Team:Amsterdam/modeling/generaldesign
From 2012.igem.org
(Difference between revisions)
(→Model definition) |
(→In theory) |
||
| Line 139: | Line 139: | ||
== In theory == | == In theory == | ||
| + | <center> | ||
| + | [[File:infer.gif|image]] <br> | ||
| + | <b>Figure 3</b>. Animation of <math>P_{1}(t)</math> over time with <math>s_{\text{off}}</math> at <math>t = 0</math>. From the measured value of <math>P_{t}</math>, <math>t</math> of measurement can be solved for as is shown using the dashed lines. The value of <math>P_{1}(t)</math> will decrease over time until it becomes 0. An interactive version of this plot has been included in the attached Mathematica file. | ||
| + | </center> | ||
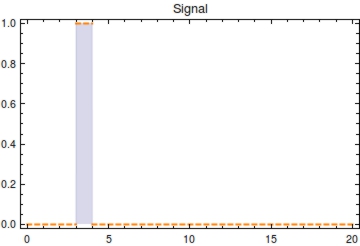
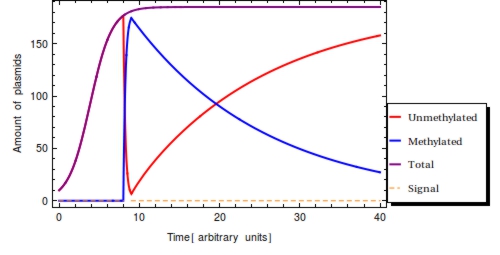
| - | + | The monotonically decreasing value of <math>F(t) = \frac{\text{methylated plasmids}}{\text{total plasmids}}</math> can be used to infer <math>s_{\text{off}}</math>, given that the degradation rate (<math>\alpha</math>) and capacity constraint <math>Ca</math> are known and constant. Also assumed is that all bits are methylated during signal presence, this implicates <math>\omega</math> is sufficient to methylate all bits during presence of the signal. Irrespective of the initial amount of plasmids, the population of plasmids within the single cell will have reached a steady state value of <math>\frac{\beta}{\alpha}</math>. As we see in the Figure 2, <math>F(t)</math> will start to decrease as a function of the degradation rate after the signal has left the medium following the following function: | |
| - | + | ||
| - | The monotonically decreasing value of <math>F(t) = \frac{\text{methylated plasmids}}{\text{total plasmids}}</math> can be used to infer, given that the degradation rate (<math>\alpha</math>) and capacity constraint <math>Ca</math> are known and constant | + | |
<math>\frac{dP_{1}}{dt} = - \alpha\ P_{1}</math> | <math>\frac{dP_{1}}{dt} = - \alpha\ P_{1}</math> | ||
Revision as of 07:05, 21 September 2012
 "
"