Team:LMU-Munich/Data/Anderson
From 2012.igem.org
| Line 44: | Line 44: | ||
{| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
| - | <font color="#000000"; size="2"><p align="justify"> '''Fig. 2:''' β-galactosidase assay (up) and growth curve (down) of the Anderson promoters J23100, J23102, J23103, J23106 in ''B. subtilis''. Results derive from one experiment and are only shown for one Bacillus clone. </p></font> | + | <font color="#000000"; size="2"><p align="justify"> '''Fig. 2:''' β-galactosidase assay (up) and growth curve (down) of the Anderson promoters J23100, J23102, J23103, J23106 in ''B. subtilis''. Results derive from one experiment and are only shown for one ''Bacillus'' clone. </p></font> |
|} | |} | ||
|} | |} | ||
|} | |} | ||
<br> | <br> | ||
| - | + | Verification of the promoter activity by β-galactosidase assays revealed that the Anderson promoters do not seem to be as weak as measured by luminescence (Fig. 1). For direct comparison we should measure a constitutive promoter, e.g. P<sub>''liaG''</sub>, in the same experiment. But we think the luminescence measurements are more reliable because they were repeated three times with two independent clones. | |
<br> | <br> | ||
<br> | <br> | ||
Revision as of 08:34, 26 September 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Anderson Promoters
Luminescence measurements
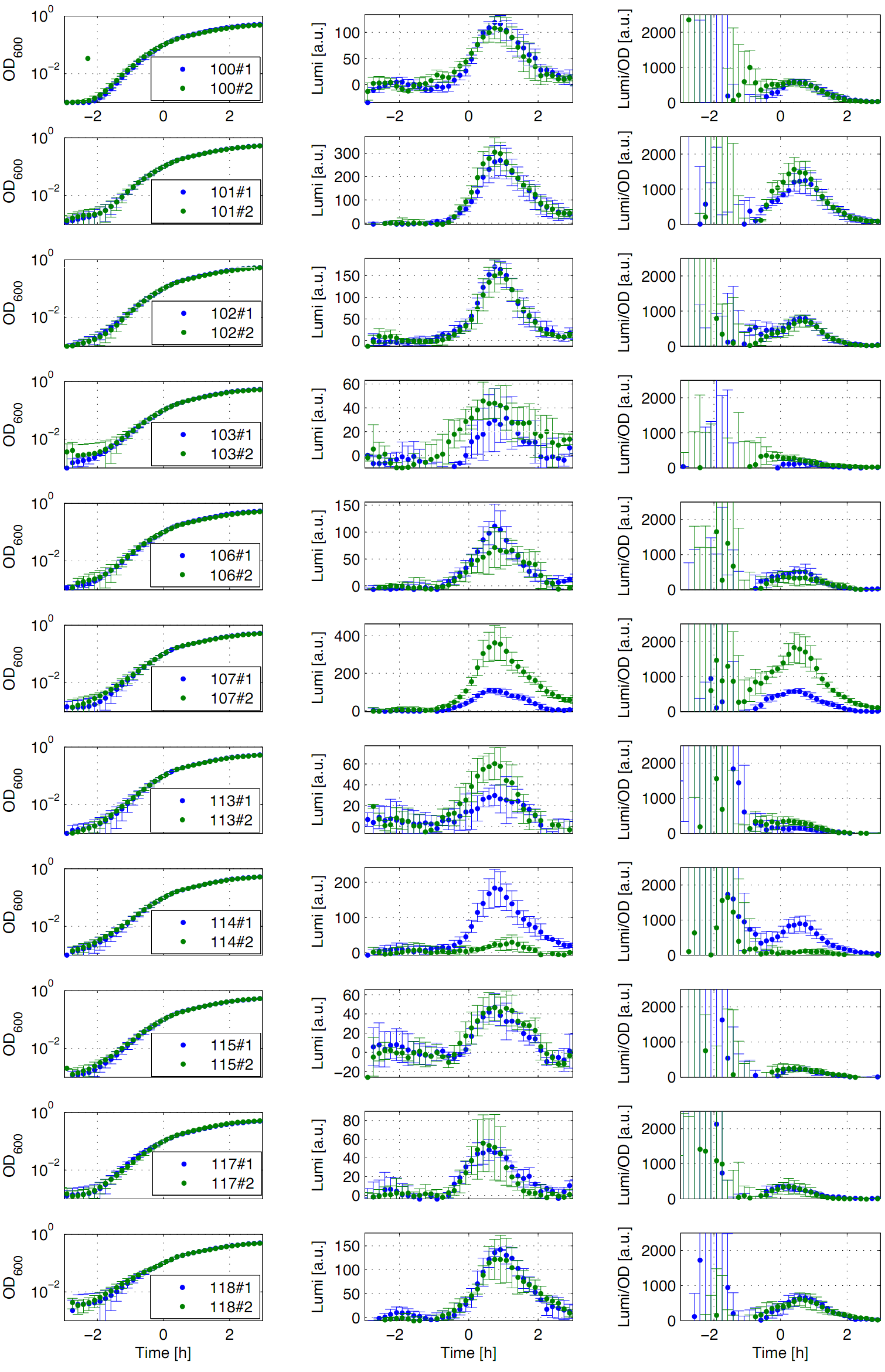
Eleven of the nineteen promoters of the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] (J23100, J23101, J23102, J23103, J23106, J23107, J23113, J23114, J23115, J23117, J23118) were evaluated. Therefore, we used the reporter vector pSBBs3C-luxABCDE from the BioBrickBox containing the lux operon ![]() as a reporter for promoter activity. The promoter activity leads to the expression of the lux operon and to the production of the enzyme luciferase. The luminescence, which is produced by the luciferase, can be measured with the plate reader Synergy2 (BioTek) (Fig.1).
as a reporter for promoter activity. The promoter activity leads to the expression of the lux operon and to the production of the enzyme luciferase. The luminescence, which is produced by the luciferase, can be measured with the plate reader Synergy2 (BioTek) (Fig.1).
All clones show a usual growth behaviour. The activity of the promoters increases during transition from log to stationary phase. The maximum (t=1h) reaches from 200Lumi/OD600 (promoter J23115) to a maximum of 1500 Lumi/OD600 for the strongest promoter (J23101). Afterwards the activity goes down to the beginning level (t=2h). The oscillation of luminescence (Lumi/OD600) in the beginning of the curves are due to the small OD600 values and do not mean high promoter activity. One clone of J23107 and J23114 shows significantly lower promoter activity. Therefore, additional clones should be measured. In comparison to all the other evaluated Bacillus promoters, these Anderson promoters showed a very low acitivity in B. subtilis.
β-galactosidase assay
To evaluate the activity not only with the lux reporter operon, four promoters of the Anderson collection were cloned into the reporter vector pSBBs1C-lacZ ![]() to do β-galactosidase assays and then to compare the results of the strength of these promoters in B. subtilis (Fig. 2). The results were compared to the results from the luminescence measurements.
to do β-galactosidase assays and then to compare the results of the strength of these promoters in B. subtilis (Fig. 2). The results were compared to the results from the luminescence measurements.
|
Verification of the promoter activity by β-galactosidase assays revealed that the Anderson promoters do not seem to be as weak as measured by luminescence (Fig. 1). For direct comparison we should measure a constitutive promoter, e.g. PliaG, in the same experiment. But we think the luminescence measurements are more reliable because they were repeated three times with two independent clones.
 "
"