Team:Colombia/Project/Experiments/Pseudomonas
From 2012.igem.org
m (→Escherichia coli K12) |
m (→Escherichia coli K12) |
||
| Line 3: | Line 3: | ||
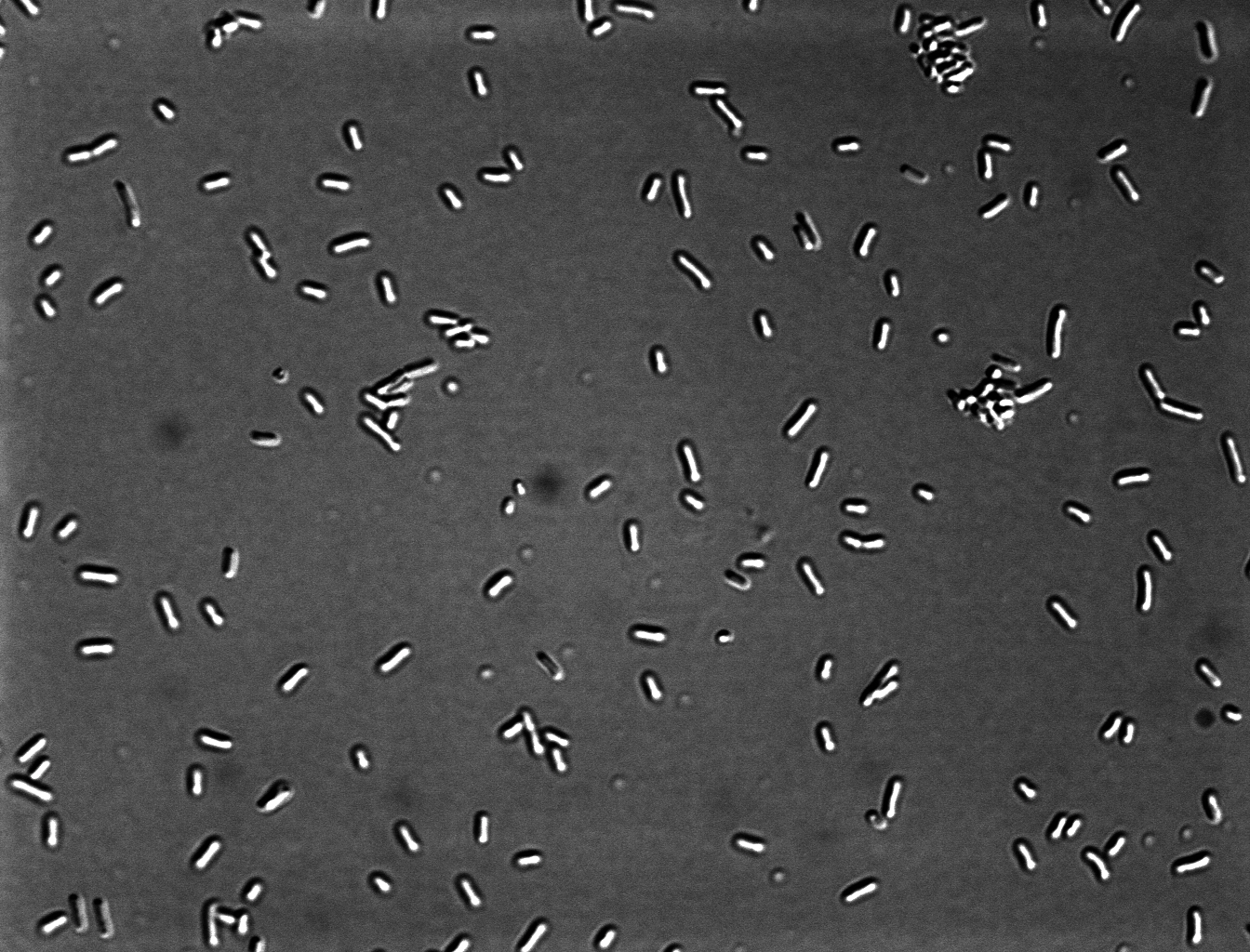
=''' ''Escherichia coli'' K12 '''= | =''' ''Escherichia coli'' K12 '''= | ||
| - | [[File:DICx60_E_coli.jpg|center| | + | [[File:DICx60_E_coli.jpg|center|300 px]] |
= ''' Toxin-Antitoxin ''' = | = ''' Toxin-Antitoxin ''' = | ||
Revision as of 15:38, 26 September 2012
Template:Https://2012.igem.org/User:Tabima
Escherichia coli K12
Toxin-Antitoxin
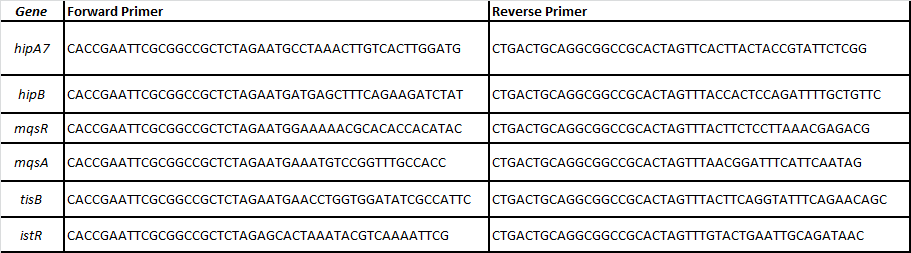
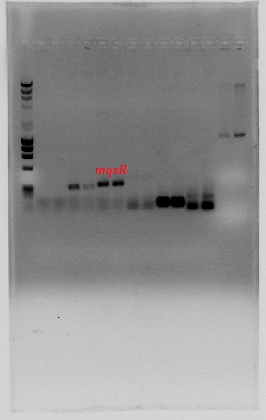
We designed specific primers for each ORF of the following toxin-antitoxin pairs related with persistence (Balaban et al, 2004; Lewis 2010; Maisonneuve et al, 2011):
-HipA7/HipB
-MqsR/MqsA
-TisB/istR
In order to enable other teams to use said toxin-antitoxin modules we cloned these parts without promoters in chloramphenicol backbones. So far we have constructed the BioBricks with the toxins [http://partsregistry.org/Part:BBa_K831000:Design HipA7] and [http://partsregistry.org/Part:BBa_K831003:Design TisB], and with the antitoxins [http://partsregistry.org/Part:BBa_K831001:Design HipB], [http://partsregistry.org/Part:BBa_K831002:Design MqsA] and [http://partsregistry.org/Part:BBa_K831004:Design istR ].
We were going to use the wild type promoters of the Toxin-Antitoxin modules; however the deterministic model gave us feedback which established that inducible promoters were much more likely to give us the results we were looking for.
For the construction of our inducible parts, we first tried to build the toxins under the control of the Lac promoter and the antitoxins under the tetR promoter. We succeeded in the construction of the parts containing the [http://partsregistry.org/Part:BBa_K831009 plac-hipB] and [http://partsregistry.org/Part:BBa_K831014:Design plac-istR] antitoxins and the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K831007 ptetR-mqsR] inducible toxin.
But due to the leakage of lac promoter and even that of the tetR promoter (has less leakage that lac) we were unable to construct the remaining toxin parts using both of those promoters. To overcome this inconvenience we decided to use the prm promoter which is inducible by the CI lambda phage protein. So far we have constructed the inducible [http://partsregistry.org/Part:BBa_K831015 prm-hipA7] BioBrick.
In order to assess the functionality of these parts, we cloned hipB under the control of lac promoter in an Escherichia coli K12 strain that contains hipA7 in its chromosome (E. coli K12 MG1655 TH1269).
As HipB neutralizes the induction of persistency caused by HipA7 in this strain, we measured the persisters frequency after induction with different concentrations of IPTG and compared it with the normal levels of persistency in this strain.
We are in the process of making inducible CI under the control of the lac promoter in order to analyze the effect of the induction of HipA7 in the persisters frequency on a wild type E. coli K12 strain.
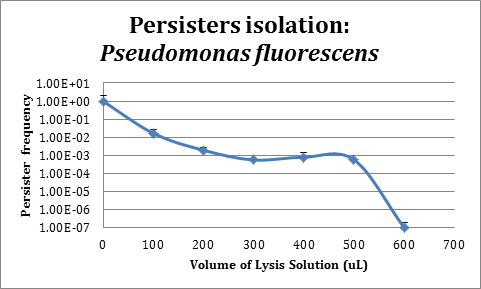
Assays in Pseudomonas fluorescens
In order to improve the stability and survival of our system we are assessing the possibility of using P. fluorescens as our final chassis. For this purpose, we first standardized the normal recount of persisters of a P. fluorescens wild type strain without induction. We used an original protocol for determining persistence based on lysis from work that is not part of iGEM (S. Cañas et al., manuscript in preparation), which we validated for use in P. fluorescens.
 "
"