Team:TU Munich/Project/Vector Design
From 2012.igem.org
(→Vector Design) |
VolkerMorath (Talk | contribs) (→Results) |
||
| Line 45: | Line 45: | ||
---- | ---- | ||
| - | === BioBricks === | + | === BioBricks backbones === |
| + | |||
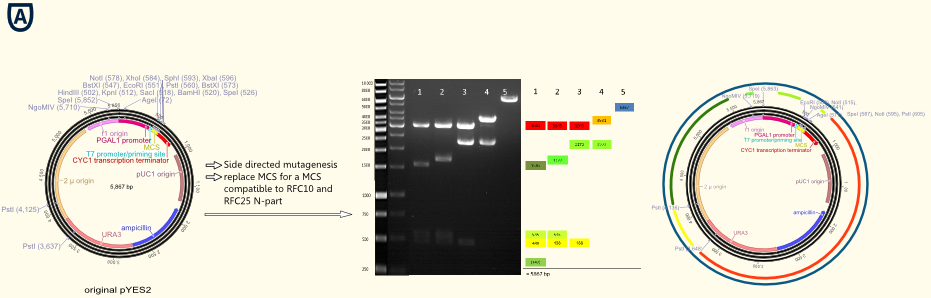
| + | [[file:TUM12_experiment_overwiew_vector.png|500px|thumb|right| Explanations on the figure: | ||
| + | |||
| + | Figure A shows the new multiple cloning site (MCS) containing the RFC 10/25 restriction sides and the DNA sequence coding for the ''Strep''-tag II. | ||
| + | |||
| + | Figure B gives an overview of all important functional elements located on the vector backbone. Upstream to the new MCS lies a T7 promoter primer binding site allowing easy forward sequencing of integrated gene constructs using the standard T7 primer. The URA 3 gene is a prototrophy marker used for the selection of transfected cells. | ||
| + | |||
| + | Figure C to E present the successfully designed BioBricks: pTUM100 simply contains the new MCS, the transcription terminator and further elements required for cloning and transfection. pTUM102 to pTUM104 contain in addition the constitutive promoters pTef1, pTef2 and ADH. On pTUM104 is the galactose inducible promoter pGAL1 located. ]] | ||
| + | |||
| + | |||
==== Gel pictures of finished constructs ==== | ==== Gel pictures of finished constructs ==== | ||
Revision as of 00:39, 27 September 2012



Contents |
Vector Design
What is the use of DNA sequences coding for valuable enzymes without the possibility to express them and analyze the protein activity?
As we planned to detect the enzymes from our biosynthetic pathways via Strep-tag II, a RFC25 compatible backbone was necessary. As no such backbone was availabe for yeast in the PartsRegistry, an important task at the beginning of our project was the design of an expression vector for yeast which is compatible to the iGEM cloning principles and standards. Based on the commercially available pYES2 vector, manufactured by Invitrogen, we created an vector containing:
- a RFC 25 compatible multiple cloning site
- a strep-tag II
- a 2µ origin for high copy number replication in Saccharomyces cerevisae
- the URA3 gene to use the uracil prototrophy of S. cerevisiae INVSc' as a selection marker for transfection
- a pUC origin for high copy number replication in E.coli
- the ß-lactamase coding gene to use ampicillin as a selection marker for cloning applications in E.coli
- a galacotsoe inducible PGal 1 promoter
- a CYC1 transcription terminator
Furthermore we designed several vectors containing our constitutive promoters Tef1, Tef2 and ADH and different additional terminators. The different versions of the vector have been successfully applied and tested in all other subprojects.
Design
vom Poster:
RFC25 COMPATIBILITY
To be able to test and quantify the expression of desired enzymes in yeast we designed an expression vector which is compatible to the iGEM RFC25 standard based on the commercially available pYES2 vector from Invitrogen.
DESIGN
This included an exchange of the original multiple cloning site by a multiple cloning site consisting of the restriction enzymes used in the iGEM RFC25 standard. Additionally the restriction sites of these enzymes in the vector backbone have been eliminated by quick-change mutagenisis. Furthermore we inserted a gene sequence coding for a strep-tag II straight behind the multiple cloning site in order to facilitate the extraction of expressed proteins. The successful introduction of these modifications has been confirmed via DNA sequencing. Additionally a second vector containing a His-tag instead of the strep-tag II and a vector giving the opportunity to test different promoters is currently planned.
Description of the original vector + 4 different modified versions.
Results
BioBricks backbones

 "
"
