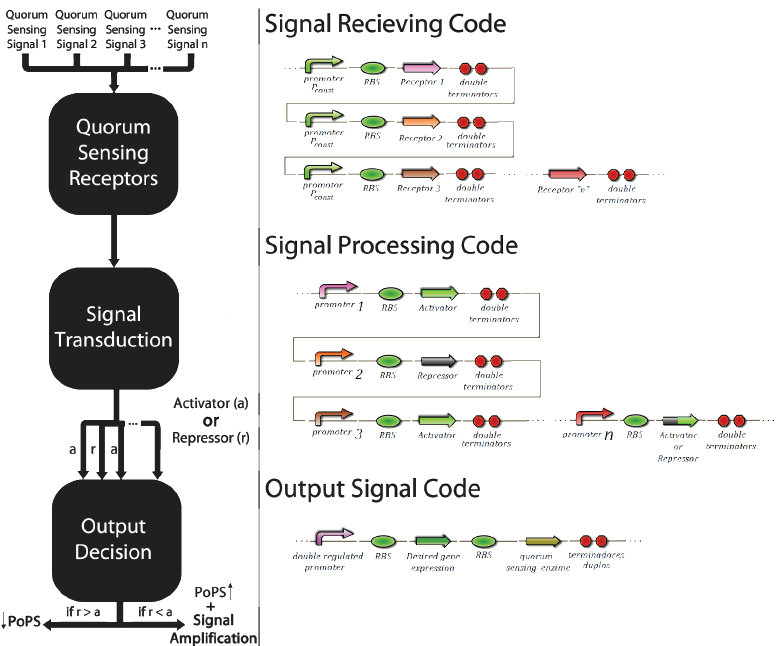
To convert the signals in activation or inhibition, we created a system of transduction of the quorum sensing signal to transcription of an activator or an inhibitor of the transcription of GFP, this will be our system activation reporter. Simultaneous inhibitions and activations of a bacterial population will be converted to a molecular competition of activators and inhibitors by the promoter that controls the production of GFP. It is this molecular competition that promotes the decision between the memories of the communication systems, associating a given input with a more similar memory. As an example, if an input activates more positions of the “X” pattern than the “O”, the competition in the pattern “X” positions will be more favorable to its activation due to the greater number of activators produced by the activated positions, while in the positions of the “O” pattern the opposite occurs, because of its small number of positions activated initially by the given input.
The promoter multi-regulated by an activator and an inhibitor is called Prm. Its inhibitor is the transcriptional factor cl434 and its activator is the cl factor. The genetic design of the positions of the patterns “X” and “O” can be seen in Figure 6.
The construction using the signal transduction containing cl434 and cl allows creating different systems of associative memory, limited only by the quantity of quorum sensingsystems available. Figure 6 shows how this generic system would work and elucidates how this system could be applied to different functions involving a genetic control.
 "
"