Team:TU Munich/Notebook/Labjournal
From 2012.igem.org
(→Tuesday, August 15th) |
(→Tuesday, August 15th) |
||
| Line 11,788: | Line 11,788: | ||
</div> | </div> | ||
| + | <div class="light_switchable_promoter"> | ||
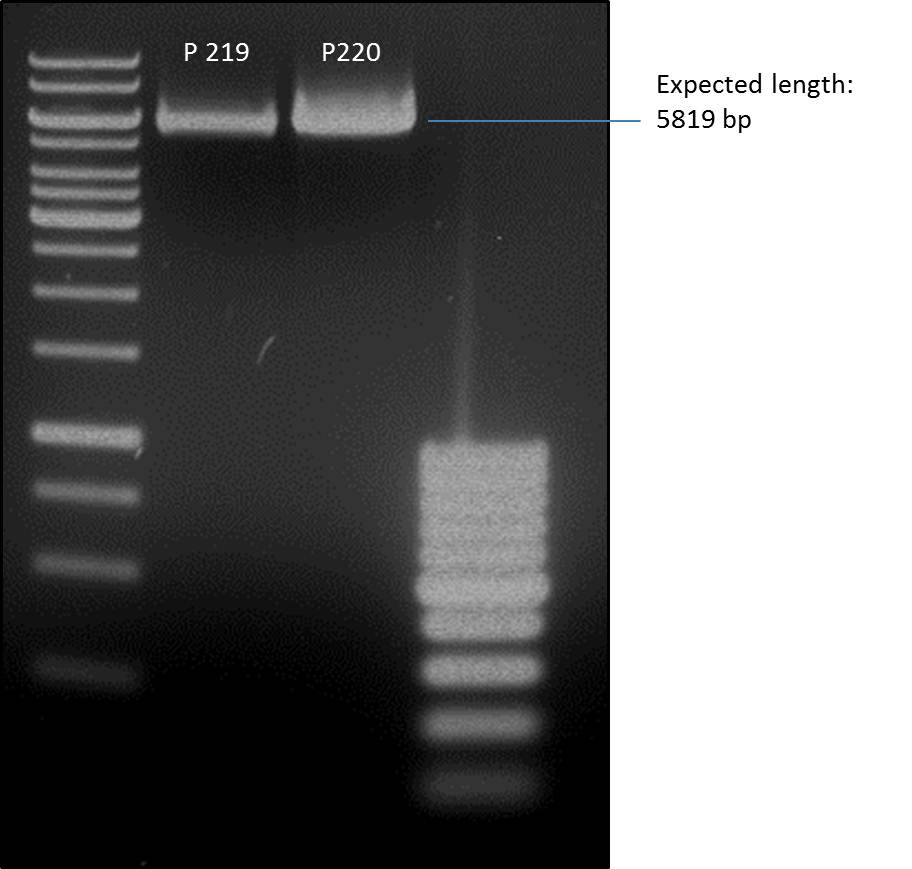
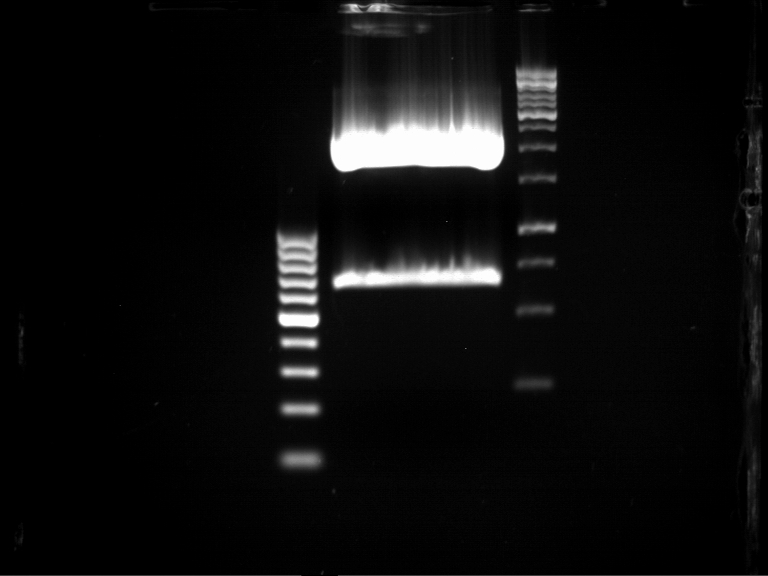
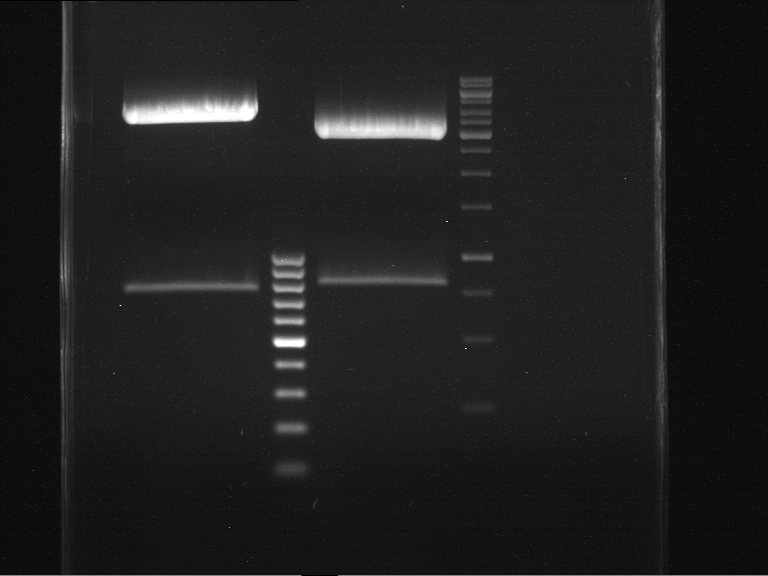
=== Analytical digestion and gelelectrophoresis of the minipreps of transformated ''E. coli'' XL1-Blue with ligated P71+P322 and P71+P323 products === | === Analytical digestion and gelelectrophoresis of the minipreps of transformated ''E. coli'' XL1-Blue with ligated P71+P322 and P71+P323 products === | ||
Revision as of 19:09, 15 August 2012



Labjournal
June
Wednesday, June 13th
Exchange of the Multiple Cloning Site of pYES2
Investigator: Saskia, Daniela
Aim of the experiment: Exchange of the Multiple Cloning Site of pYES2
Hybridisation of the primers O1 with O2 and O3 with O4
Primer preparation:
- centrifugation
- dilution in the denoted quantity in bidest. water (concentration = 100 pmol/µl)
- centrifugation
Hybridisation:
| volume | reagent |
| 34 µl | ddH2O |
| 5 µl | PNK-buffer |
| 2.5 µl | Primer O1 |
| 2.5 µl | Primer O2 |
| 1 µl | PNK (10mM) |
| volume | reagent |
| 34 µl | ddH2O |
| 5 µl | PNK-buffer |
| 2.5 µl | Primer O3 |
| 2.5 µl | Primer O4 |
| 1 µl | PNK (10mM) |
- 30 min 37 °C
- 10 min 80 °C
- put the Thermo Block with the mixture in a styrofoam box and let it cool down over night
Thursday, June 14th
Exchange of the Multiple Cloning Site of pYES2
Investigator: Saskia, Daniela
Aim of the experiment: Exchange of the Multiple Cloning Site of pYES2
Digestion of pYES2 with HindIII and XbaI
| volume | reagent |
| 12 µl | Miniprep (pYES2 SH 1.7.3 with a concentration of 87.8 ng/µl) |
| 5 µl | NEB2 |
| 5 µl | 10x BSA |
| 2 µl | XbaI (10 U/µl) |
| 2 µl | HinIII (10U/µl) |
| 24 µl | ddH2O |
Incubation: 37 °C, 1.75 h
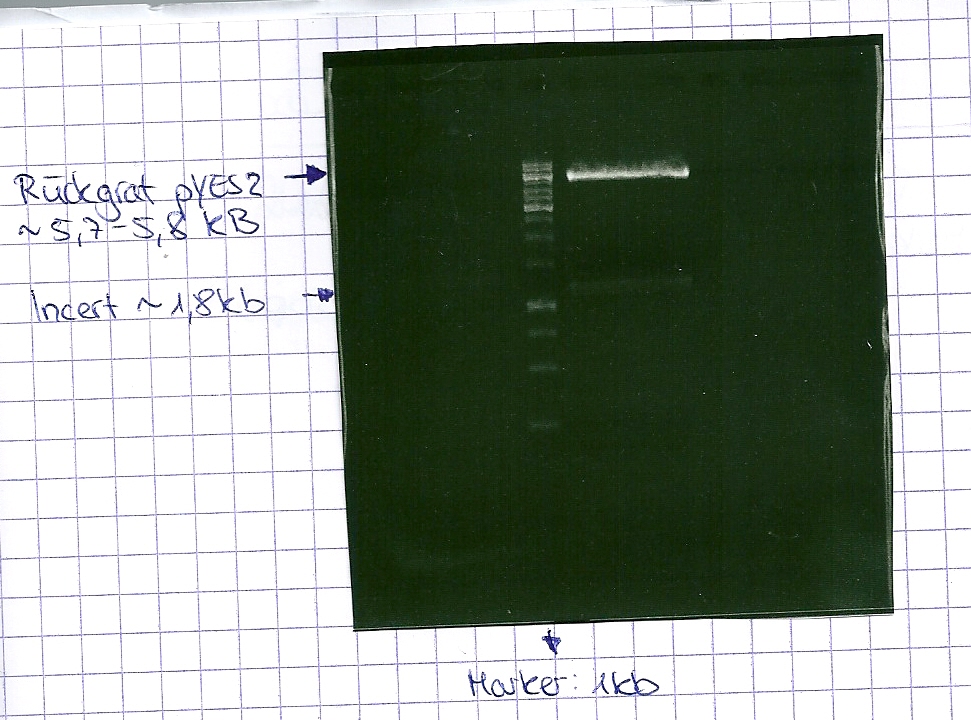
DNA preparative gel electrophoresis
- gel: 1% with LMP-agarose
- band 1: 10 µl DNA ladder (1kb)
- band 2: 50 µl probe + 5 µl loading dye
- 70 V, 90 min
Gelextration
- cut the bands (5.7-5.8 kb) and split it in two eppis
- m1=165.3 mg
- m2=211.1 mg
- QIAquick Gel Extractrion Kit
- eppi1: 495.9 µl QG-buffer
- eppi2: 633.3 µl QG-buffer
- step 6 was left out
- step 9: 30µl buffer, 4 min incubation
- the product was named P5
Transformation of plasmids from Prof. Schwab in E.coli XL-1 Blue
Investigator: Lara, Andrea
Aim of the experiment: Preparation of the plasmids for transformation
Overnight cultures of cells with limonenesynthase-plasmid from Prof. Schwab
- resuspend 50 µl / 200 µl of competent cells with 50 ml LB medium
- add 0,1 ml Ampicillin (100 µg/ml) and 0,28 ml Chloramphenicol (170 µ/ml) for strain 108
- add 0,07 ml Kanamycin (50 µg/ml) and 0,28 ml Chloramphenicol (170 µ/ml) for strain 106
- incubate at 37 °C
Friday, June 15th
Exchange of the Multiple Cloning Site of pYES2
Investigator: Saskia, Daniela
Aim of the experiment: Exchange of the Multiple Cloning Site of pYES2
Analytical DNA gel electrophoresis
- gel: 1.2 %
- band 1: 10 µl DNA ladder (1kb)
- band 2: 3 µl pYES2 digested + 7 µl TAE-buffer + 1 µl loading dye
- band 3: 3 µl O5 + 7 µl TAE-buffer + 1 µl loading dye
- band 4: 3 µl O6 + 7 µl TAE-buffer + 1 µl loading dye
- band 5: 10 µl DNA ladder (100 bp)
Ligation of Plasmid P5 (pYES2 digested) with the hybridized primers O5 and O6
| volume | reagent |
| 4 µl | pYES2 digested (P5) |
| 1 µl | O5 |
| 1 µl | O6 |
| 2 µl | T4-ligase buffer (10x) |
| 0,5 µl | T4 DNA-ligase |
| 11.5 µl | ddH2O |
Negative control
| volume | reagent |
| 4 µl | pYES2 digested (P5) |
| 2 µl | T4-ligase buffer (10x) |
| 0,5 µl | T4 DNA-ligase |
| 13.5 µl | ddH2O |
- water bath 16 °C
- after 3 h a probe for the transformation was taken
- the rest was ligated over the weekend
Transformation of E. coli with ligated products (P6)
- competent cells: SHXL1 Blue (by Simon)
- Transformation with ligation product (P6) and negative control
results:
- P6 (100 µl): 1 clone
- P6 (concentrated): 30 clones
- negative control (100 µl): 0 clones
- negative control (concentrated): 6 clones
Transformation of plasmids from Prof. Schwab into E.coli XL-1 Blue
Investigator: Andrea
Aim of the experiment: Preparation of the plasmids for transformation
Determination of the concentration with Nano Drop
| Sample | concentration [ng/µl] |
| P3 | 1353 |
| P4 | no result |
- the strain 106 culture was not grown satisfying and were incubated for 2 more days
Miniprep of pGex-4T-1 of strain 108 from Prof. Schwab
- see QIAprep Spin Miniprep Kit
- stored as P3 (-20 °C)
Sunday, June 17th
Exchange of the Multiple Cloning Site of pYES2
Investigator: Saskia, Daniela
Aim of the experiment: Exchange of the Multiple Cloning Site of pYES2
Picking clones for Miniprep
- 10 clones of transformed E.coli with P6 were picked
- medium: 5ml LB with Amp
Monday, June 18th
Exchange of the Multiple Cloning Site of pYES2
Investigator: Saskia, Daniela
Aim of the experiment: Exchange of the Multiple Cloning Site of pYES2
Miniprep of transformed E.coli with P6
- QIAprepS Spin Miniprep Kit
- step 3: invert 2-3 times (don't shake to avoid destruction of genomic DNA)
- the 10 Minipreps were named: P7 - P16
Determination of the concentration with Nano Drop
| Sample | concentration [ng/µl] | 260/280 |
| P7 | 157.6 | 2.32 |
| P8 | 207.3 | 1.63 |
| P9 | 171.2 | 2.02 |
| P10 | 183.1 | 1.57 |
| P11 | 160.4 | 2.2 |
| P12 | 179.9 | 1.75 |
| P13 | 179.2 | 2.07 |
| P14 | 188.3 | 1.6 |
| P15 | 166.7 | 2.05 |
| P16 | 174.6 | 2.08 |
Controll digestion with HindIII XbaI and NgoMIV
- Samples P7-P16: 2.5 µl
- Negative controll pYES SH 1.7.3: 2.5 µl
- Master Mix HindIII and XbaI: 17,5 µl for a 20 µl preparation
| volume | reagent |
| 3 µl | HindIII |
| 3 µl | XbaI |
| 24 µl | NEB2 |
| 24 µl | 10x BSA |
| 156 µl | ddH2O |
- Master Mix NgoMIV: 17,5 µl for a 20 µl preparation
| volume | reagent |
| 6 µl | NgoMIV |
| 24 µl | NEB4 |
| 180 µl | ddH2O |
Incubation: 37 °C, 1.5 h
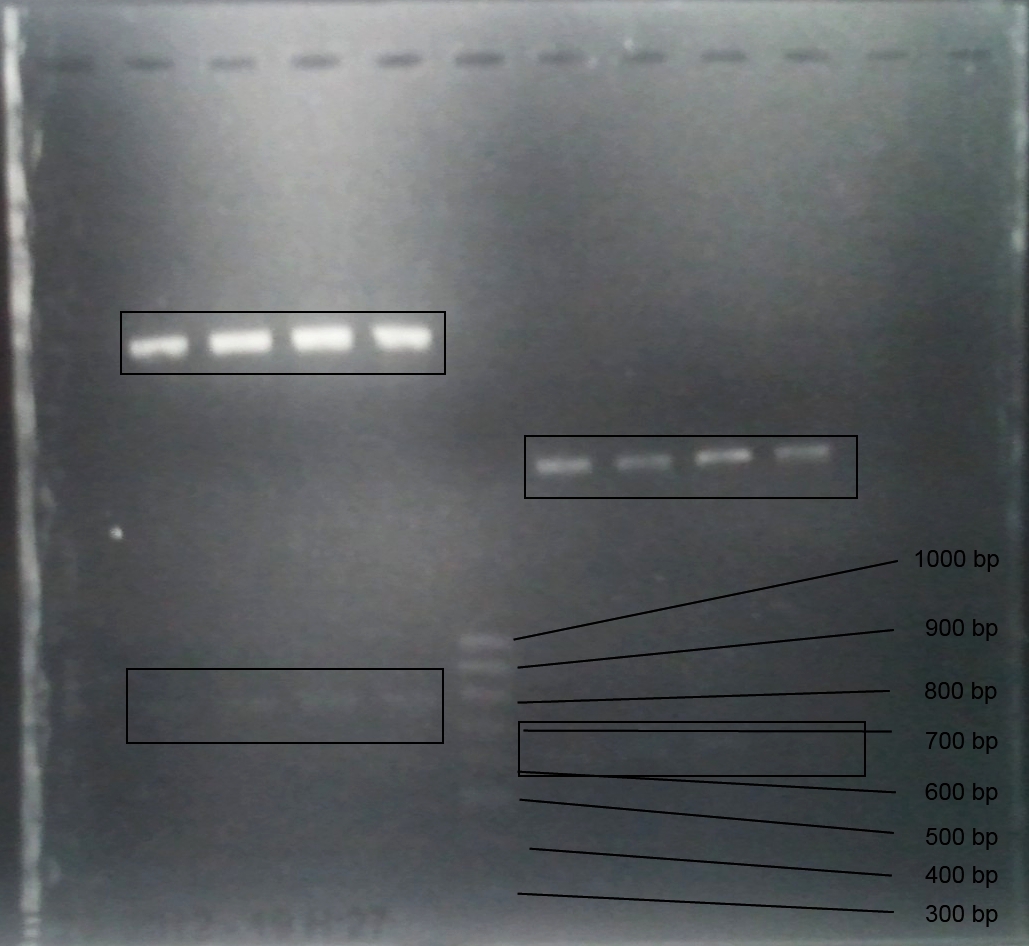
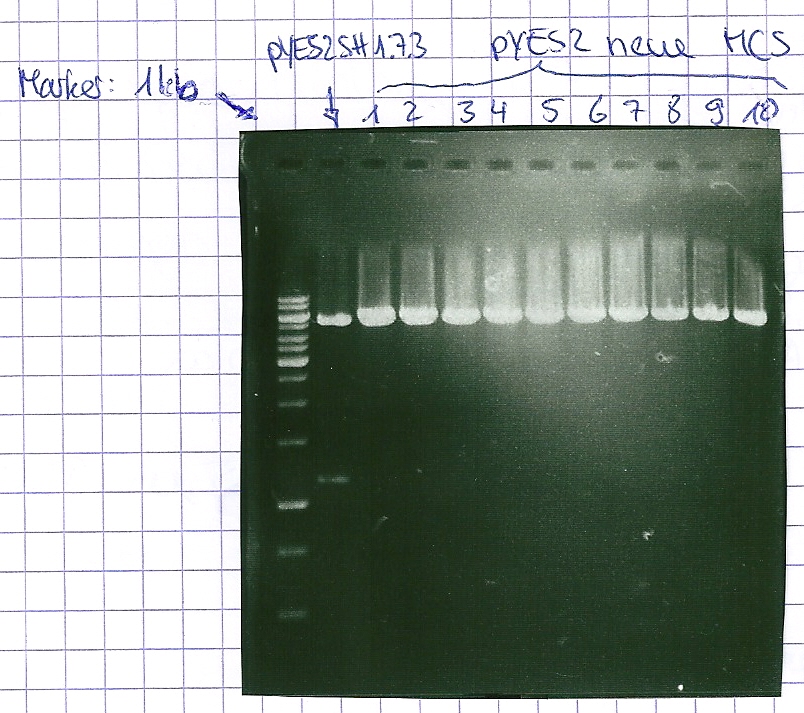
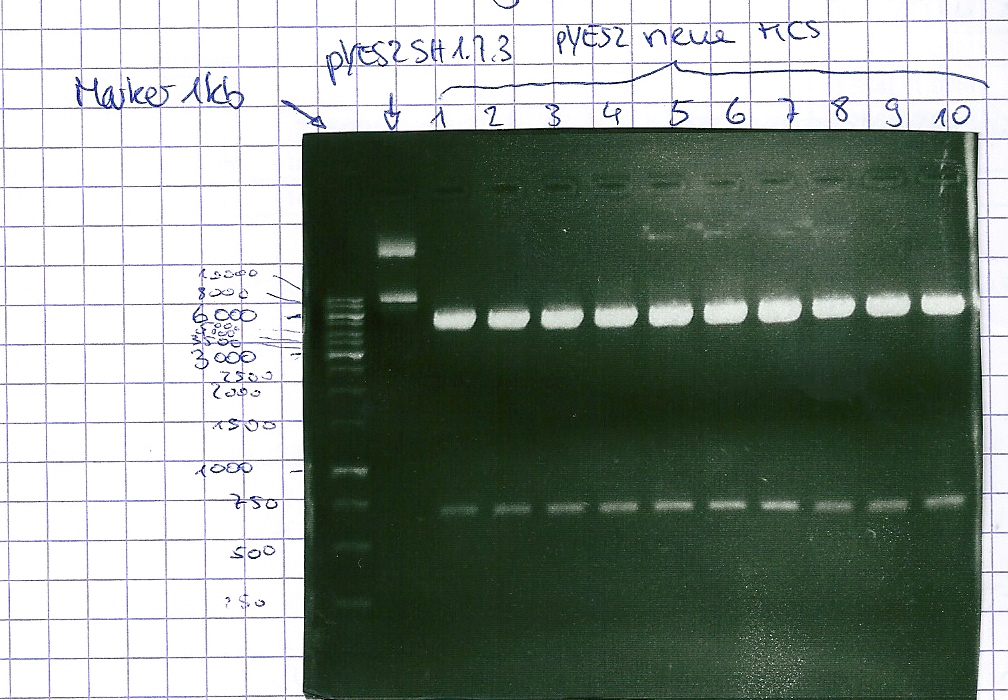
Analytical gel electrophoresis of P7-P16
- gel: 1.5 %
gel 1:
- band 1: 10 µl DNA ladder (1 kb)
- band 2: 3 µl pYES2 SH 1.7.3 digested with HindIII and XbaI + 7 µl TAE buffer + 1 µl loading dye
- band 3 - 12: 3 µl P7-P16 digested with HindIII and XbaI + 7 µl TAE buffer + 1 µl loading dye
gel 2:
- band 1: 10 µl DNA ladder (1 kb)
- band 2: 3 µl pYES2 SH 1.7.3 digested with NgoMIV + 7 µl TAE buffer + 1 µl loading dye
- band 3 - 12: 3 µl P7-P16 digested with NgoMIV + 7 µl TAE buffer + 1 µl loading dye
Tuesday, June 19th
Transformation of BBa_I742111 (Limonenesynthase) into E.coli XL-1 Blue
Investigator: Andrea
Aim of the experiment: Transformation
- for each Biobrick 100 µl cells were used and pooled together with 2 µl of plasmid DNA
- Incubation on ice for 30 min
- 5 min heat shock at 37 °C
- cells were prefilled with 1 ml of LB-medium and incubated in a cell-culture shaker at 37 °C for 45 min
- 100 µl of these cell suspension were plated on antibiotic selection plates (Ampicillin)
- cell suspension was centrifuged at 13000 rpm for 60 s for resuspending the pellet with 100 µl LB and plating also
- incubation at 37 °C overnight
Wednesday, June 20th
Exchange of the Multiple Cloning Site of pYES2
Investigator: Saskia, Daniela
Aim of the experiment: Exchange of the Multiple Cloning Site of pYES2
Sequencing of P13 and P14: pYES2 with new MCS sequencing primer:
- 1.6 µM forward primer O9
- DNA P13 and P14
The Multiple Cloning Site was exchanged successfully!!!
Transformation of BBa_I742111 (Limonenesynthase) into E.coli XL-1 Blue
Investigator: Daniela
Aim of the experiment: Transformation
Picking of Clones
- 6 clones were picked
- Incubation at 37 °C in LB + Amp
Thursday, June 21st
Transformation of BBa_I742111 and plasmids from Prof. Schwab into E.coli XL-1 Blue
Investigator: Lara, Andrea
Aim of the experiment: Controll of Transformation
Controll digestion
- Sample P3
| volume | reagent |
| 14 µl | Plasmid-DNA |
| 0,25 µl | NcoI |
| 2 µl | Buffer Tango (Fermentas) |
| 0,25 µl | HindIII |
| 2 µl | Buffer Red (Fermentas) |
| 1,5 µl | ddH2O |
- Sample P4
| volume | reagent |
| 6 µl | Plasmid-DNA |
| 0,25 µl | EcoRI |
| 2 µl | Buffer EcoRI (Fermentas) |
| 0,25 µl | NotI |
| 2 µl | Buffer Orange (Fermentas) |
| 9,5 µl | ddH2O |
- Sample Biobrick-clones
| volume | reagent |
| 5 µl | Plasmid-DNA |
| 0,25 µl | EcoRI |
| 2 µl | Buffer EcoRI (Fermentas) |
| 0,25 µl | PstI |
| 2 µl | Buffer Orange (Fermentas) |
| 10,5 µl | ddH2O |
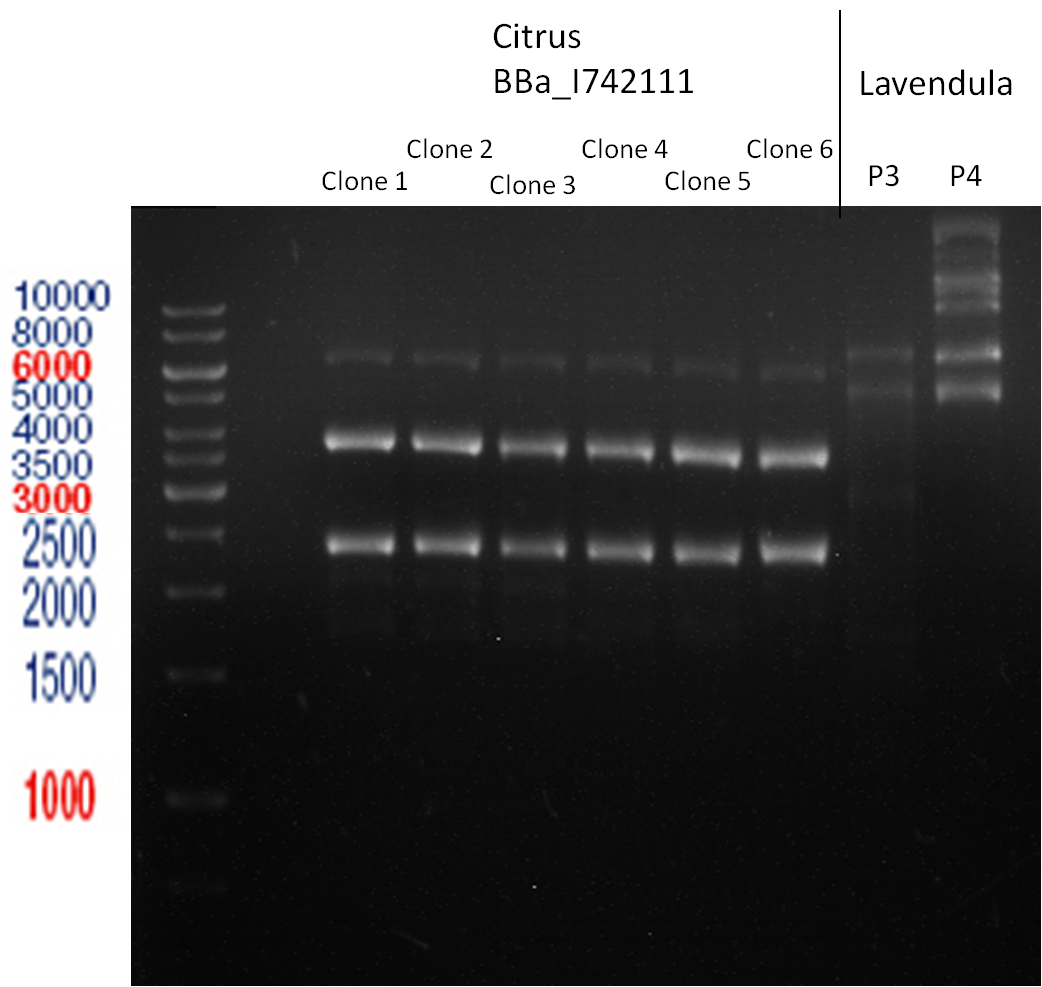
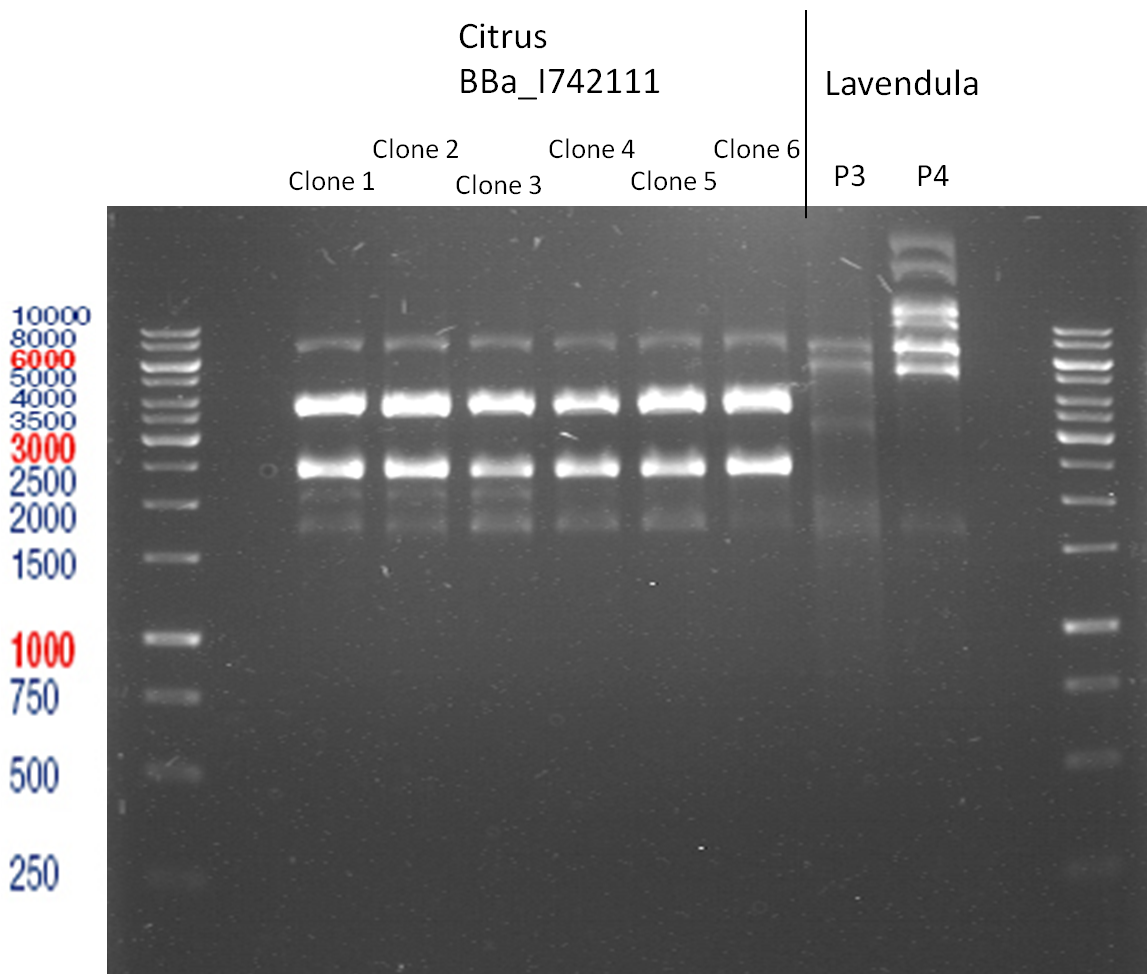
Analytic Gelelectrophoresis
Friday, June 22nd
Transformation of E.coli XL1-Blue with pKS2µHyg-PAL-4Cl-CHS
Investigator: Ingmar, Volker
Aim of the experiment: Plasmid amplification
Operation Sequence:
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the Plasmid pKS2µHyg-PAL-4Cl-CHS
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Saturday, June 23rd
Quick Change mutagenis to remove NgoMIV from pYES2
Investigator: Ingmar, Volker
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
PCR
Reaction batch
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P7 pYes2_RFC25 MCS 1.1 template |
| 0.5 µl | 1:10 dilution of O38 (10 pmol/µL) |
| 0.5 µl | 1:10 dilution of O39 (10 pmol/µL) |
| 17 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 15 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 6 min |
- The vector resulting from the PCR-product was named pYes2_RFC25 MCS 1.2.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the Plasmid P7 pYes2_RFC25 MCS 1.2
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Sunday, June 24th
Miniprep of E.coli XL1-Blue with pKS2µHyg-PAL-4Cl-CHS
Investigator: Ingmar, Volker
Aim of the experiment: Plasmid purification
Operation Sequence:
- A single clone of E.coli XL1-Blue with pKS2µHyg-PAL-4Cl-CHS was picked an transferred to 5 ml LB Amp on saturday evening. Incubation overnight at 37°C 180 rpm.
- Using a Quiagen kit a miniprep of the overnight culture was done.
Quick Change mutagenis to remove NgoMIV from pYES2
Investigator: Ingmar, Volker
Aim of the experiment: Removal of a NgoMIV restriction site in the backbone of pYes2.
Operational sequence:
- A single clone of E. coli pYes2_RFC25 MCS 1.2 was picked an transferred to 5 ml LB Amp. Incubation overnight at 37°C 180 rpm.
Transformation of 2 Biobricks into E. coli XL1-Blue
Investigator: Jeffery Truong
Aim of the experiment: Transformation of Biobricks into E. coli for plasmid propagation for PCR with new RFC pre- and suffix primer in order to do protein fusions.
- 2 Biobricks from the distribution kit were used: First, LexA (BBa_K105005, Plate 3 Well 9E) in the pSB1A2 plasmid with ampicillin resistance and second, the heme oxygenase (BBa_I15008, Plate 2 Well 13J) in the pSB2K3 plasmid with kanamycin resistance.
- 10 µL of autoclaved H2O were added to each well on the distribution kit. The well immediately turned red which means that one does it right.
- The now resuspended DNA liquids were transferred into a new ERG on ice.
- CaCL2 competent E. coli XL1-Blue cells from the stock were gently defrezed on ice.
- For each Biobrick 100 µL cells were used and pooled together with 2 µL of plasmid DNA in a ERG on ice.
- Incubation on ice for 30 min.
- 5 min heat shock at 37 °C.
- Each ERG now is transferred in a new ERG prefilled with 1 mL of LB-medium and incubated in a cell-culture shaker at 37 °C for 45 min.
- 100 µL of these cell suspension were plated on antibiotic selection plates (Ampicillin for LexA and Kanamycin for heme oxygenase).
- The rest of the cell suspension is centrifuged at 13000 rpm for 60 s and the supernatant is discarded.
- The pellet is resuspended with 100 µL for each ERG and is plated on another antibiotic selection plate
- These 4 plates were put at 37 °C overnight
Monday, June 25th
Miniprep of E. coli XL1-Blue with pYes2_RFC25 MCS 1.2
Investigator: Alois, Martin
Aim of the experiment: proof of successful removal of NgoMIV in the backbone of pYes2
Operation Sequence:
- Mini prep of pYes2 1.2. The resulting purified DNA is P33.
- Control digest of pYes2_RFC25 MCS 1.2 and p13 (+ analytical gel electrophoresis: 90 V, 1 h:
* 15 µl ddH20 * 2 µl NEBuffer4 * 0,5 µl NgoMIV * 2,5 µl pYes2 1.2/p13 * 37°C, 1 h.
Quick Change mutagenis to remove SpeI from pYES2_RFC25 MCS 1.2
Investigator: Ingmar, Volker
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
PCR
Reaction batch
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P33 template |
| 0.5 µl | 1:10 dilution of O44 (10 pmol/µL) |
| 0.5 µl | 1:10 dilution of O45 ((10 pmol/µL) |
| 17 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 15 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 6 min |
- The procedure was furthermore applied to P13 and P14.
- The vector resulting from the PCR-product was named pYes2_RFC25 MCS 1.3.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- This operation sequence was applied to the PCR prducts of P33, P13 and P14 respectively.
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Picking of E. coli cells on antibiotic selection plates: pSB1A2 plasmid with BBa_K105005 (LexA) and pSB2K3 plasmid BBa_I15008 (heme oxygenase)
Investigator: Jeffery Truong
Aim of the experiment: Picking colonies from transformed E. coli XL1-Blue, 4x picked for each Biobrick.
- pSB1A2 plasmid with BBa_K105005 (LexA): Colonies were on both ampicillin selection plates, the one with diluted cell suspension and the one with concentrated E. coli cell suspension. Typical E. coli colony morphology. Picking was performed on the plate with diluted cell suspension.
- pSB2K3 plasmid BBa_I15008 (heme oxygenase): Colonies were only on the kanamycin selection plate with concentrated cell suspension. The one with diluted susepension was empty. Typical but very small E. coli colonies. Picking was performed from the first plate.
- Picked pipette tips was transferred into a special cell-culture tubes with air-permeable, but sterile cover. In each tube 4 mL of LB-medium + ampicillin (???)(for pSB1A2) or kanamycin (35 mg/mL) (for pSB2K3).
- 4 colonies for each Biobrick was picked; total: 8 tubes overnight culture.
- These tubes were transferred in a cell culture shaker at 37 °C and were incubated overnight
Tuesday, June 26th
Quick Change mutagenis to remove SpeI from pYes2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Removal of a SpeI restriction site in the backbone of pYes2.
Operational sequence:
- For each transformation of the PCR-products of P14 and P33 a single clone was picked an transferred to 6 ml LB Amp. Incubation overday at 37°C 180 rpm. The transfomation with the PCR product of P13 was not successfull. Therfore no clone could be picked.
- Using a Quiagen kit a miniprep of the overnight culture was done.
- The resulting purified DNA was aliquoted in new tubes labeled as follows:
PCR product of P33(transformation done by Ingmar): P29
PCR product of P33(transformation done by Saskia&Jara): P30
PCR product of P14(transformation done by Ingmar): P31
PCR product of P14(transformation done by Saskia&Jara): P32
- Afterwards a control digestion of P29-P32 was done.
Reaction batch
| Plasmid | P29 | P30 | P31 | P32 |
| NEB4 buffer | 2 µl | 2 µl | 2 µl | 2 µl |
| DNA | 2,5 µl | 2,5 µl | 5 µl | 5 µl |
| SpeI-HF | 0.25 µl | 0.25 µl | 0.25 µl | 0.25 µl |
| NgoMIV | 0.25 µl | 0.25 µl | ||
| ddH2O | 15 µl | 15 µl | 12.75 µl | 12.75 µl |
| Sum | 20 µl | 20 µl | 20 µl | 20 µl |
- Incubation at 37 °C for 1h.
- Verification of control digest by agarose gel electrophoresis:
20 µl of each digest product was mixed with 2 µl DNA loading buffer and loaded into the gel. The separation process lasted 1h at 90 V.
Verification of the PCR products P30, P31 and P33
Investigator: Saskia, Jara
Aim of the experiment: Verification of the PCR produts P30, P31 and P33
Nano Drop
| Sample | concentration [ng/µl] | 260/280 |
| P33 | 1072.6 | 1.01 |
| P30 | 1588 | 1.28 |
| P31 | 926.2 | 0.82 |
Analytical gel electrophoresis
- gel: 1 %
- band 1: 10 µl DNA ladder (1kb)
- band 2: P30
- band 3: P33
- band 4: P31
Control of the competent cells and transformation with P20
Investigator: Saskia, Jara
Aim of the experiment: Control of the competent cells and transformation with P20 Transformation
- competent cells: by Simon and Ingmar
- plasmid: P20
result:
- successful transformation: red colonies
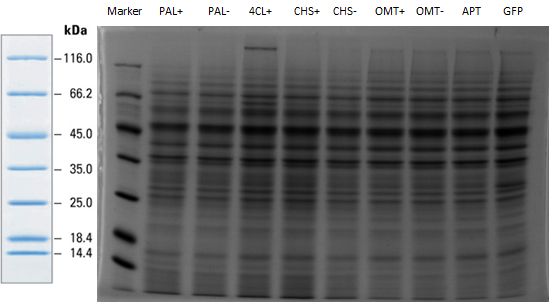
PCR of PAL, 4CL, CHS, OMT (Coumaryl-CoA)
Investigator: Daniela, Mary
Determination of concentration of plasmids (Nanodrop): c(pKS2µHyg-PAL-4CL-CHS) = 500 ng/µl c (pOMT) = 20 ng/µl
PCR:
| Name of tube | Enzyme | consensus (+)/ consensless (-) | used Oligos |
| CHS - | CHS | - | O13 and O24 |
| CHS + | CHS | + | O23 and O24 |
| PAL - | PAL | - | O15 and O16 |
| PAL + | PAL | + | O22 and O16 |
| OMT - | OMT | - | O17 and O26 |
| OMT + | OMT | + | O25 and O26 |
| 4CL - | 4CL | - | O19 and O20 |
| 4CL + | 4CL | + | O21 and O20 |
Reaction batch
| volume | reagent |
| 5 µl | 10x Pfu Ultra II buffer |
| 4 µl | dNTP's (each 2.5 mM) |
| 0.5 µl | Pfu Ultra II (2.5 U/µL) |
| 5 µl | 1:10 dilution of used forward primers (10µM) |
| 5 µl | 1:10 dilution of used reversed primers (10µM) |
| 1 µl | DNA (pKS2µHyg-PAL-4CL-CHS 50 ng/µL or pOMT 20 ng/µL) |
| 29.5 µL | bidest. sterile Water |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 5 min (and adding Pfu Ultra after 3 min) |
| 2 | 30 | 95°C | 30 sec |
| 46°C | 2.5 min | ||
| 72°C | 1.5 min | ||
| 3 | 72°C | 5 min |
PCR purification
- Purification was done using QIAquick PCR Purification Kit (250)
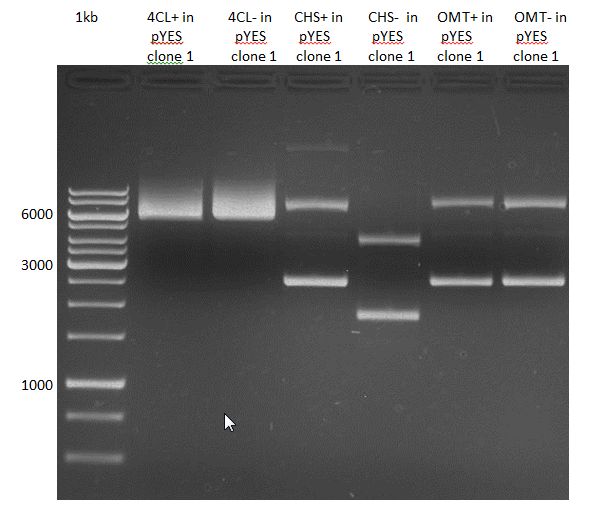
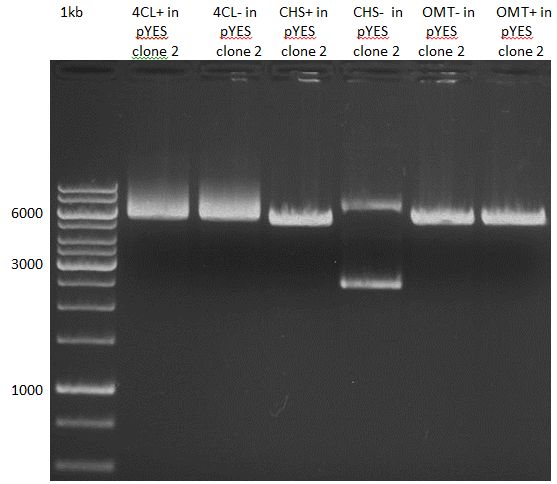
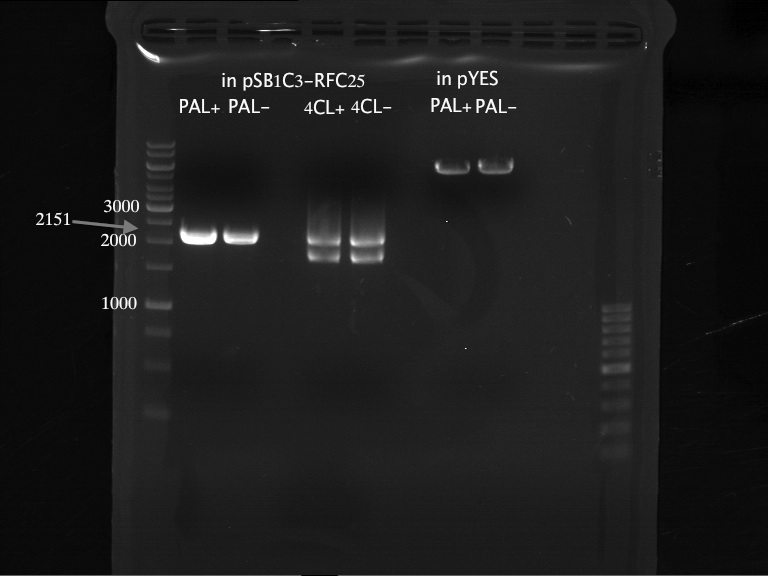
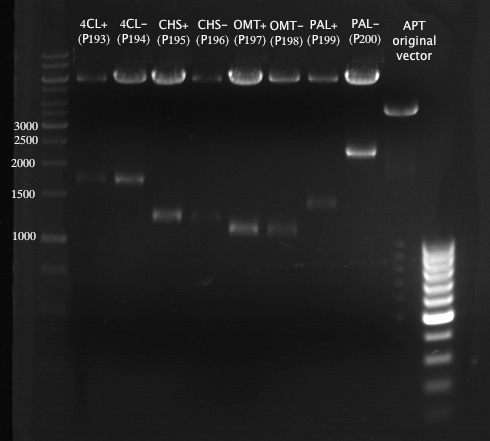
Analytical Gel Electrophoresis:
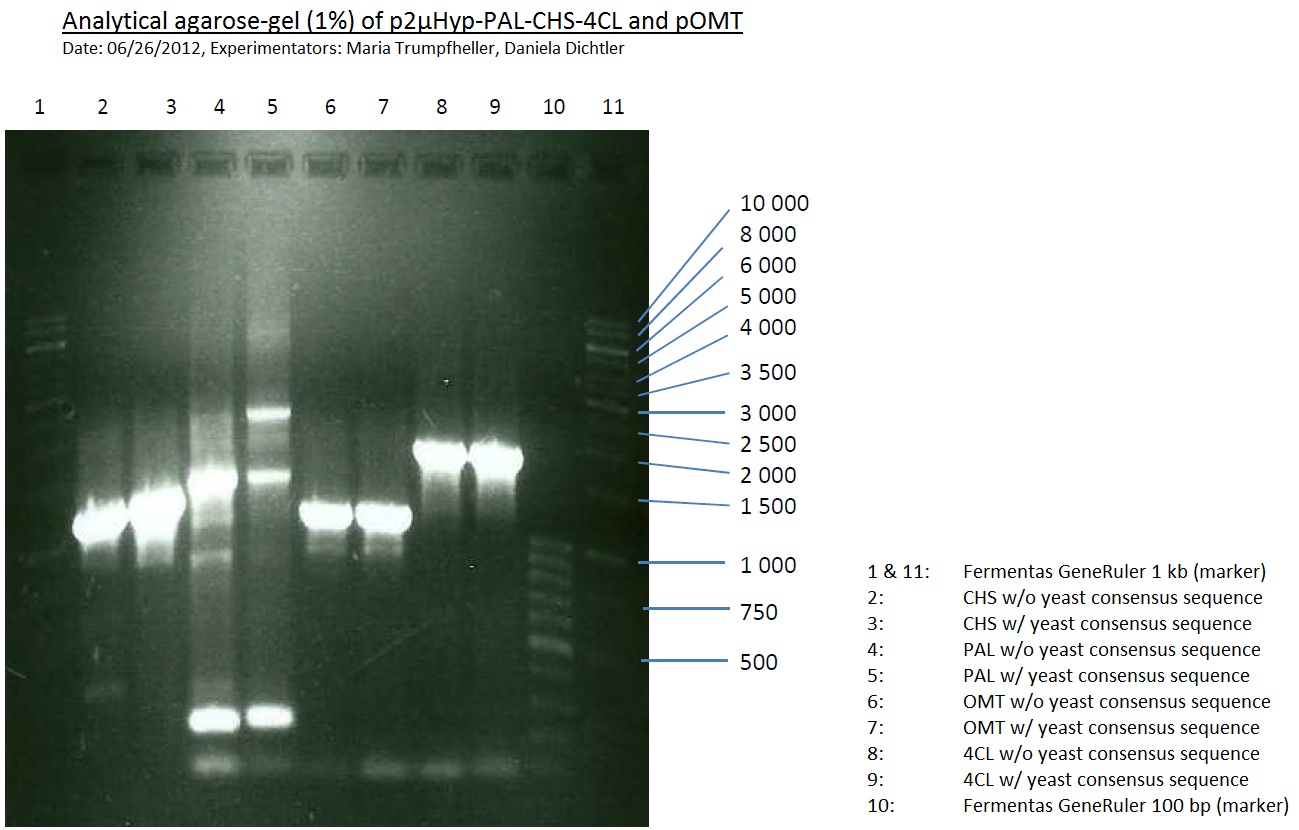
-> going on with CHS, 4CL and OMT; the PCR of PAL will be repeated
Miniprep and analytical gel of picked transformed overnight culture with pSB1A2 plasmid with BBa_K105005 (LexA) pSB2K3 plasmid BBa_I15008 (heme oxygenase)
Investigator: Jeffery Truong, Georg Schützinger
Aim of the experiment: Plasmid isolation from the picked transformed overnight E. coli cells with pSB1A2 plasmid with BBa_K105005 (LexA) pSB2K3 plasmid BBa_I15008 (heme oxygenase).
- The LB-medium with antibiotics of every tube was opaque which means that the picked cells were successfully inoculated.
- Centrifugation step at 5000 rpm for 10 min at 16 °C.
- Every single step now was performed on ice.
- Miniprep (Qiagen Qiaprep spin) after manufacturer's protocoll.
- Analytical restriction master mix was prepared after following scheme (Using XbaI and PstI):
- 4.4 µL XbaI
- 4.4 µL PstI
- 17.6 µL Tango-buffer (10x)
- 132 µL ELGA H2O
- 17.5 µL from the master mix was poooled together with 2.5 µL of plasmid DNA. That corresponds to 2.5 µL of plasmid DNA, 0.25 µL XbaI, 0.25 µL PstI, 2 µL Tango-buffer (10x), 15 µL ELGA H2O.
- Incubation at 37 °C for 120 min on a ERG heating unit.
- BUT: error was performed during preparing the digested plasmid DNA for analytical gelelectrophoresis in the dilution step. 1:10 dilution of analyctical probe with DNA loading buffer:
- 3.3 µL sample (contains already 1x loading buffer!)
- 0.7 µL loading buffer (?x)
- 6 µL TAE-buffer
- Should have done: 3 µL sample + 1 µL loading buffer (10x) + 6 µL TAE-buffer (1x)
- DNA-ladder preperation: 10 µL ladder stock solution + 10 µL DNA loading buffer + 80 µL TAE-buffer. 10 µL of this solution was pipetted in one gel pocket of the prepared 1% agarose gel including ehtiudiumbromid.
- 20 µL of each samples were also pipetted into the gel pockets.
- The gel pockets were pipetted after following scheme:
| Heme oxygenase (colony 1) | Heme oxygenase (colony 2) | Heme oxygenase (colony 3) | Heme oxygenase (colony 4) | DNA-ladder | LexA (colony 1) | LexA (colony 2) | LexA (colony 3) | LexA (colony 4) |
- Gel electrophoresis at 90 V
- After 20 min the resolution was still poor; 20 min longer.
- Analytical Gel okay, but samples interchanged
Wednesday, June 27th
Friday, June 29th
Preparative digest of PCR-products of 4CL, CHS and OMT
Investigator: Katrin, Mary
each digestion will dure 2.5h at 37°C
- CHS: digestion with Xba1 and HF-Age1 (both NEB)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer NEB4 |
| 0.5µl | BSA |
| 1µl | Xba1 (NEB; 20u/µl) |
| 1µl | HF-Age1 (NEB; 20u/µl) |
| 17.5µl | bidest. sterile H2O |
- OMT: digestion with Xba1 and HF-Age1 (both NEB)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer NEB4 |
| 0.5µl | BSA |
| 1µl | Xba1 (NEB; 20u/µl) |
| 1µl | HF-Age1 (NEB; 20u/µl) |
| 17.5µl | bidest. sterile H2O |
- 4CL: digestion with Xba1 and Pst1 (both Fermentas)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer Tango |
| 2µl | Xba1 (Fermentas; 10u/µl) |
| 3µl | Pst1 (Fermentas; 10u/µl) |
| 15µl | bidest. sterile H2O |
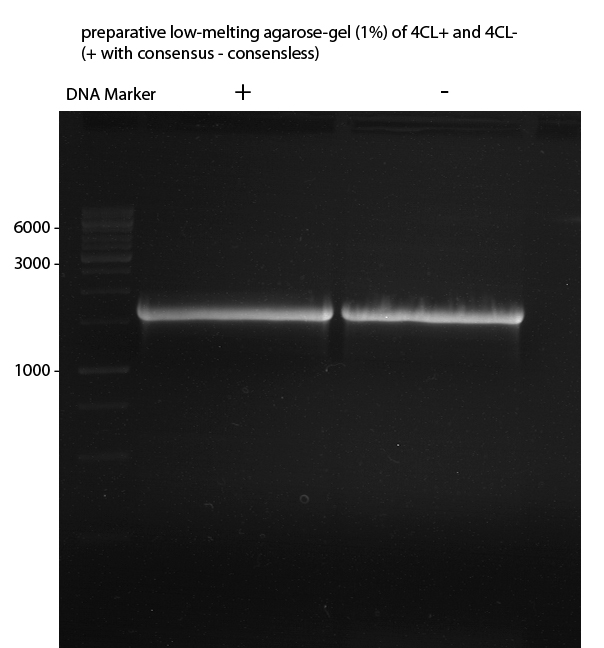
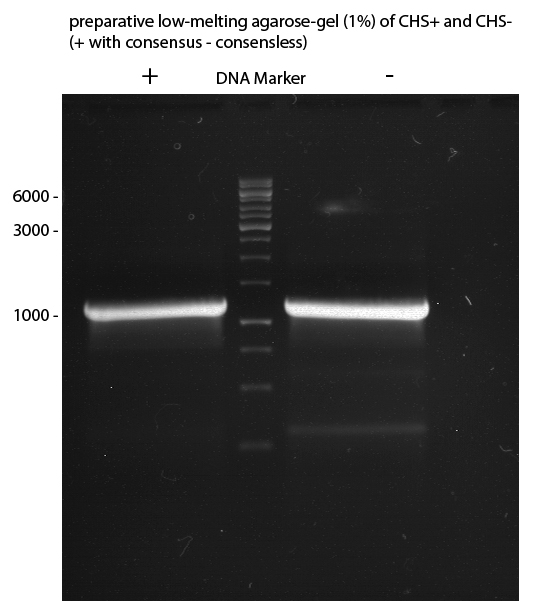
Preparative Gelelectrophoresis of PCR-products of 4CL, CHS
Investigator: Katrin, Mary
Gelextraction of 4CL+, 4CL-, CHS+, CHS- (bands are as expected; +=with consensus-sequence, -=without consensus-sequence)
DNA-purification with Kit from Quiagen
Quick Change mutagenesis to remove PstI in URA3 from pYES2_RFC25 MCS 1.2
Investigator: Ingmar
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
PCR
Reaction batch
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P29 template |
| 0.5 µl | 1:10 dilution of O40 (10 pmol/µL) |
| 0.5 µl | 1:10 dilution of O41 ((10 pmol/µL) |
| 17 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 15 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 6.5 min |
- The procedure was furthermore applied to P31.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- This operation sequence was applied to the PCR prducts of P29 and P31 respectively.
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Saturday, June 30th
Quick Change mutagenis to remove PstI in the URA 3 gene from pYes2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Removal of a PstI restriction site in the backbone of pYes2.
Operational sequence:
- For each transformation of the PCR-products of P29 and P30 a single clone was picked an transferred to 6 ml LB Amp. Incubation overnight at 37°C 180 rpm. The transfomation with the PCR product of P31, P32 and P33 was not successfull. Therfore no clone could be picked from these plates and four instead of one clone was picked from the plates containing the transformations of P29.
July
Sunday, July 1st
Quick Change mutagenis to remove PstI in the URA 3 gene from pYes2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Removal of a PstI restriction site in the backbone of pYes2.
Operational sequence:
- Using a Quiagen kit a miniprep of the overnight culture was done.
- The resulting purified DNA was aliquoted in new tubes labeled as follows:
1st PCR product of P29: P34
2nd PCR product of P29: P35
3rd PCR product of P29: P36
4th PCR product of P29: P37
PCR product of P30: P38
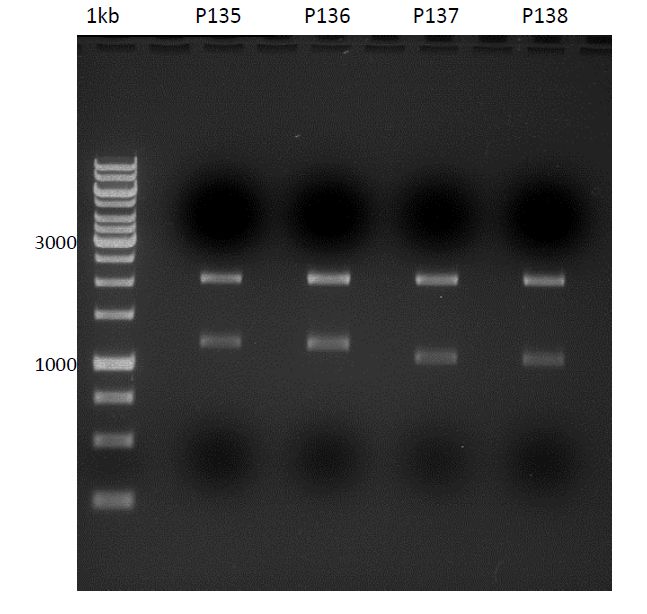
- Afterwards a control digestion of P34-P38 was done.
Reaction batch
| Plasmid | P34 | P35 | P36 | P37 | P38 |
| Fermentas 10x R buffer | 0.5 µl | 0.5 µl | 0.5 µl | 0.5 µl | 0.5 µl |
| DNA | 2 µl | 2 µl | 2 µl | 2 µl | 5 µl |
| PstI | 0.25 µl | 0.25 µl | 0.25 µl | 0.25 µl | 0.25 µl |
| ddH2O | 17.25 µl | 17.25 µl | 17.25 µl | 17.25 µl | 14.25 µl |
| Sum | 20 µl | 20 µl | 20 µl | 20 µl | 20 µl |
- Incubation at 37 °C for 1h.
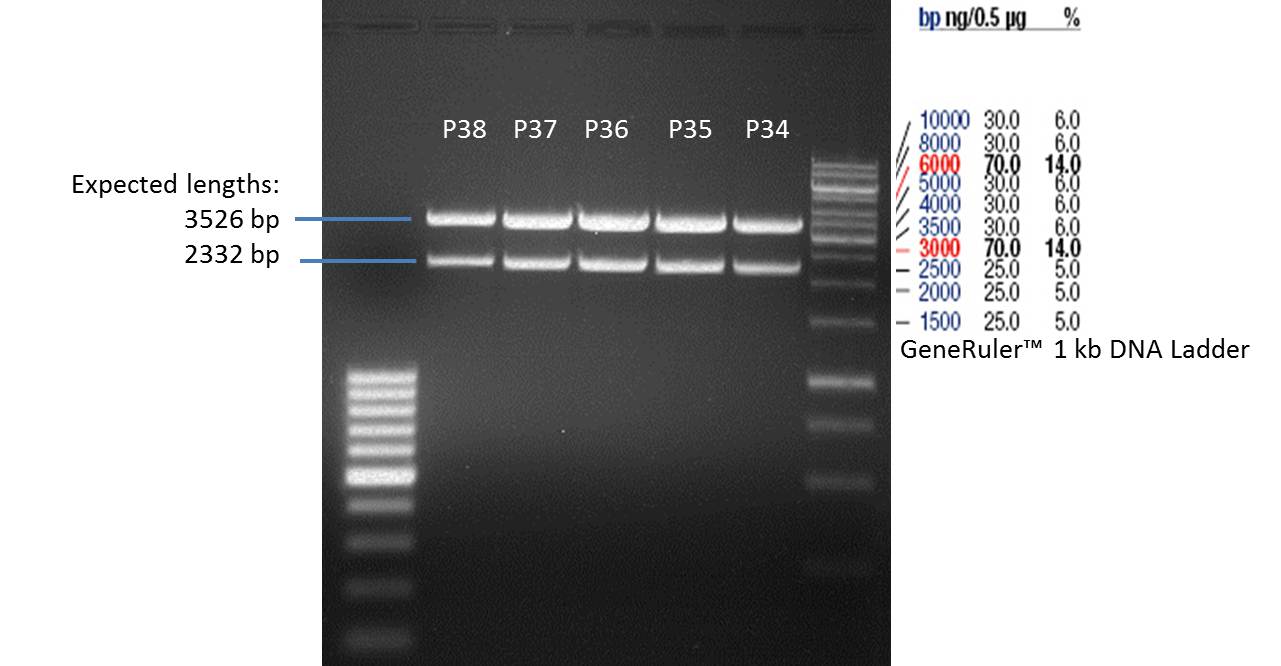
- Verification of control digest by agarose gel electrophoresis:
20 µl of each digest product was mixed with 4 µl 6x DNA loading buffer and loaded into the gel. The separation process lasted 1h at 90 V.
- All digest products show the expected two bonds at 3526 bp and at 2332 bp. The Miniprep product P35 was chosen to be used for the further Quickchange Mutagenesis.
Quick Change mutagenesis to remove PstI in the 2µ ori from pYES2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P35 template |
| 0.5 µl | 1:10 dilution of O42 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P35 template |
| 0.5 µl | 1:10 dilution of O43 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 6 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Monday, July 2nd
Repetition of PCR of PAL
Investigator: Mary
Reaction batch
| volume | reagent |
| 5 µl | 10x Pfu Ultra II buffer |
| 4 µl | dNTP's (each 2.5 mM) |
| 0.5 µl | Pfu Ultra II (2.5 U/µL) |
| 5 µl | 1:10 dilution of used forward primers (10µM) |
| 5 µl | 1:10 dilution of used reversed primers (10µM) |
| 1 µl | DNA (pKS2µHyg-PAL-4CL-CHS 50 ng/µL) |
| 29.5 µL | bidest. sterile Water |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 2 min (and adding Pfu Ultra after 2 min) |
| 2 | 30 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 72°C | 2.5 min | ||
| 3 | 72°C | 5 min |
PCR of LexA with primers including the RFC25 pre- and suffix
Investigator: Jeffery Truong, Georg Schützinger
Aim of the experiment: The Biobrick BBa_K105005 (LexA) has a RFC10 pre- and suffix, but we need RFC25 pre- and suffix for protein fusion, so we have to do a PCR with primer containing the RFC25 pre- and suffix.
- The received forward and reverse primer TUM12-LexA-fw and TUM12-LexA-rv are resuspended in 204 µL and 221 µL ELGA water as described in the data sheet to get a final primer concentration of 100 pmol/µL=100 µM. For the PCR reaction mixture we took 0.5 µL of these resuspended primer and add 4.5 µL of ELGA water to get a final primer concentration of 10 µM.
- Clone 3 of BBa_K105005 (LexA) has beed choosen for the PCR (ERG No. P23).
PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer |
| 1 µl | 10 µM Reverse Primer |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA (BBa_K105005) from P23 (Clone 3) |
| 35.75 µL | ELGA Water |
| =50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 55 °C | 60 s | |
| 68 °C | 60 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | overnight |
Tuesday, July 3rd
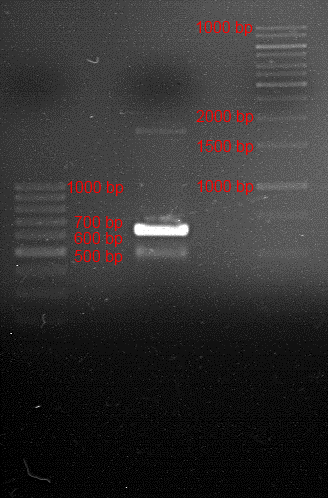
Analytic gelelectrophoresis of cleaned up PCR product from LexA with primer containing RFC25 pre- and suffix
Investigator: Jeffery Truong
Aim of the experiment: Analytical gelelectrophoresis of cleaned up PCR product from LexA (BBa_K105005) with primer containing the RFC25 pre- and suffix (TUM12-LexA-fw and TUM12-LexA-rv).
- The clean-up of the PCR product LexA (BBa_K105005) with primer containing the RFC25 pre- and suffix (TUM12-LexA-fw and TUM12-LexA-rv) was performed with the QIAquick PCR Purification Kit from Qiagen after manufacturer's protocoll.
- 5 µL of the purificated PCR product was taken to perform a analytical gelelectrophoresis to verify the success of the PCR.
- 1% agarose gel containing ethidium bromide was used for the analytical gelelectrophoresis.
- The analytical gelelectrophoresis was performed for 60 min at 90 V.
- Scheme of the gel:
| 100 bp ruler | PCR product of BBa_105005 | 1000 bp ruler |
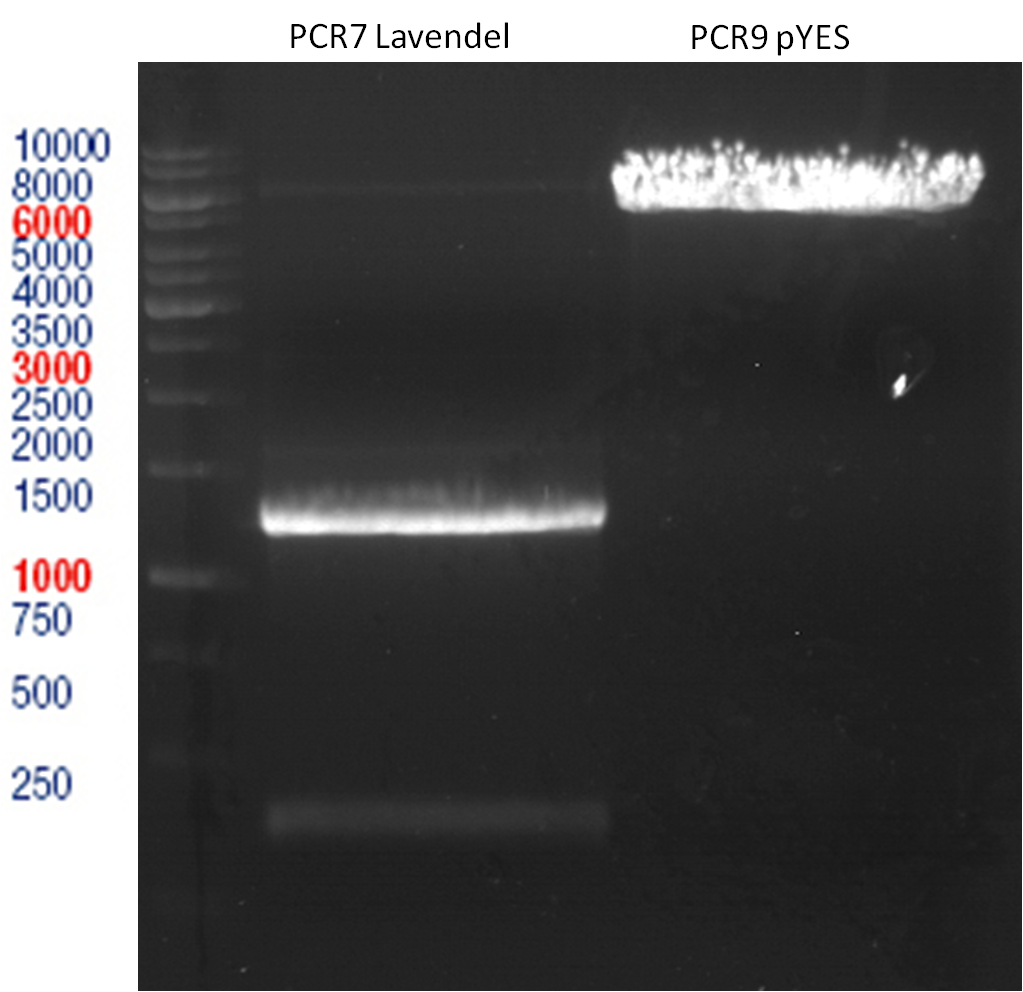
LS: Plating of Schwab expression stains #106 & #108 which contain lavendula limonene synthase
Investigator: Lara Kuntz
Aim of the experiment: To get colonies of BL21 strains containing lavendula LS for amplification and subsequent plasmid extraction.
- Schwab strain #106 was plated on a chloramphenicol containing LB plate. #108 was plated on a amp+chlp containing LB plate. The plates were incubated at 37°C over night.
Quick Change mutagenis to remove PstI in the 2µ Ori from pYes2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Removal of a PstI restriction site in the backbone of pYes2.
Operational sequence:
- From the transformation of the PCR-product of P35 two single clones were picked an transferred to 6 ml LB Amp. Incubation overnight at 37°C 180 rpm.
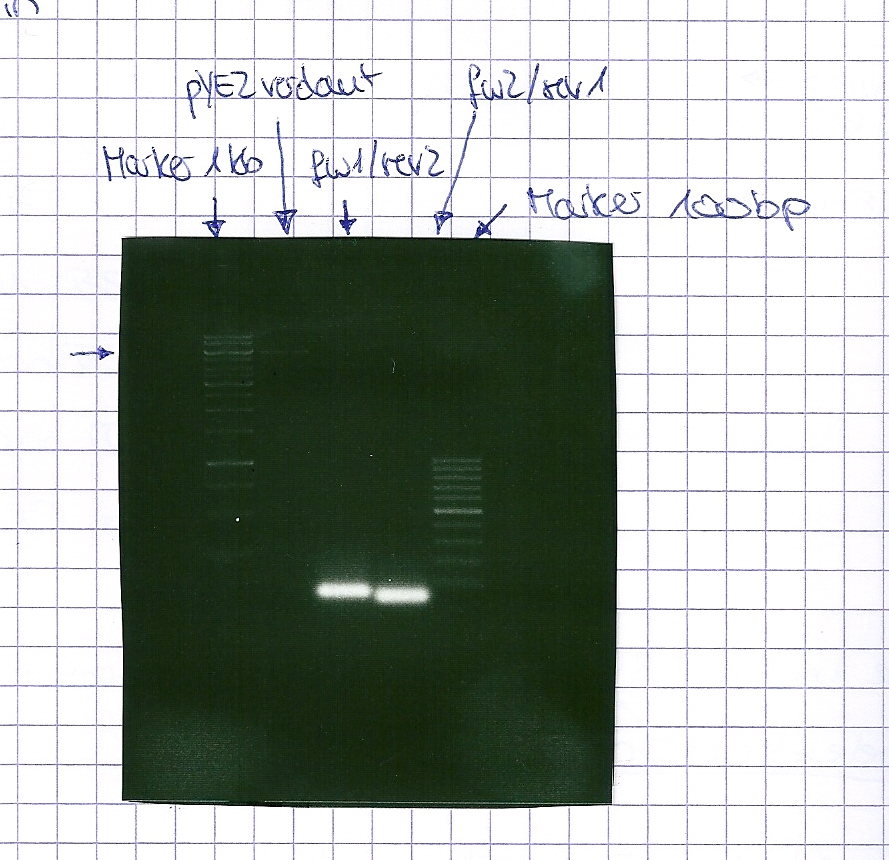
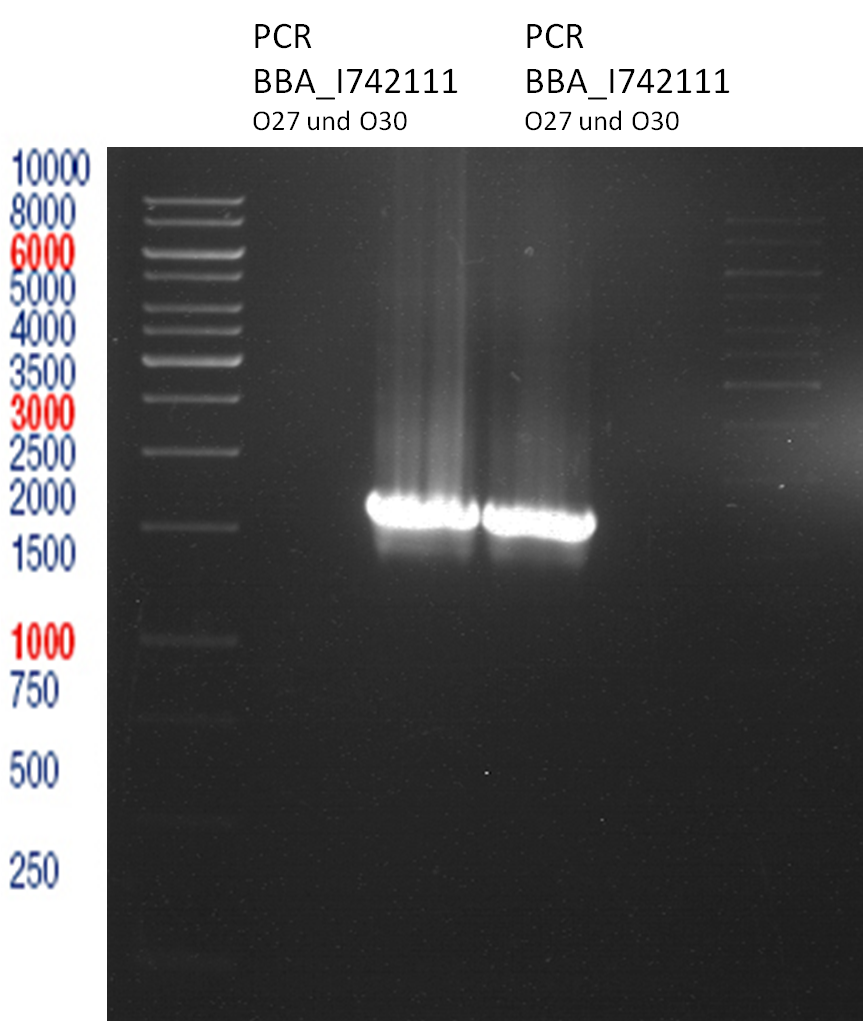
PCR of BBa_I742111 (Limonenesynthase) Clone 3 (Trafo 19.06.12)
Investigator: Andrea
PCR used forward primer with consensus sequence; PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer O27 |
| 1 µl | 10 µM Reverse Primer O30 |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µl) |
| 1 µl | Plasmid DNA (BBa_I742111) Clone 3 |
| 35.75 µl | dd water |
| 50 µL | TOTAL |
PCR used forward primer without consensus sequence; PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer O28 |
| 1 µl | 10 µM Reverse Primer O30 |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µl) |
| 1 µl | Plasmid DNA (BBa_I742111) Clone 3 |
| 35.75 µl | dd water |
| 50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 47 °C | 30 s | |
| 68 °C | 1,75 min | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | 1 h |
Analytical Gelelectrophoresis
- 5 µl DNA + 1 µl loading buffer
Wednesday, July 4th
Quick Change mutagenis to remove PstI in the 2µ Ori from pYes2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Removal of a PstI restriction site in the backbone of pYes2.
Operational sequence:
- Using a Quiagen kit a miniprep of the overnight culture was done.
- The resulting purified DNA was aliquoted in new tubes labeled as follows:
1st Transformation of P35: P43
2nd Transformation of P35: P44
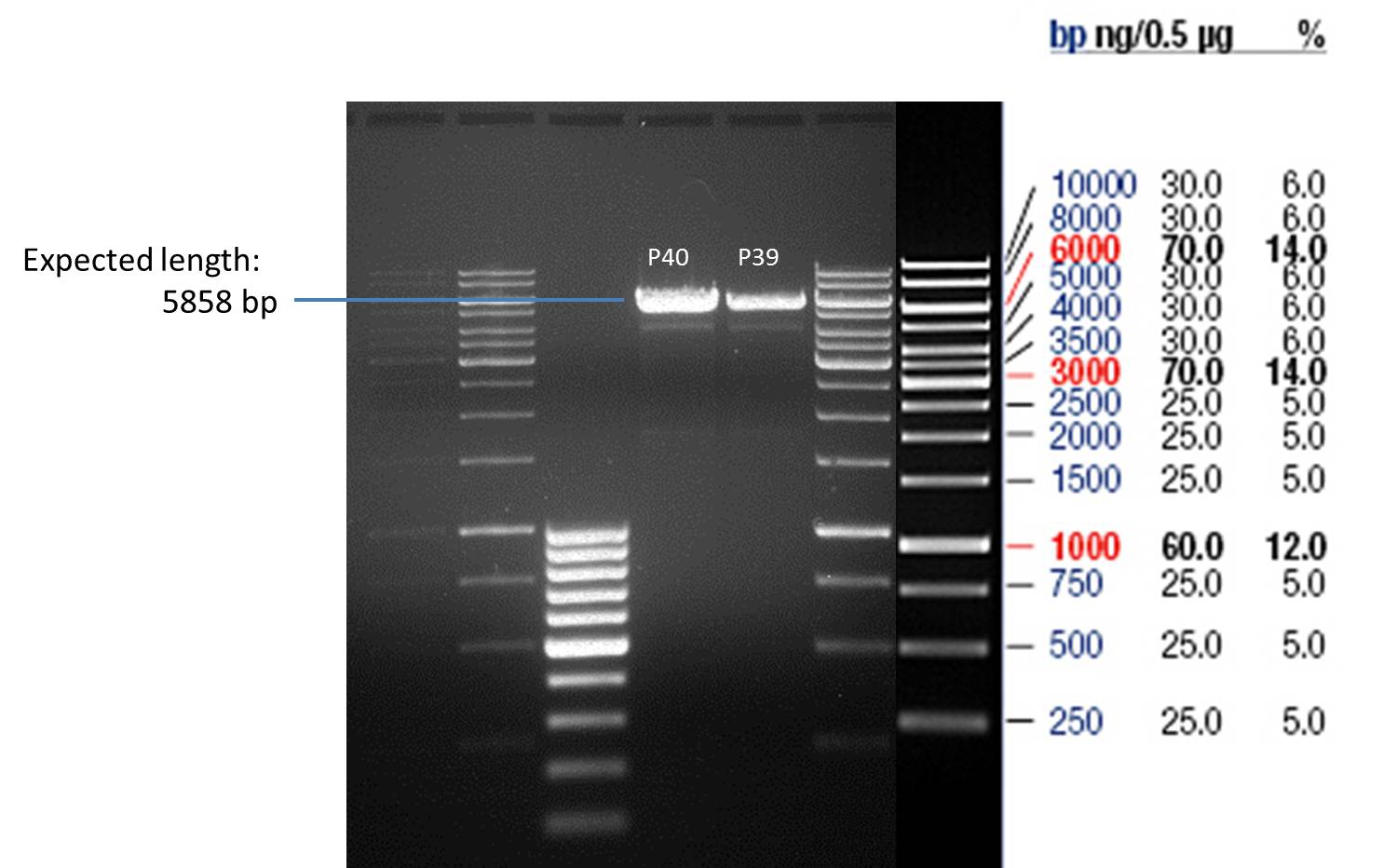
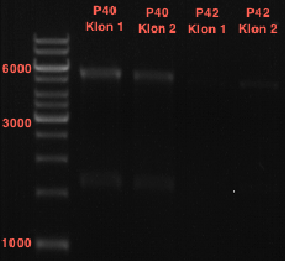
- Afterwards a control digestion of P43 and P44 was done.
Reaction batch
| Plasmid | P43 | P44 |
| Fermentas 10x R buffer | 0.5 µl | 0.5 µl |
| DNA | 5 µl | 5 µl |
| PstI | 0.25 µl | 0.25 µl |
| ddH2O | 14.25 µl | 14.25 µl |
| Sum | 20 µl | 20 µl |
- Incubation at 37 °C for 1h.
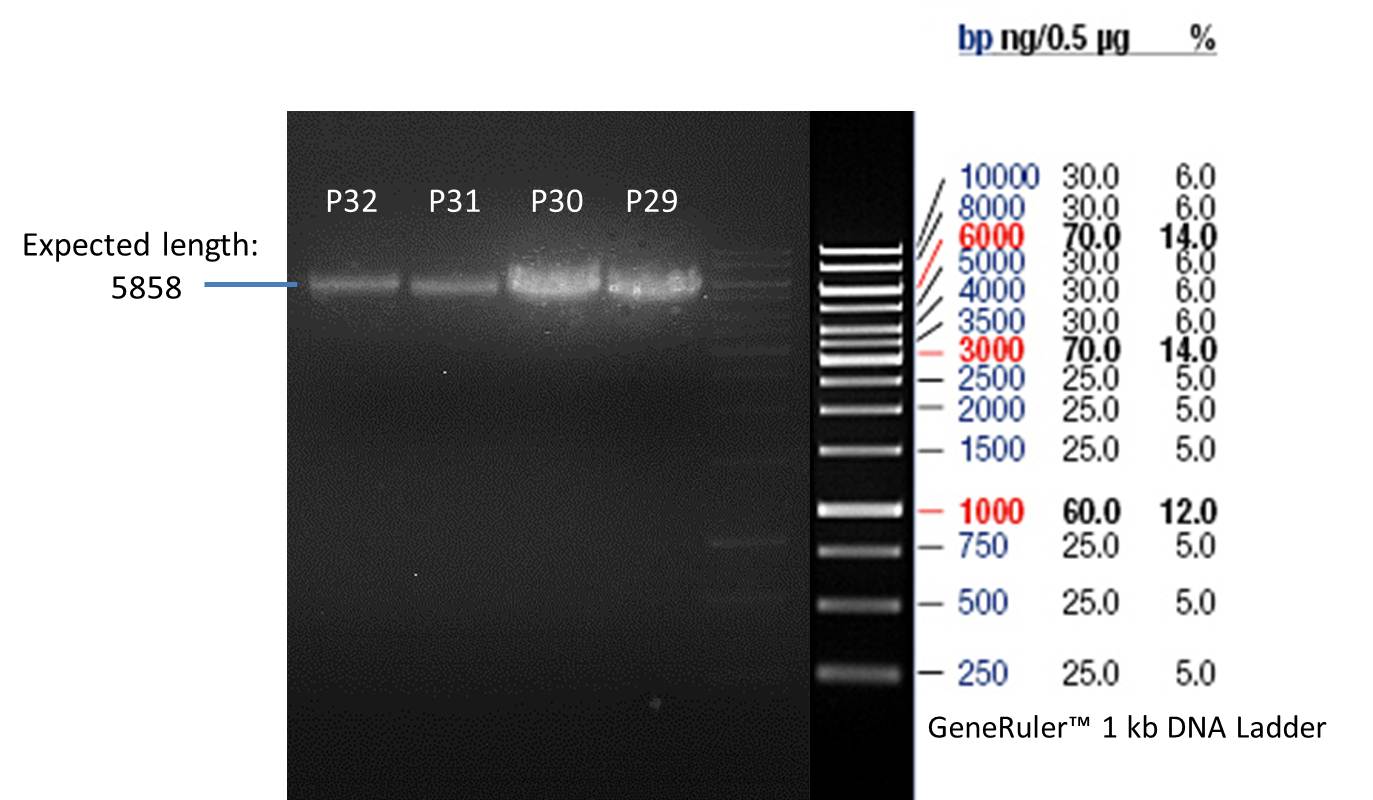
- Verification of control digest by agarose gel electrophoresis:
20 µl of each digest product was mixed with 4 µl 6x DNA loading buffer and loaded into the gel. The separation process lasted 1h at 90 V.
- All digest products show the expected bond at 5858 bp. The Miniprep product P40 was chosen to be used for the further Quickchange Mutagenesis.
Quick Change mutagenis to insert Ala in front of the Strep - tag II in pYES2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector; operating purfication possibility via Strep-tag II.
PCR
Reaction batch
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P44 |
| 1 µl | 1:10 dilution of O54 (10 pmol/µL) |
| 1 µl | 1:10 dilution of O55 ((10 pmol/µL) |
| 16 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 16 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 6 min |
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Analytical Gelelektrophoresis of PCR-Products of PAL
Investigator: Mary
Aim of the experiment: purification and testing if PCR was successful
Purification of PCR-Products with purification kit from quiagen analytical gelelectrophoresis of PAL+, PAL-; expected band at 2,1 kb
Analytical Gel Electrophoresis: File:TUM12 20120704 PAL-PCR v2.tiff
-> PCR was not successful, no band at 2,1 kb
-> next steps: new Design of Primer and repetition of PCR with new primers
Picking of clones of Schwab expression stains #106 & #108
Investigator: Andrea
Aim of the experiment: Getting clones of cells with plasmids containing the gene for limonenesynthase and amplification of these clones for further plasmid preparation.
Picking of Clones
- 2 clones of every stain were picked
- Incubation at 37 °C in LB + Amp (#108) / LB + Kan (#106)
Preparative Gelelectrophoresis of PCR-products of OMT
Investigator: Mary, Katrin
Aim of the experiment: Purification of the previously digested DNA, test if digestion was successful
picture follows!
Gelextraction of OMT- and OMT+ (bands are as expected; +=with consensus-sequence, -=without consensus-sequence)
DNA-purification with Kit from Quiagen
Thursday, July 5th
Preparation of plasmids containing lavendula LS
Investigator: Lara
Aim: Purify Schwab plasmids containing lavendula limonene synthase
Experiment was conducted using Qiagen Plasmid Miniprep Kit.
P39: Plasmid from Schwab strain #106 (1); 35 ng/µl
P40: Plasmid from Schwab strain #106 (2); 72 ng/µl
P41: Plasmid from Schwab strain #108 (1); 240 ng/µl
P42: Plasmid from Schwab strain #108 (2); 120 ng/µl
Restriction digest of Schwab plasmids
Investigator: Lara
Aim: To check whether extracted plasmids from Schwab expression strains #106 & #108 are OK.
Digest of plasmid from strain #106 was conducted with the following protocoll:
500 ng Plasmid DNA
0,25 µl Nco1
0,25 µl Hind 3
2 µl Buffer Tango
dd H2O to a total volume of 20 µl.
Digest of plasmid from strain #108 was conducted with the following protocoll:
500 ng Plasmid DNA
0,25 µl EcoR1
0,25 µl Not1
2 µl Buffer Orange
dd H2O to a total volume of 20 µl.
(All enzymes and buffers were from Fermentas).
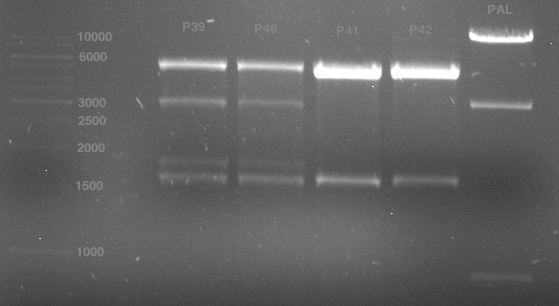
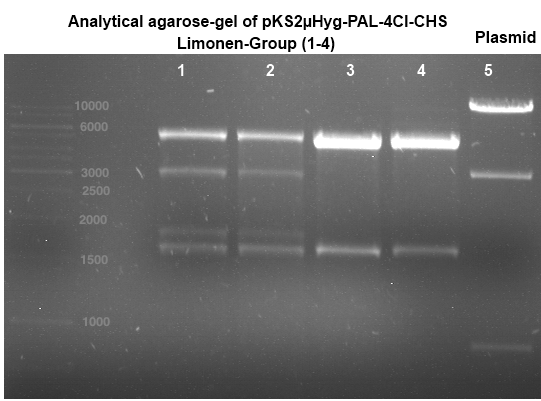
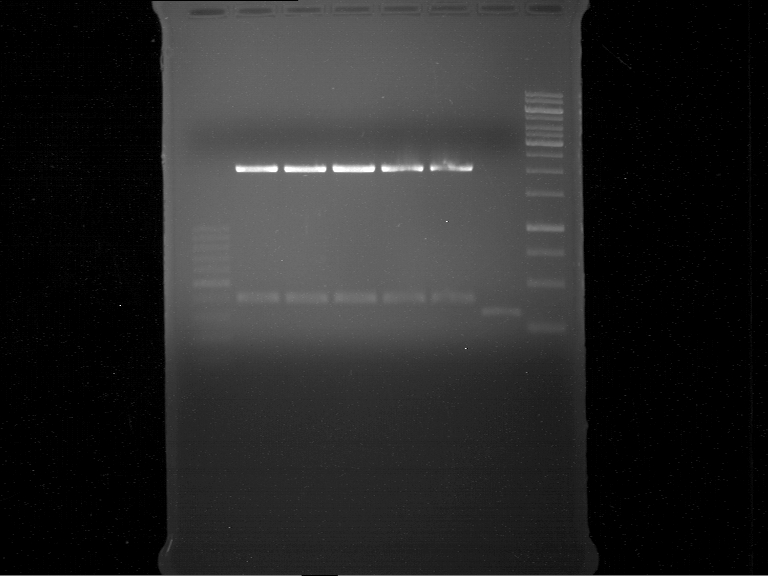
Analytical gel electrophoresis of digested Schwab plasmids
Investigator: Lara Aim: Check plasmids for insert.
Plasmids were digested, 2 µl loading dye (10x) was added to each sample. Gel was loaded in the following order:
1. 6 µl gene ruler 1 kb, 2.-6. 11 µl of: P39, P40, P41, P42, Coumaryl-Plasmid (Katrin)
Analytical Gelelektrophoresis of plasmid pKS2µHyg-PAL-4Cl-CHS
Investigator: Katrin
Aim of the experiment: testing if PAL is part of the plasmid that was sent to us (troubleshooting because PCR of PAL was not successful)
digestion took 1 h at 37°C
- digestion with ApaI (Fermentas)
| volume | reagent |
| 9.3 µl | plasmid pKS2µHyg-PAL-4Cl-CHS (miniprep) |
| 2 µl | Buffer B |
| 0.5 µl | ApaI (Fermentas; 10u/µl) |
| 8.2 µl | bidest. sterile H2O |
analytical gelelectrophoresis: expected band at 3,14 kb (Gal-PAL-XK)
-> experiment was successful, PAL is part of the plasmid pKS2µHyg-PAL-4Cl-CHS
Quick Change mutagenis to insert Ala in front of the Strep - tag II in pYES2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector; operating purfication possibility via Strep-tag II.
Operational sequence:
- From the transformation of the PCR-product of P44 two single clones were picked an transferred to 6 ml LB Amp. Incubation overday at 37°C 180 rpm.
- Using a Quiagen kit a miniprep of the overnight culture was done.
- The resulting purified DNA was aliquoted in new tubes labeled as follows:
1st picked clone of P44: P45
2nd picked clone of P44: P46
Minipreparation of biobricks BBa_J52028, BBa_E2030, BBa_E2020
Investigator: Martin, Alois
Aim: Getting "reporter proteins"
- 10 µl water war added to the well of the distribution kit (=> red)
- BBa_J52028: GFP with PEST191 tag => p51
- BBa_E2030: EYFP, yeast optimized => p52
- BBa_E2020: ECFP, yeast optimized => p53
- Transformation + Minipreparation (Qiagen Plasmid Miniprep Kit)
Friday, July 6th
Quick Change mutagenis to insert Ala in front of the Strep - tag II in pYES2_RFC25 MCS
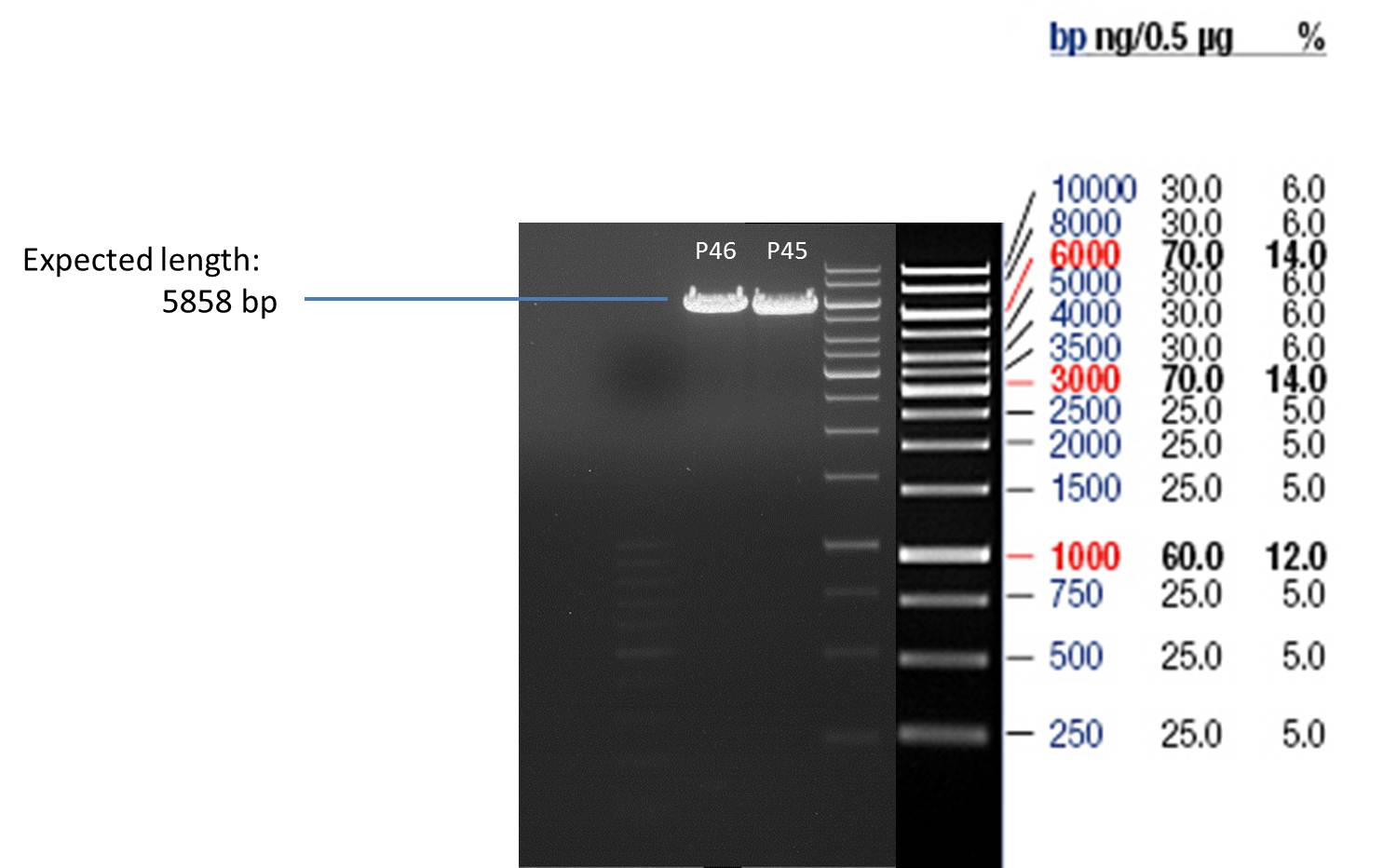
- Afterwards a control digestion of P45 and P46 was done.
Reaction batch
| Plasmid | P45 | P46 |
| Fermentas 10x O buffer | 2 µl | 2 µl |
| DNA | 3 µl | 3 µl |
| PstI | 0.25 µl | 0.25 µl |
| ddH2O | 14.75 µl | 14.75 µl |
| Sum | 20 µl | 20 µl |
- Incubation at 37 °C for 1h.
- Verification of control digest by agarose gel electrophoresis:
20 µl of each digest product was mixed with 4 µl 6x DNA loading buffer and loaded into the gel. The separation process lasted 1h at 90 V.
- All digest products show the expected bonds at 5858 bp. The Miniprep product P45 was chosen to check the insertion of Ala in front of the strep-tag II via DNA sequencing.
The results of the sequencing are shown below:
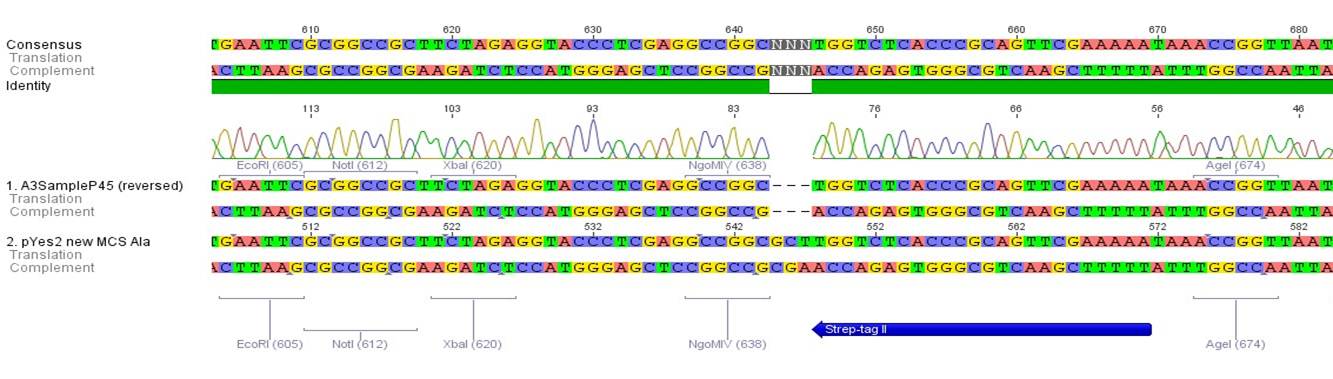 The sequencing results show that the insertion of Alanin in front of the Strep-tag II was not successful.
The sequencing results show that the insertion of Alanin in front of the Strep-tag II was not successful.
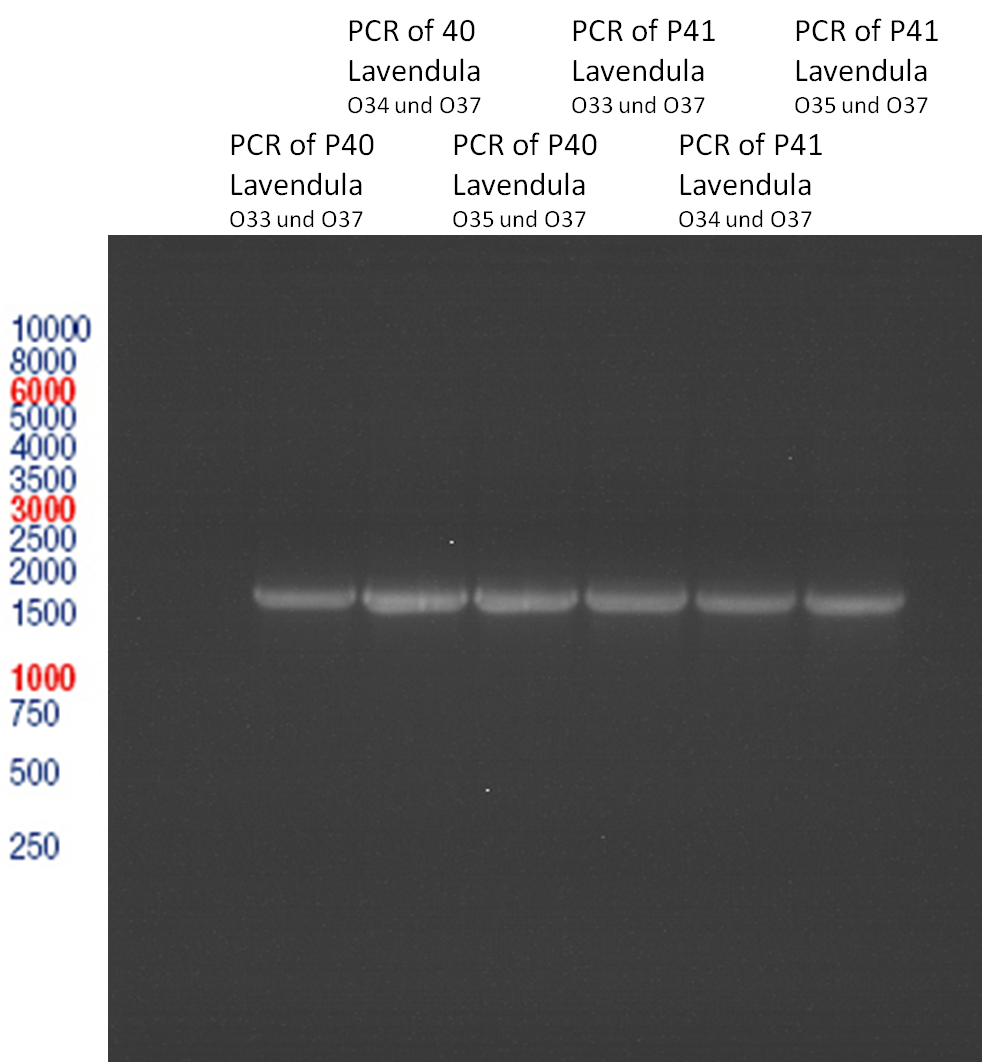
PCR of Schwab plasmid DNA to amplify gene for lavendula LS
Instructor: Lara
Aim: PCR of Schwab plasmids with primers which were designed to amplify the lavendula LS gene and to add RFC25 restriction sites.
3 different primer combinations were used:
1. O33/O37
2. O34/O37
3. O35/O37
Each primer combination was used for plasmid DNA amplification of P40 and P41.
PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer |
| 1 µl | 10 µM Reverse Primer |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Final: 1.25 units/50 µl) |
| 1 µl | Plasmid DNA (BBa_I742111) Clone 3 |
| 35.75 µl | dd water |
| 50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 50 °C | 30 s | |
| 68 °C | 1,75 min | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | 1 h |
Minipreparation of biobricks BBa_J52028, BBa_E2030, BBa_E2020
Investigator: Martin, Alois
Aim: Getting "reporter proteins"
- 10 µl water war added to the well of the distribution kit (=> red)
- BBa_J52028: GFP with PEST191 tag => p51
- BBa_E2030: EYFP, yeast optimized => p52
- BBa_E2020: ECFP, yeast optimized => p53
- Transformation + Minipreparation (Qiagen Plasmid Miniprep Kit)
Saturday, July 7th
Quick Change mutagenesis to insert Ala in front of the Strep-tag II in pYES2_RFC25 MCS
Investigator: Ingmar
Aim of the experiment: Generation of an RFC 25 compatible version of the pYes2 Vector.
- As the sequencing of the first attempt to introduce Ala in front of the Strep-tag II did not show a successfull insertion of Ala, the quickchange mutagense was performed once again with a modified setup. Presumably a formation of primer dimers was responsable for the experiment's results. Therefore the second PCR was operated in two steps as shown below.
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P44 template |
| 0.5 µl | 1:10 dilution of O54 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P44 template |
| 0.5 µl | 1:10 dilution of O55 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 6 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Transformation of E.coli XL1 blue with ADH1 promoter, ADH1 terminator and TEF2 promoter
Investigator: Georg
Monday, July 9th
Phycocyanobilin (PCB) extraction from dried Spirulina platensis powder (part 1/4)
Investigator: Jeff, Alois, Martin
Aim of the experiment: Phycocyanobilin (PCB) is a cofactor neeeded for the funtion of phytochrome B. Phycocyanobilin is covalently bound to Cys457 of phytochrome B. Saccharomyces cerevisiae does not contain endogenous PCB. For proof of concept PCB should be added to the medium. In the following experiment, PCB is extracted from dried Spirulina platensis powder.
Operational sequence:
- 50 g of Spirulina platensis powder was (from concept-vitalprodukte.de) resuspended in 1.5 l of H2O (30 mg/l) in a beaker covered with aluminium foil.
- Stirring for 10 min at RT.
- Green Spirulina suspension was divided in 6 centrifuge bottles, covered in aluminium foil.
- Centrifation at 10500 rpm at 4 °C (SLA-3000 rotor, Thermo Scientific) for 1 h.
- Supernatant was discarded
- 25 ml of MeOH added to each bottle and was heavily shaked to resuspend the pellet for the next cleaning step with MeOH.
- Each bottle with the resuspended pellet were filled with MeOH to a final volume of 250 ml and shaked again to fully resuspend the pellet.
- Centrifugation at 10500 rpm at 4 °C (SLA-3000 rotor, Thermo Scientific) for 10 min.
- Supernatant was discarded.
- The last 4 steps were repeated until the supernatant of the washed pellet was colorless or cyanblue but not green anymore (Regulary, it takes 3 or 4 times). Pellet should be cyanblue now.
- The pellet of the 6 centrifuge tubes was collected in a sole centrifugation tube, covered in aluminium foil tube, by scratching it out with a small spoon.
- The remaining rest of the pellet which cannot be scratched out were resuspended in a small amount of MeOH and were transformed from tube to tube with a interim shaking step.
- This suspension was transferred into the tube with the scratched-out pellet.
- Centrifugation at 10500 rpm at 4 °C (SLA-3000 rotor, Thermo Scientific) for 10 min.
- Supernatant was discarded.
- Finally washed pellet was stored, wrapped in foil overnight for the methanolysis next day.
Tuesday, July 10th
Phycocyanobilin (PCB) extraction from dried Spirulina platensis powder (part 2/4)
Investigator: Jeff, Alois, Martin
Aim of the experiment: Phycocyanobilin (PCB) is a cofactor neeeded for the funtion of phytochrome B. Phycocyanobilin is covalently bound to Cys457 of phytochrome B. Saccharomyces cerevisiae does not contain endogenous PCB. For proof of concept PCB should be added to the medium. In the following experiment, PCB is extracted from dried Spirulina platensis powder.
Operational sequence:
- Washed pellet from the day before was resuspended in 500 ml MeOH.
- Heat suspension in a 500 ml flask in a water bath at 70 – 75 ºC with a condensing coil cooled with water for 5 – 8 hrs.
- After this, the suspension was transferred into a new centrifuge tube and centrifuged at 10500 rpm at 4 °C (SLA-3000 rotor, Thermo Scientific) for 20 min.
- The supernatant was decanted trough a filter paper into new centrifugation tube and stored, wrapped in a aluminium foil, at -20 °C.
- The pellets also was stored, wrapped in a aluminium foil, at -20 °C.
Plating of received E. coli containing biobricks
Investigator: Jeff
Aim of the experiment: The received biobricks were already transformed in E. coli and were in an agar stabs. These E. coli cells were transferred with an inoculation loop on antibiotic selection plates and were incubated over night.
Operational sequence:
- Bacterias containing plasmids with biobricks were transferred with a sterile inoculation loop on antibiotic plates and were incubated at 37 °C overnight.
- The biobricks were: BBa_K207001 in pSB1A2, BBa_K243006 in BBa_K157000, BBa_K300001 in K300007 (part for other subproject), BBa_K268000 in pSB6A0 (part for other subproject), BBa_K365005 RFC25 in pSB1C3, BBa_K365000 in pSB1C3, BBa_K207000 in pSB3K3, BBa_K165031 in pSB1AK3
Wednesday, July 11th
Phycocyanobilin (PCB) extraction from dried Spirulina platensis powder (part 3/4)
Investigator: Jeff, Alois, Martin
Aim of the experiment: Phycocyanobilin (PCB) is a cofactor neeeded for the funtion of phytochrome B. Phycocyanobilin is covalently bound to Cys457 of phytochrome B. Saccharomyces cerevisiae does not contain endogenous PCB. For proof of concept PCB should be added to the medium. In the following experiment, PCB is extracted from dried Spirulina platensis powder.
Operational sequence:
- The pellet after the first methanolysis of the day before was undergone a second methanolysis step to ensure high efficiacy of PCB extraction. Operational sequence was performed like the first methanolysis including the centrifugation and filtering step. The pellet after the second methanolysis should be more colorless and the filtered supernatant was freezed, like the one from the first methanolysis, at -20 °C, wrapped in aluminium foil
Transformation of E. coli XL1-Blue with pGADT7 and pGBKT7 plasmid for Yeast-two-hybrid (Y2H)
Investigator: Jeff
Aim of the experiment: pGADT7 and pGBKT7 are plasmids containing the transcriptional activation domain or the DNA binding domain of the transcription activator Gal4. pGADT7 contains the transcriptional activation domain of gal4; we want to clone the the first 100 residues of Pif3 (BBa_K365000) into this plasmid. As a result we have a fusion contruct of Gal4 and Pif3, which is nescassary for the light-switchable promoter system. pGBKT7 is for backup, if the ordered biobricks are not working.
Operational sequence:
- E. coli XL1-Blue are transformed with pGADT7 and pGBKT7 seperately after standard protocol of our lab.
Introducing new Saccharomyces cerevisiae strain (Y190 strain) from Schwab lab for Y2H
Investigator: Jeff
Aim of the experiment: The Y190 Saccharomyces cerevisiae is a special strain for Yeast-two-hybrid. This strain carries a Gal4 and Gal80 deletion to higher the signal/noise-ratio of protein-protein interactions. Reporter for protein-protein interactions are HIS3, lacZ and MEL1 and are encoded in the genomic DNA. Transformation markers are trp1, leu2 and cyhR2 and are encoded on the transformation plasmids.
Operational sequence:
- The freshly plated Y190 cells are transferred with a inoculation loop from the original plate on a new YPD agar plate.
- After 2 days the the plate was put in 4 °C.
Repetition of PCR of PAL
Investigator: Katrin, Daniela
Reaction batch
| volume | reagent |
| 5 µl | 10x Pfu Ultra II buffer |
| 4 µl | dNTP's (each 2.5 mM) |
| 0.5 µl | Pfu Ultra II (2.5 U/µL) |
| 5 µl | 1:10 dilution of used forward primers (10µM) |
| 5 µl | 1:10 dilution of used reversed primers (10µM) |
| 1 µl | DNA (pKS2µHyg-PAL-4CL-CHS 50 ng/µL) |
| 29.5 µL | bidest. sterile Water |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 2 min (and adding Pfu Ultra after 2 min) |
| 2 | 30 | 95°C | 30 sec |
| 52°C | 1 min | ||
| 72°C | 2.5 min | ||
| 3 | 72°C | 5 min |
PCR purification
- Purification was done using QIAquick PCR Purification Kit (250)
Analytical gelelektrophoresis of PCR-Products of PAL
-> PCR was not successful, no band at 2,1 kb (picture follows)
Preparative Digest of pYES_iGEM
Investigator: Katrin, Daniela
Digestion of P50 with Xbal and NgOMIV
| volume | reagent |
| 10 µl | P50 (concentration: 264.5 ng/µl) |
| 2.5 µl | NEB4 |
| 2.5 µl | 10x BSA |
| 1 µl | XbaI (20 U/µl) |
| 2 µl | NgOMIV (10U/µl) |
| 7 µl | ddH2O |
Incubation: 37 °C, 3 h
Digestion of P50 with Xbal and PstI
| volume | reagent |
| 10 µl | P50 (concentration: 264.5 ng/µl) |
| 5 µl | Tango buffer |
| 2 µl | XbaI |
| 3 µl | PstI |
| 7.5 µl | ddH2O |
Incubation: 37 °C, 3 h
DNA preparative gel electrophoresis
- gel: 1% with LMP-agarose
- band 1: P50 digested with XbaI and PstI
- band 2: P50 digested with XbaI and NgOMIV
- 70 V, 90 min
Gelextration
- cut the bands
- QIAquick Gel Extractrion Kit was used
- step 6 was left out
- step 9: 30µl buffer, 4 min incubation
Ligation of digested P50 with digested PCR-products of PCR 15-20 (4CL, CHS and OMT)
Investigator: Katrin, Daniela
Concentration (Nano Drop:
4CL+ (PCR 15) = 40.3 ng/µl
4CL- (PCR 16) = 37.3 ng/µl
CHS+ (PCR 17) = 46.1 ng/µl
CHS- (PCR 18) = 51.2 ng/µl
OMT+ (PCR 19) = 22.3 ng/µl
OMT- (PCR 20) = 16.7 ng/µl
P50 digested with XbaI and PstI = 23.9 ng/µl (the digested Plasmid has the number P71)
P50 digested with XbaI and NgOMIV = 28.4 ng/µl (the digested Plasmid has the number P72)
- required volumes were calculated using Lab Tools
4CL+ (PCR 15) with P50 digested with XbaI and PstI
| volume | reagent |
| 5.27 µl | P50 digested |
| 2.73 µl | 4CL+ (PCR 15) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
4CL- (PCR 16) with P50 digested with XbaI and PstI
| volume | reagent |
| 5.13 µl | P50 digested |
| 2.87 µl | 4CL- (PCR 16) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
CHS+ (PCR 17) with P50 digested with XbaI and NgOMIV
| volume | reagent |
| 5.84 µl | P50 digested |
| 2.16 µl | CHS+ (PCR 17) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
CHS- (PCR 18) with P50 digested with XbaI and NgOMIV
| volume | reagent |
| 6 µl | P50 digested |
| 2 µl | CHS- (PCR 18) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
OMT+ (PCR 19) with P50 digested with XbaI and NgOMIV
| volume | reagent |
| 4.45 µl | P50 digested |
| 3.55 µl | OMT+ (PCR 19) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
OMT- (PCR 20) with P50 digested with XbaI and NgOMIV
| volume | reagent |
| 3.87 µl | P50 digested |
| 4,13 µl | OMT- (PCR 20) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
Negative control
| volume | reagent |
| 5 µl | P50 digested with XbaI and PstI or NgOMIV |
| 3 µl | ddH2O |
| 1 µl | T4 DNA-ligase |
| 1 µl | T4-ligase buffer (10x) |
- water bath 16 °C
Thursday, July 12th
Genextraction and analytical digestion of CYC1-terminator, TEF1-promoter, and PGK1-Promoter from pSB1C3 and pSB1A2
Investigator: Georg
- Genes from overnight cultures were extracted, using the Quiaprep gene extraction Kit. Extracted Plasmids then were analytically digested with XbaI and PstI (Fermentas).
Analytical digestion XbaI, PstI
| Chemical | Volume |
| XbaI | 2,5 µl |
| PstI | 2,5 µl |
| 10xBuffer Tango | 2 µl |
| Plasmid-DNA | 2,5 µl |
| ddH2O | 11,5 µl |
- Analytical gel was run (1% Agarose).
Phycocyanobilin (PCB) extraction from dried Spirulina platensis powder (part 4/4)
Investigator: Martin, Jeff, Alois
Aim of the experiment: Phycocyanobilin (PCB) is a cofactor neeeded for the funtion of phytochrome B. Phycocyanobilin is covalently bound to Cys457 of phytochrome B. Saccharomyces cerevisiae does not contain endogenous PCB. For proof of concept PCB should be added to the medium. In the following experiment, PCB is extracted from dried Spirulina platensis powder.
Operational sequence:
- The supernatants (~1l in total) of the two previous experiments were pooled and put into a rotary evaporater in order to acquire a concentrate of approximately 100ml.
- The settings of the rotary evaporater: ~160 millibars, waterbath temperature of 25-30°C
- Furthermore, there were measures to be taken to protect the solution from direct sunlight: blinds down, aluminum foil wrapped around the waterbath
- Afterwards we transfered the Methanol solution into a separating funnel and added another 100ml of aqua dest..
- By means of adding chloroform we created two phases. The lower (chloroform) phase with the solved Phycocyanobilin was separated and put into another rotary evaporater flask. This step was repeated three times in order to get all the Phycocanobilin into the next step
- Then the chloroform was completely removed in the rotary evaporater (same settings as before), the pure (?) Phycocyanobilin in the flask was solved in 60 ml DMSO, transfered into a new flask and frozen at -20 °C.
Picking of transformated (pGADT7 and pGBKT7) E. coli cells on antibiotic selection plates
Investigator: Jeff
Aim of the experiment: pGADT7 and pGBKT7 are plasmids containing the transcriptional activation domain or the DNA binding domain of the transcription activator Gal4. pGADT7 contains the transcriptional activation domain of gal4; we want to clone the the first 100 residues of Pif3 (BBa_K365000) into this plasmid. As a result we have a fusion contruct of Gal4 and Pif3, which is nescassary for the light-switchable promoter system. pGBKT7 is for backup, if the ordered biobricks are not working.
Operational sequence:
- A E. coli colony was picked for each plasmid and was transferred in a tube containing 4 ml of LB-medium containing antibiotics (Amp for pGADT7 and Kan for pGBKT7).
- Overnight culture at 37 °C in a cell-culture shaker.
Miniprep of transformated E. coli from overnight culture (8 plasmids containing biobricks)
Investigator: Andrea
Aim of the experiment: Miniprep of transformated E. coli from overnight culture to get the plasmids with biobricks
NanoDrop Measure
| Plasmid | Concentration |
| P73 | 41.6 ng/µl |
| P74 | 85.2 ng/µl |
| P75 | 72.4 ng/µl |
| P76 | 254.3 ng/µl |
| P77 | 74.4 ng/µl |
| P78 | 50.0 ng/µl |
| P79 | 46.2 ng/µl |
| P80 | 116.7 ng/µl |
| P81 | 77.1 ng/µl |
| P82 | 65.5 ng/µl |
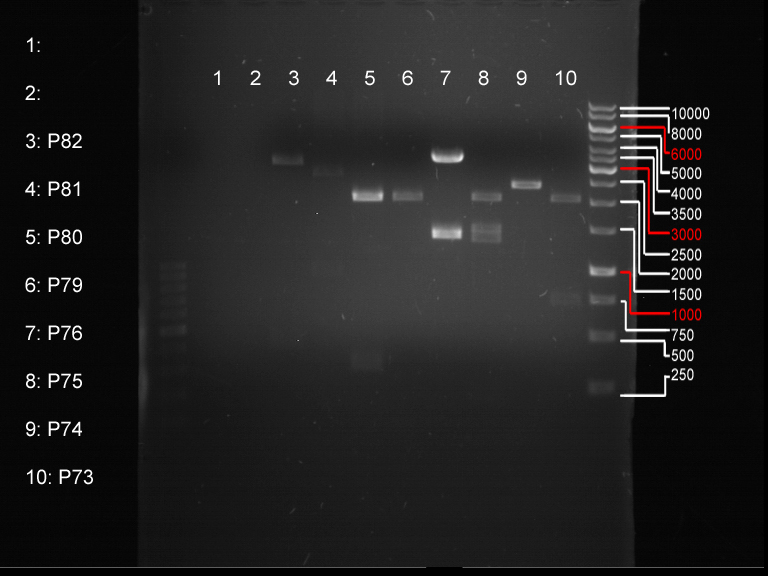
Analytical digestion and gelelectrophoresis of biobricks (8 plasmids containing biobricks)
Investigator: Jeff
Aim of the experiment: Checking whether the biobricks are part of the plasmid-backbone.
Operational sequence:
- Reaction batch for each plasmid:
| Reagent | Volume in µl |
| Tango Buffer 10x | 4 µl |
| XbaI (Fermentas) | 0.25 µl |
| PstI (Fermentas) | 0.5 µl |
| Plasmid DNA | 2.5 µl |
| ddH2O | 12.75 µl |
| TOTAL | 20 µl |
- Incubation at 37 °C for 1 h 45 min.
- Analytical gelelectrophoresis at 90 V for 1 h.
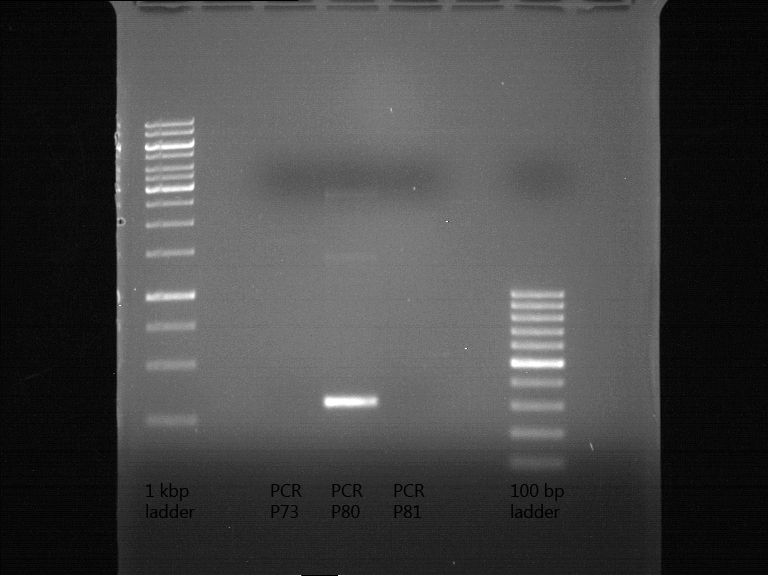
- Order of gel-pockets:
| 1 kbp ladder | P73 | P74 | P75 | P76 | P79 | P80 | P81 | P82 | 100 bp ladder |
| BAD | OK | OK | OK | OK | OK | BAD | OK |
- P73 and P81, both parts from Havard university are bad. To exclude errors, one should pick another colony for each plasmid and do again the analytical digestion and gelelectrophoresis after the miniprep.
Transformation of Ligationproducts of pYES2 + OMT, 4Cl and CHS in E.coli
Investigator: Mary
- adding 5µl ligation product in 100µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (Withouth antibiotica) and incubate at 37°C, 30 min, 180 rpm
- plate on agar with Ampicillin over night
Repetition of PCR of PAL
Investigator: Daniela
Aim of the experiment: Repetition of PCR of PAL (so far not successful) with the use of different polymerase and try of 3 temperatures
only with PAL+, 3 different temperatures (see cycling parameters) and one batch at 52.5 °C with DMSO
Reaction batch
| volume | reagent |
| 10 µl | 5x Herculase II reaction buffer |
| 0,5 µl | dNTP Mix (each dNTP 2.5 mM) |
| 1 µl | Herculase II fusion DNA Polymerase |
| 1,25 µl | 1:10 dilution of used forward primers (O22) |
| 1,25 µl | 1:10 dilution of used reversed primers (O59) |
| 1 µl | 1:10 dilution of DNA (P19=pKS2µHyg-PAL-4CL-CHS 50 ng/µL) -> 5 ng/µL |
| 35 µL | ddH2O |
Reaction batch with DMSO
| volume | reagent |
| 10 µl | 5x Herculase II reaction buffer |
| 0,5 µl | dNTP Mix (each dNTP 2.5 mM) |
| 1 µl | Herculase II fusion DNA Polymerase |
| 1,25 µl | 1:10 dilution of used forward primers (O22) |
| 1,25 µl | 1:10 dilution of used reversed primers (O59) |
| 1 µl | 1:10 dilution of DNA (pKS2µHyg-PAL-4CL-CHS 50 ng/µL) -> 5 ng/µL |
| 1,5 µl | DMSO (3% des Ansatzes) |
| 33.5 µL | ddH2O |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 2 min |
| 2 | 30 | 95°C | 30 sec |
| 52.5°C | 30 sec | ||
| 72°C | 4 min | ||
| 3 | 72°C | 3 min |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 2 min |
| 2 | 30 | 95°C | 30 sec |
| 45°C | 30 sec | ||
| 72°C | 4 min | ||
| 3 | 72°C | 3 min |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 2 min |
| 2 | 30 | 95°C | 30 sec |
| 56.2°C | 30 sec | ||
| 72°C | 4 min | ||
| 3 | 72°C | 3 min |
Analytical gel electrophoresis
- no bands at all (picture follows)
- possibly due to low concentration of P19 (5 ng/µl)
Ligation of P93 (pSB1C3 digested with XbaI and AgeI) with digested PCR-products of PCR 17-20 (CHS and OMT)
Investigator: Daniela
Concentration (Nano Drop:
CHS+ (PCR 17) = 46.1 ng/µl
CHS- (PCR 18) = 51.2 ng/µl
OMT+ (PCR 19) = 22.3 ng/µl
OMT- (PCR 20) = 16.7 ng/µl
P93 (digested with XbaI and AgeI) = 4.8 ng/µl
CHS+ (PCR17) with P93 digested with XbaI and AgeI
| volume | reagent |
| 7.04 µl | P93 |
| 0.96 µl | CHS+ (PCR 17) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
CHS- (PCR 18) with P93 digested with XbaI and AgeI
| volume | reagent |
| 7.13 µl | P93 |
| 0.87 µl | CHS- (PCR18) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
OMT+ (PCR19) with P93 digested with XbaI and AgeI
| volume | reagent |
| 6.19 µl | P93 |
| 1.81 µl | OMT+ (PCR19) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
OMT- (PCR20) with P50 digested with XbaI and AgeI
| volume | reagent |
| 5.75 µl | P93 |
| 2.25 µl | OMT- (PCR20) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
Negative control
| volume | reagent |
| 5 µl | P93 |
| 3 µl | ddH2O |
| 1 µl | T4 DNA-ligase |
| 1 µl | T4-ligase buffer (10x) |
- water bath 16 °C
- products were named P94-P97
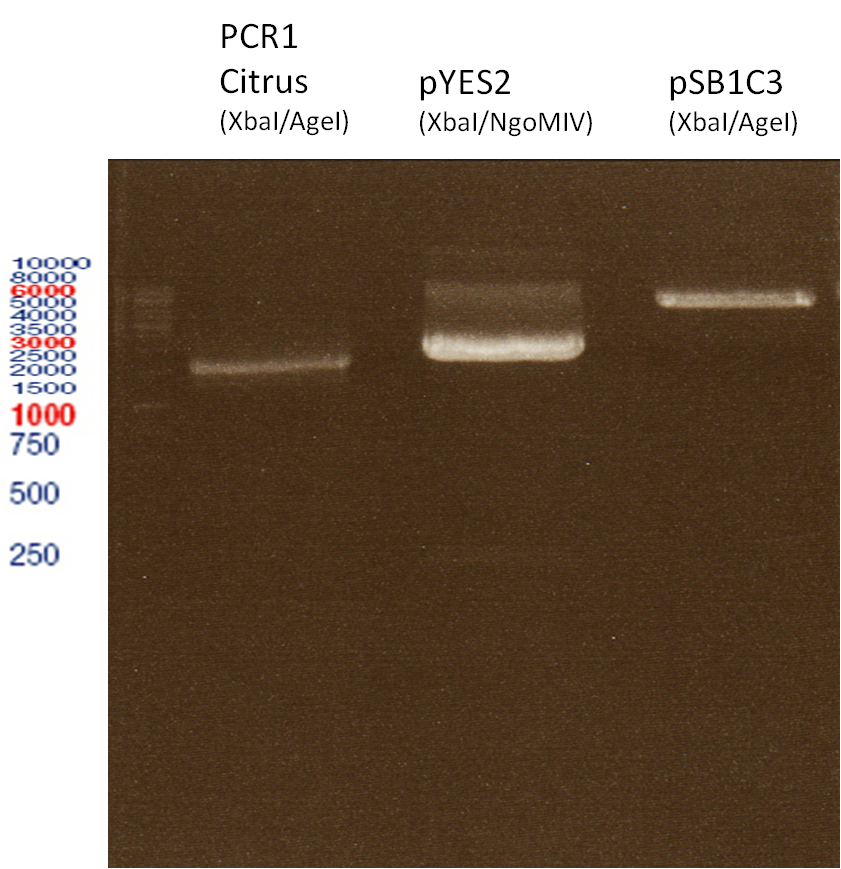
Preparative digest of PCR1-PCR7 and P50
Investigator: Andrea
Digestion of P50 with XbaI and NgoMIV
| volume | reagent |
| 20 µl | P50 |
| 4 µl | NEB Buffer |
| 0.4 µl | BSA |
| 1 µl | XbaI (10 U/µl) |
| 2 µl | NgoMIV (10 U/µl) |
| 27.4 µl | ddH2O |
Digestion of PCR1-PCR7 with XbaI and AgeI
| volume | reagent |
| 25 µl | PCR-Product |
| 5 µl | NEB Buffer |
| 0.5 µl | BSA |
| 1 µl | XbaI (10 U/µl) |
| 1 µl | AgeI (10 U/µl) |
| 32.5 µl | ddH2O |
Incubation: 37 °C, 3 h
Preperative gel electrophoresis
Investigator: Andrea
Aim of the experiment: Analytical gel electrophoresis of products from restriction digest of plasmids PCR1 - PCR7 and PCR9
Ligation of digested PCR1-PCR7 (digested with XbaI and AgeI) with pSB1C3 (digested with XbaI and AgeI) and with pYESnew (digested with XbaI and NgoMIV)
Investigator: Andrea
Concentration (Nano Drop:
LIMS Citrus (PCR 1) = 13.4 ng/µl
LIMS Citrus (PCR 2) = 10.3 ng/µl
LIMS Lavendula (PCR 3) = 2.2 ng/µl
LIMS Lavendula (PCR 3) = 9.9 ng/µl
LIMS Lavendula (PCR 3) = 9.6 ng/µl
LIMS Lavendula (PCR 3) = 9.8 ng/µl
LIMS Lavendula (PCR 3) = 12.3 ng/µl
P50 (digested with XbaI and NgoMIV) = 10.3 ng/µl
P93 (digested with XbaI and AgeI) = 4 ng/µl
PCR1 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.89µl | P50 |
| 3.11 µl | PCR 1 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR2 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.38 µl | P50 |
| 3.62 µl | PCR2 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR3 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 1.64 µl | P50 |
| 6.36 µl | PCR3 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR4 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.3 µl | P50 |
| 3.7 µl | PCR4 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR5 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.26 µl | P50 |
| 3.74 µl | PCR5 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR6 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.26 µl | P50 |
| 3.74 µl | PCR6 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR7 with P50 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 7.83 µl | P50 |
| 3.27 µl | PCR7 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
Negative control
| volume | reagent |
| 5 µl | P50 |
| 3 µl | ddH2O |
| 1 µl | T4 DNA-ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR1 with P93 digested with XbaI and AgeI
| volume | reagent |
| 4.89 µl | P93 |
| 3.11 µl | PCR1 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR2 with P93 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.38 µl | P93 |
| 3.62 µl | PCR2 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR3 with P93 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 1.52 µl | P93 |
| 6.48 µl | PCR3 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR4 with P93 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.3 µl | P93 |
| 3.7 µl | PCR4 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR5 with P93 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.26 µl | P93 |
| 3.74 µl | PCR5 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR6 with P93 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.26 µl | P93 |
| 3.74 µl | PCR6 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
PCR7 with P93 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 4.72 µl | P93 |
| 3.28 µl | PCR7 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
- water bath 16 °C
- products were named P100-P115
Friday, July 13th
PCR of P73, P80 and P81 to add RFC25 pre- and suffix
Investigator: Jeff, Saskia
Aim of the experiment: Introducing RFC25 pre- and suffix into parts of P73, P80 and P81.
Operational sequence:
- PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer (For P73: O46 (TUM12-PhyBGal4bd-fw); for P80: O31 (TUM12-LSPS-fw); for P81: O50 (TUM12-Phyb-fw)) |
| 1 µl | 10 µM Reverse Primer (For P73: O47 (TUM12-PhyBGal4bd-rv); for P80: O32 (TUM12-LSPS-rv ); for P81: O51 (TUM12-Phyb-rv)) |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA (P80 or P73 or P81) |
| 35.75 µL | ELGA Water |
| =50 µL | TOTAL |
- The gradient PCR program was performed after following scheme with following conditions (Tm=58 ΔG=5 °C; P73 in row 9(=61.1 °C); P80 in row 1 (=53.0 °C); P81 in row 7(=58.4 °C):
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| Tm=58 °C; ΔG=5 °C | 150 s | |
| 68 °C | 60 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | overnight |
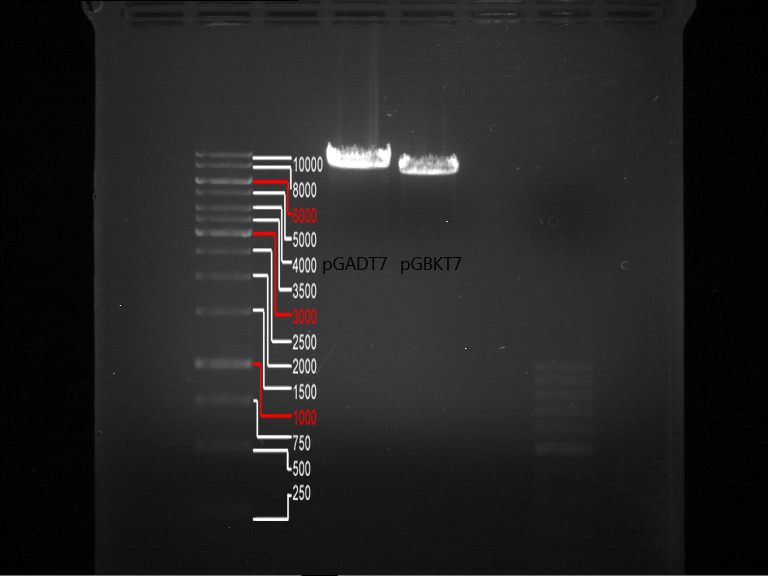
Miniprep, analcytical digestion and gelelectrophoresis of pGADT7 and pGBKT7
Investigator: Jeff, Saskia
Aim of the experiment:Miniprep, analcytical digestion and gelelectrophoresis of pGADT7 (tube P98, EcoRI and BamHI)and pGBKT7 (tube P99, BamHI and PstI)
Operational sequence:
- Operated after standard protocol of the lab for analytical digestion and gelelectrophoresis.
- Gel OK, like expected
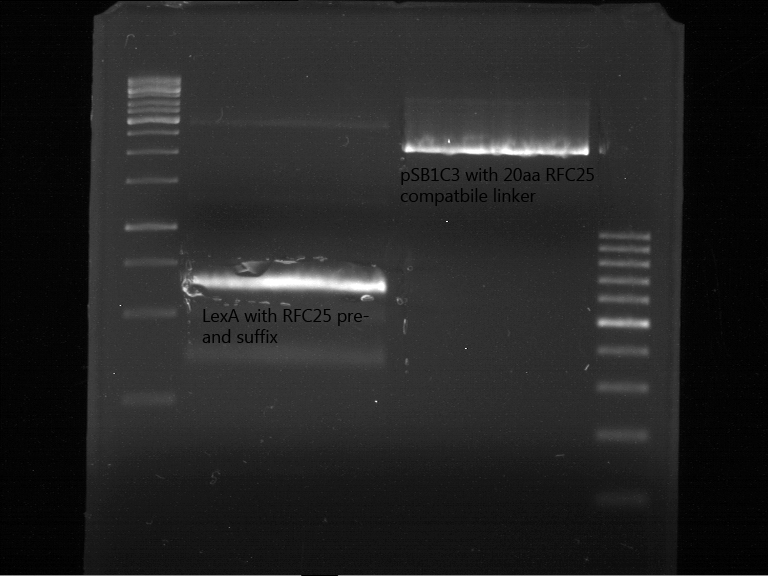
Preperative digestion and gelelectrophoresis of P79 and O56???
Investigator: Jeff, Saskia, Georg
Aim of the experiment: Preperativ gelelectrophoresis and gel of digested PCR product of LexA (NgoMIV+PstI) and pSB1C3 containing the RFC25 compatible 20aa linker with RFC25 pre- and suffix (AgeI and PstI).
Operational sequence:
- Operated after standard protocol of the lab for preperative digestion and gelelectrophoresis.
- Gel OK, like expected.
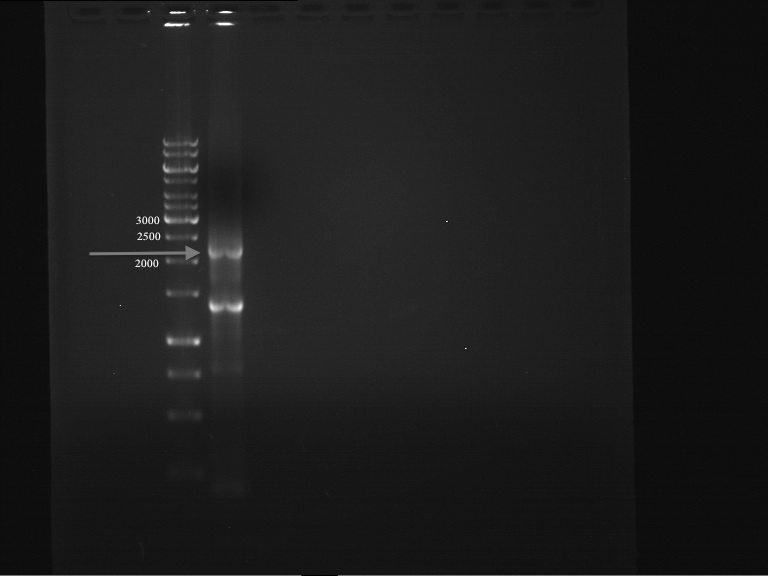
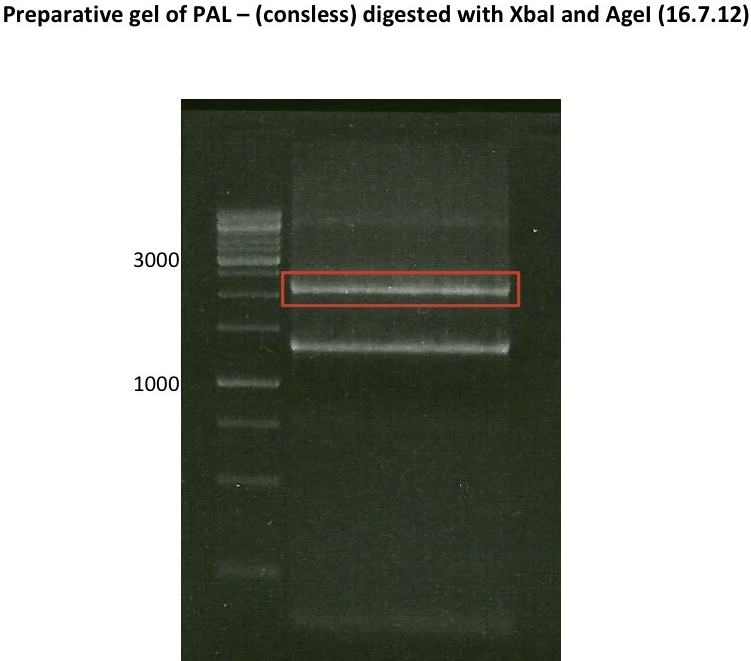
Preparative digest of P19
Investigator:Daniela, Ingmar
Digestion of P19 with ApaI
| volume | reagent |
| 20 µl | P19 (concentration: 53.8 ng/µl) |
| 4 µl | Buffer B |
| 2 µl | ApaI (10 U/µl) |
| 14 µl | ddH2O |
Incubation: 37 °C, 3 h
DNA preparative gel electrophoresis
- gel: 1% with LMP-agarose
- band 1: P50 digested with XbaI and PstI
- band 2: P50 digested with XbaI and NgOMIV
- 70 V, 90 min
Gelextration
- cut the bands
- QIAquick Gel Extractrion Kit was used
- step 6 was left out
- step 9: 30µl buffer, 4 min incubation
Gradient PCR of PAL to optimize the primer annealing temperature
Investigator: Ingmar
Aim of the experiment: As all PCR experiments to amplify the PAL gene contained in P19 failed, we decided to run a gradient PCR to optimize the primer annealing temperature.
PCR
Reaction batch
| volume | reagent |
| 1 µl | 10x Pfu Ultra II buffer |
| 0.5 µl | Plasmid P19 template |
| 0.5 µl | 1:10 dilution of O15 (10 pmol/µL) |
| 0.5 µl | 1:10 dilution of O59 (10 pmol/µL) |
| 0.25 µl | dNTP mix |
| 7 µl | ddH2O |
| 0.25 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 29 | 95°C | 30 sec |
| gadient 33-47.5 °C | 1 min | ||
| 68°C | 3 min | ||
| 3 | 1 | 68°C | 5 min |
| 4 | 1 | 4°C | infinity |
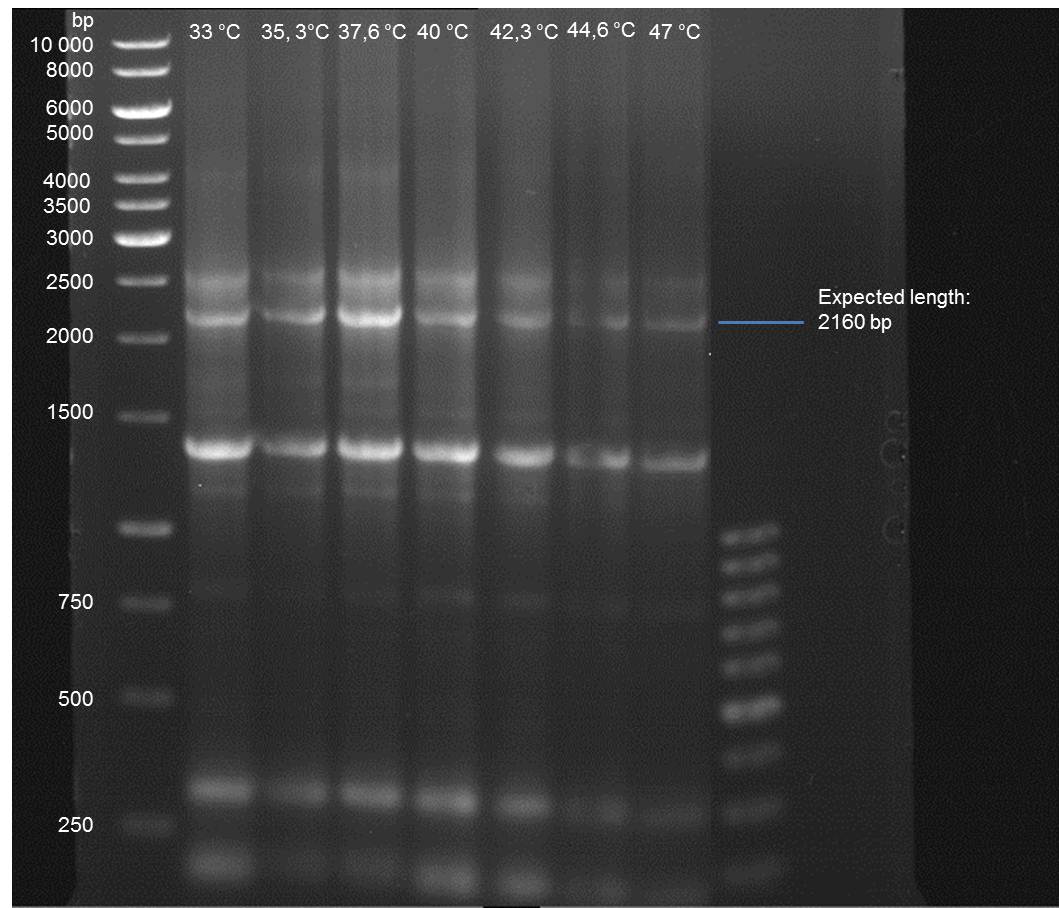
- Verification of the PCR by agarose gel electrophoresis:
10 µl of each PCR tube was mixed with 2 µl 6x DNA loading buffer and loaded into the gel. The separation process lasted 1h at 90 V in a 1% agarose gel.
- A bond at the expected length of 2160 bp appears at 47 °C. Therefore a PCR using this primer annealing temperature will done on sunday.
Transformation of P94 - P97 (ligationproducts of pSB1C3 with CHS and OMT respectively) in E.coli
Investigator: Daniela
- adding 5µl ligation product (P94-P97) in 100µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (Withouth antibiotica) and incubate at 37°C, 30 min, 180 rpm
- plate on agar with Ampicillin over night
wrong antibiotica was used!!! Repetition follows!!! Nevertheless, some colonies could be observed.
Ligation of LIMS in pYESnew and pSB1C3
Investigator:Andrea
Transformation
- adding 5µl ligation product (PCR1-PCR7) in 100µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 30 min, 180 rpm
- plate on agar with Ampicillin (pYESnew) or Chloramphenicol (pSB1C3) over night
Preparation of YPD medium
Investigator: A lot of Alois, Martin
Manual for YPD production
Yeast Extract Peptone Dextrose Medium (1 liter)
- 1% yeast extract
- 2% peptone
- 2% dextrose (D-glucose)
1. Dissolve the following in 1000 ml of water:
- 10 g yeast extract
- 20 g peptone
- 20 g dextrose (see note below if making plates)
2. Autoclave for 20 minutes on liquid cycle.
3. Store medium at room temperature. The shelf life is approximately one to two months.
Sunday, July 15th
PCR purification of PCR products of P73, P80 and P81 and analytical gelelectrophoresis
Investigator: Jeff
Aim of the experiment: PCR was performed to introduce restriction sites to the gene of interest. MfeI and BamHI for P73 and RFC25 pre- and suffix for P80 and P81. The aim is to contruct fusion proteins with the aid of these restriction sites.
Operational sequence:
- PCR cleaning with Qiagen PCR purification kit after manufacturer's protocol.
- Analytical gelelectrophoresis (1% agarose) at 90 for 60&min.
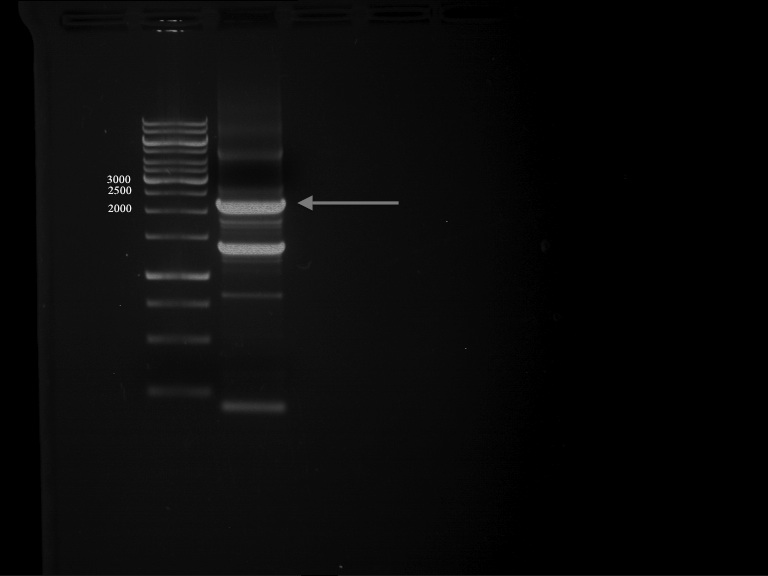
PCR of PAL consless using the optimized primer annealing temperature of 47 °C
Investigator: Ingmar
Aim of the experiment: Amplification of the PAL gene contained in P19.
PCR
Reaction batch
| volume | reagent |
| 5 µl | 10x Pfu Ultra II buffer |
| 2.5 µl | Plasmid P19 template |
| 2.5 µl | 1:10 dilution of O15 (10 pmol/µL) |
| 2.5 µl | 1:10 dilution of O59 (10 pmol/µL) |
| 1.25 µl | dNTP mix |
| 35 µl | ddH2O |
| 1.25 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 29 | 95°C | 30 sec |
| 47 °C | 1 min | ||
| 68°C | 3 min | ||
| 3 | 1 | 68°C | 5 min |
| 4 | 1 | 4°C | infinity |
PCR successful - Band at about 2,1 kb :
Monday, July 16th
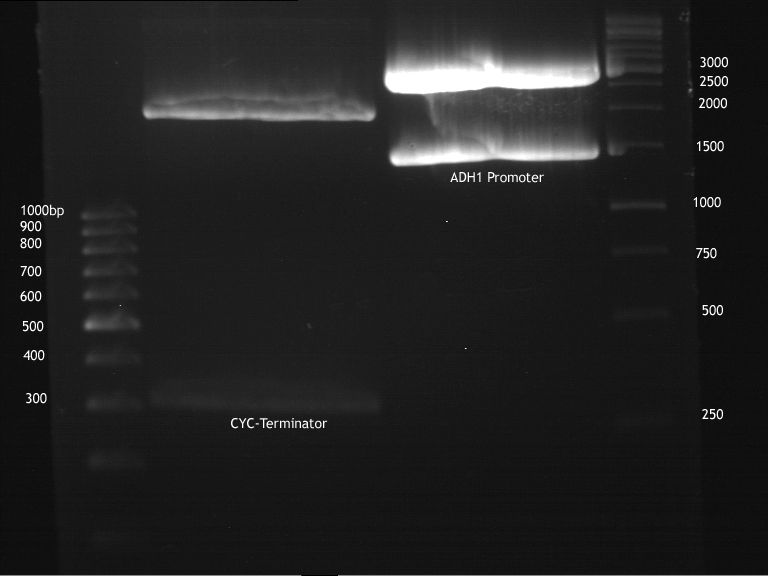
Preparative digestion and gelelectrophoresis of P86 and P55
Investigator:Georg Aim of the experiment: Digestion of CYC1-Terminator, ADH1-Promoter for ligation
- Preparative digestion after manufacturer's advice (NEB) with 20 u PstI and 20 u XbaI in 1x Tango buffer with 25 µl DNA and 4 µl NEB4 10x buffer and 0,4 µl 100x BSA. Water was added to a volume of 40 µl. Restriction time was 3 hours at 37 °C; 3 hours.
- Preparative gelelectrophoresis after laboratory's standart protocol. (70 V, 90 min)
- Gel was stored at -20 °C.
Preperative digestion and gelelectrophoresis of P98 and PCR26
Investigator: Jeff, Georg
Aim of the experiment: Construction of Gal4AD-Pif3, a part of the light-switchable promoter system.
Operational sequence:
- Preperative digestion after manufacturer's (Fermentas) advise for double-digestion for EcoRI+BamHI and MfeI(MunI)+BamHI; EcoRI+BamHI: Buffer 2X Tango™, 2-fold excess of BamHI; MfeI(MunI)+BamHI: Buffer G.
- Preperative gelelectrohphoresis after laboratory's standard protocol. (70 V, 90 min)
Plating of received E. coli containing biobrick BBa_K165055
Investigator: Jeff
Aim of the experiment: The received biobrick BBa_K165055 (LexA binding sites + mCYC + Kozak + YFPx2 + ADH1 terminator) were already transformed in E. coli and were in an agar stabs. These E. coli cells were transferred with an inoculation loop on antibiotic selection plates and were incubated over night.
Operational sequence:
- Bacterias containing plasmids with biobricks were transferred with a sterile inoculation loop on antibiotic plates and were incubated at 37 °C overnight.
- The biobricks was BBa_K165055 in BBa_J63009 plasmid (AmpR, low copy plasmid)
Preparatory culture of S. cerevisiae
Investigator: Alois, Martin
Aim of the experiment: In order to have sufficient viable cells to inoculate an experimental culture in which we want to compare the growth rate of s. cerevisiae with a strain of brewing yeast (34/70 s. pastorianus weihenstephan) we aim to establish a preparatory over night culture.
Operational sequence: A sample of -80°C stored s. cerevisiae (glycerol stock) is used to inoculate 20ml of YPD-medium. This is shaken overnight at 30°C.
Miniprep of plasmids containing lavendula limonene synthase/citrus limonene synthase
Investigator: Lara
Aim of the experiment: Extract plasmids (pYESnew and pBS1C3) that contain limonene synthase/citrus limonene synthase.
Miniprep with Qiagen Kit; Plasmids p116-p122. Restriction digest with Xba1 and Pst1.
Plasmid DNA concentrations:
p116: 285 ng/µl
p117: 337 ng/µl
p118: 520 ng/µl
p119: 370 ng/µl
p120: 320 ng/µl
p121: 320 ng/µl
p122: 335 ng/µl
Restriction digest of p116-p122
Investigator: Lara
Aim of the experiment: Analytical restriction digest of plasmids p116, p117, p118, p119, p120, p121, p122.
| volume | reagent |
| 2 µl | DNA |
| 2 µl | Buffer 4 |
| 0,25 µl | Xba1 |
| 0,25 µl | Pst1-HF |
| 15,3 µl | ddH2O |
Incubation: 37 °C; 1,5 h
Restriction digest with Xba1 and Pst1-HF.
Analytical gel electrophoresis
Investigator: Lara
Aim of the experiment: Analytical gel electrophoresis of products from restriction digest of plasmids p116, p117, p118, p119, p120, p121, p122.
p116: PCR1/pYES
p117: PCR2/pYES
p118: PCR4/pYES
p119: PCR4/pSB1C3
p120: PCR5/pYES
p121: PCR6/pYES
p122: PCR7/pYES
Midiprep of pSB1C3 (RFC25-compatible)
Investigator:Mary
Aim of the experiment: Get a lot of pSB1C3 for further experiments
Midipreparation of the pellet from over night cultures (E.coli, Transformed with pSB1C3 - RFC25; name in registry: K365005)
was done with the Midiprep Kit from Qiagen
Result was named as P123 and had the concentration: 469,5 ng/µl (100µl at all)
PCR-Products of PAL (consless): digestion, extraction and ligation
investigator: Daniela, Mary
Aim of the experiment: digestion of PCR-Products to gain the correct cutting sites for ligation in pYES2 and pSB1C3 afterwards
Purification of PCR-Products with PCR-Purification Kit
- digestion with Xba1 and HF-Age1 (both NEB)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer NEB4 |
| 0.5µl | BSA |
| 1µl | Xba1 (NEB; 20u/µl) |
| 1µl | HF-Age1 (NEB; 20u/µl) |
| 17.5µl | bidest. sterile H2O |
digestion took 3h at 37°C
- extraction from preparative gel:
PCR of PAL+cons using the optimized primer annealing temperature of 47 °C
Investigator: Daniela
Aim of the experiment: Amplification of the PAL cons gene contained in P19.
PCR
Reaction batch
| volume | reagent |
| 5 µl | 10x Pfu Ultra II buffer |
| 2.5 µl | Plasmid P19 template |
| 2.5 µl | 1:10 dilution of O22 (10 pmol/µL) |
| 2.5 µl | 1:10 dilution of O59 (10 pmol/µL) |
| 1.25 µl | dNTP mix |
| 35 µl | ddH2O |
| 1.25 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 29 | 95°C | 30 sec |
| 47 °C | 1 min | ||
| 68°C | 3 min | ||
| 3 | 1 | 68°C | 5 min |
| 4 | 1 | 4°C | infinity |
PCR was succesful, band at 2,1kb:
Transformation of P94 - P97 (ligationproducts of pSB1C3 with CHS and OMT respectively) in E.coli
Investigator: Daniela
- adding 5µl ligation product (P94-P97) in 100µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 45 min, 180 rpm
- plate on agar with chloramphenicol over night
-> were succesfull: some colonies were grown
Tuesday, July 17th
Picking of transformed E. coli containing biobrick BBa_K165055
Preparative digest of P123 (pSB1C3 RFC25)
Investigator: Mary, Daniela
Digestion of P123 with Xbal/PstI and XbaI/AgeI
| volume | reagent |
| 23.1 µl | ddH2O |
| 4 µl | NEB4 |
| 0.4 µl | 10x BSA |
| 1 µl | XbaI (20 U/µl) |
| 1.5 µl | PstI (10U/µl) |
| 10 µl | P123 |
| volume | reagent |
| 23.35 µl | ddH2O |
| 4 µl | NEB4 |
| 0.4 µl | 10x BSA |
| 1 µl | XbaI (20 U/µl) |
| 1.25 µl | AgeI (10U/µl) |
| 10 µl | P123 |
Incubation: 37 °C, 3 h
DNA preparative gel electrophoresis
- gel: 1% with LMP-agarose
- band 1: P123 digested with XbaI and PstI
- band 2: P123 digested with XbaI and AgeI
- 70 V, 90 min
Gelextration
- cut the bands
- QIAquick Gel Extractrion Kit was used
- step 6 was left out
- step 9: 30µl buffer, 4 min incubation
Products were names as follows:
- P123 digested with XbaI and PstI: P132
- P123 digested with XbaI and AgeI: P133
PCR-Products of PAL (with consensussequence): digestion and extraction
investigator: Daniela, Mary
Aim of the experiment: digestion of PCR-Products to gain the correct cutting sites for ligation in pYES2 and pSB1C3 afterwards
Purification of PCR-Products with PCR-Purification Kit
- digestion with Xba1 and HF-Age1 (both NEB)
| volume | reagent |
| 25µl | PCR-product |
| 5µl | Buffer NEB4 |
| 0.5µl | BSA |
| 1µl | Xba1 (NEB; 20u/µl) |
| 1µl | HF-Age1 (NEB; 20u/µl) |
| 17.5µl | bidest. sterile H2O |
digestion took 3h at 37°C
- extraction from preparative gel:
Picking Clones of CHS and OMT in pSB1C3-RFC25
investigator: Daniela, Mary
Aim of the experiment: preculture over night for the miniprep next day
Repetition of ligation of PCR 15-20 (4CL, CHS and OMT) with pYES (P71 and P72 respectively)
Investigator: Mary, Daniela
Concentration (Nano Drop:
4CL+ (PCR 15) = 40.3 ng/µl
4CL- (PCR 16) = 37.3 ng/µl
CHS+ (PCR 17) = 46.1 ng/µl
CHS- (PCR 18) = 51.2 ng/µl
OMT+ (PCR 19) = 22.3 ng/µl
OMT- (PCR 20) = 16.7 ng/µl
P50 digested with XbaI and PstI = 23.9 ng/µl (the digested Plasmid has the number P71)
P50 digested with XbaI and NgOMIV = 28.4 ng/µl (the digested Plasmid has the number P72)
- required volumes were calculated using Lab Tools
4CL+ (PCR 15) with P71
| volume | reagent |
| 5.27 µl | P71 |
| 2.73 µl | 4CL+ (PCR 15) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
4CL- (PCR 16) with P71
| volume | reagent |
| 5.13 µl | P71 |
| 2.87 µl | 4CL- (PCR 16) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
CHS+ (PCR 17) with P72
| volume | reagent |
| 5.84 µl | P72 |
| 2.16 µl | CHS+ (PCR 17) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
CHS- (PCR 18) with P72
| volume | reagent |
| 6 µl | P72 |
| 2 µl | CHS- (PCR 18) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
OMT+ (PCR 19) with P72
| volume | reagent |
| 4.45 µl | P72 |
| 3.55 µl | OMT+ (PCR 19) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
OMT- (PCR 20) with P72
| volume | reagent |
| 3.87 µl | P72 |
| 4,13 µl | OMT- (PCR 20) |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
Negative control
| volume | reagent |
| 5 µl | P71 or P72 |
| 3 µl | ddH2O |
| 1 µl | T4 DNA-ligase |
| 1 µl | T4-ligase buffer (10x) |
- water bath 16 °C
Transformation of ligationproducts of 4CL, OMT and CHS respectively in pYES in E.coli
Investigator: Mary,Daniela
- adding 5µl ligation product in 100µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 45 min, 180 rpm
- plate on agar with ampicillin over night
results:
- CHS+ in pYES (P72) 100 µl: 7 clones
- CHS- in pYES (P72) 100 µl: 9 clones
- CHS+ in pYES (P72) Pellet: 111 clones
- CHS- in pYES (P72) Pellet: 77 clones
- 4CL+ in pYES (P71) 100 µl: 9 clones
- 4CL- in pYES (P71) 100 µl: 3 clones
- 4CL+ in pYES (P71) Pellet: 47 clones
- 4CL- in pYES (P71) Pellet: 45 clones
- OMT+ in pYES (P72) 100 µl: 7 clones
- OMT- in pYES (P72) 100 µl: 5 clones
- OMT+ in pYES (P72) Pellet: 30 clones
- OMT- in pYES (P72) Pellet: 53 clones
- negative control P71 100 µl: 14
- negative control P71 Pellet: 71
- negative control P72 100 µl: 7
- negative control P72 Pellet: 64
Inoculation of YPD medium with S. cerevisiae and brewing yeast strain 34/70 (part 1/3)
investigator: Maddin, Aloisius
Aim of the experiment: Discrimination of growing ability in wort medium
4 different worts:
- 21%, without hop
- 21%, without hop, autoclaved
- 16%, with hop
- 16%, with hop, autoclaved
were diluted with ELGA to a solution of 12%. These 4 different worts were inoculated with 5ml of preparatory culture of S. cerevisiae and 1ml of brewing yeast strain 34/70 (provided by the Forschungsbrauerei, Weihenstephan). Start OD was measured at 550nm (for table see next experiment). Those 8 flasks were incubated over night at room temperature and 130 rpm.
Wednesday, July 18th
Ligation of plasmid pSB1C3 containing 20aaLinker with LexA and plasmid containing Gal4AD with Pif3 for fusion protein construction
Miniprep of overnight culture from picked transformed E. coli containing biobrick BBa_K165055
Inoculation of YPD medium with S. cerevisiae and brewing yeast strain 34/70 (part 2/3)
Investigators: Martin, Alois
Aim of the experiment: Discrimination of growing ability in wort medium.
After 12h of cultivating the optical density (OD) at 550nm was determined:
| Assay | Yeast | Autoclaved | Hop | OD 0h | OD 12h | delta OD |
| A1 | S. cerevisae | No | No | 1.2 | 2.2 | + 1.0 |
| A2 | S. cerevisae | Yes | No | 2.0 | 2.6 | + 0.6 |
| A3 | S. cerevisae | No | Yes | 1.8 | 2.4 | + 0.6 |
| A4 | S. cerevisae | Yes | Yes | 2.4 | 2.7 | + 0.3 |
| B1 | 34/70 | No | No | 1.6 | 2.5 | + 0.9 |
| B2 | 34/70 | Yes | No | 3.0 | 2.9 | - 0.1 |
| B3 | 34/70 | No | Yes | 2.2 | 2.5 | + 0.3 |
| B4 | 34/70 | Yes | Yes | 2.7 | 2.8 | + 0.1 |
After 12h of cultivating the S. cerevisiae is growing quite well. There is a tendency that the brewing yeast strain 34/70 is not capable to grow sufficiently in autoclaved (e.g. proteinfree) medium. Since the 34/70 was stored at 4°C over 5 days, directly used to inoculate the medium (no preparatory culture) and we only used 1ml to inoculate with (compared to 5ml of the preparatory culture of S. cerevisiae - because of obvious differences in opacity of the inoculation media) there is not yet a conclusion to be drawn. We decided to incubate for another 24h at least.
new Miniprep of P50 pYES2
Investigators: Andrea
NanoDrop concentration: 303.6 ng/µl (260/280: 1.89)
Transformation of P50 (pYes) in E.coli
Investigator: Katrin
Aim of the experiment: Get more P50(pYes) for further experiments
- adding 2-3 µl of plasmid P50 in 100µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (withouth antibiotica) and incubate at 37°C, 30 min, 180 rpm
- plate on agar with Ampicillin over night
Ligation of 4CL and PAL in pSB1C3-RFC25 and ligation of PAL in pYES
Investigator: Katrin, Daniela
Concentration (Nano Drop:
4CL+ (PCR 15) = 40.3 ng/µl
4CL- (PCR 16) = 37.3 ng/µl
PAL+ (PCR 32) = 25 ng/µl
PAL- (PCR 33) = 16.8 ng/µl
P132 (pSB1C3-RFC25 digested with Xbal and Pstl) = 33.5 ng/µl
P133 (pSB1C3-RFC25 digested with Xbal and AgeI) = 41.8 ng/µl
- required volumes were calculated using Lab Tools
4CL+ (PCR 15) with P132 (pSB1C3-RFC25)
| volume | reagent |
| 2.64 µl | P132 |
| 5.36 µl | 4CL+ (PCR 15) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
4CL- (PCR 16) with P132 (pSB1C3-RFC25)
| volume | reagent |
| 2.51 µl | P132 |
| 5.49 µl | 4CL- (PCR 16) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
PAL+ (PCR 32) with P133 (pSB1C3-RFC25)
| volume | reagent |
| 1.29 µl | P133 |
| 6.71 µl | PAL+ (PCR 32) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
PAL- (PCR 33) with P133 (pSB1C3-RFC25)
| volume | reagent |
| 0.91 µl | P133 |
| 7.09 µl | PAL- (PCR 33) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
PAL+ (PCR 32) with P72 (pYES)
| volume | reagent |
| 3.55 µl | P72 |
| 4.45 µl | PAL+ (PCR 32) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
PAL- (PCR 33) with P72 (pYES)
| volume | reagent |
| 2.79 µl | P72 |
| 5.21 µl | PAL- (PCR 33) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
Negative control:
| volume | reagent |
| 5 µl | P72, P132 or P133 |
| 12 µl | ddH2O |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
- water bath 16 °C
Miniprep of CHS and OMT in pSB1C3-RFC25
Investigator: Daniela
Aim of the experiment: Extract plasmids (pSB1C3-RFC25) that contain CHS and OMT
QIAprepS Spin Miniprep Kit
- step 3: invert 2-3 times (don't shake to avoid destruction of genomic DNA)
the Minipreps were named as follows:
- P135: CHS+ in pSB1C3-RFC25 (c = 150.9 ng/µl)
- P136: CHS- in pSB1C3-RFC25 (c = 204 ng/µl)
- P137: OMT+ in pSB1C3-RFC25 (c = 179.3 ng/µl)
- P138: OMT- in pSB1C3-RFC25 (c = 146.7 ng/µl)
Control digest of CHS and OMT in pSB1C3-RFC25
Investigator: Katrin, Daniela
Aim of the experiment: Test whether the ligation of CHS and OMT in pSB1C3 was successful
| volume | reagent |
| 2.5 µl | P135 / P136 / P137 / P138 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 14.8 µl | ddH2O |
- pSB1C3: 2070 bp
- CHS: 1173 bp
- OMT: 1059 bp
It seems as if the ligation was successful. However it is strange that the band of OMT is lower that the one of CHS. we had the wrong information about the length of OMT -> the right length of OMT is 1059 bp, so everything is fine :)
Picking clones of 4CL, CHS and OMT in pYES
investigator: Katrin, Daniela
Aim of the experiment: preculture over night for the miniprep next day
Thursday, July 19th
Transformation of P124+PCR29 and P126+PCR31
Investigator: Daniela
Aim of the experiment: Transformation of the ligation of P124+PCR29 and P126+PCR31 overnight, to see if ligation has been successful.
Operational sequence:
- P124+PCR29 ->Chlp
- negative control: NK1 ->Chlp
- P126+PCR31 ->Amp
- negative control: NK2 ->Amp
- adding 5µl ligation product in 100 µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 30 min for ampicillin and 45 min for chloramphenicol, 180 rpm
- plate on agar with ampicillin or chloramphenicol over night
EDIT from 20.07.2012: Unfortunately, no colonies for P124+PCR29 can be identified but colonies on from the concentrated pellet of P126+PCR31 ligation, BUT more but very small colonies on the control plate from the pellet. For further identification the colonies of the ligation has been picked on 21.07.2012 for miniprep and analytical digestion and gelelectrophoresis.
Restriction digest of P79
Investigator: Roman
Aim of the experiment: Double digest of p79 with Xba1 and Age1 to prepair it for ligation with an N- terminal nuclear localization signal sequence
40 µl composition
- 18 µl plasmid DNA
- 1 µl Age1
- 1 µl Xba1
- 4 µl NEBuffer 4 (10x)
- 15,6 µl ddH2O
- 0,4 µl BSA (100x)
The mixture was incubated over night at 37 °C
Oligohybridization of single-stranded DNA for creating SV40 nuclear localization signal for fusion proteins for translocating them into the nucleus
Investigator: Jeff
Aim of the experiment: Oligohybridization of single-stranded DNA (TUM12-SV40-fw and TUM12-SV40-rv)for creating SV40 nuclear localization signal for fusion proteins for translocating them into the nucleus
Operational sequence:
- 25 µL of 100 pM of TUM12-SV40-fw and 25 µL of 100 pM TUM12-SV40-rv in one ERG
- Heating up to 95 °C for 30 min
- Cooling at RT in a styropor box overnight.
Inoculation of YPD medium with S. cerevisiae and brewing yeast strain 34/70 (part 3/3)
Investigators: Andrea, (Martin, Alois)
Aim of the experiment: Discrimination of growing ability in wort medium.
After 40h of cultivating the optical density (OD) at 550nm was determined:
| Assay | Yeast | Autoclaved | Hop | OD 0h | OD 12h | OD 40h | delta OD |
| A1 | S. cerevisae | No | No | 1.2 | 2.2 | 2.5 | + 1.3 |
| A2 | S. cerevisae | Yes | No | 2.0 | 2.6 | 2.7 | + 0.7 |
| A3 | S. cerevisae | No | Yes | 1.8 | 2.4 | 2.6 | + 0.8 |
| A4 | S. cerevisae | Yes | Yes | 2.4 | 2.7 | 2.8 | + 0.4 |
| B1 | 34/70 | No | No | 1.6 | 2.5 | 2.8 | + 1.2 |
| B2 | 34/70 | Yes | No | 3.0 | 2.9 | 3.0 | 0 |
| B3 | 34/70 | No | Yes | 2.2 | 2.5 | 2.8 | + 0.6 |
| B4 | 34/70 | Yes | Yes | 2.7 | 2.8 | 2.9 | + 0.2 |
After 40h of cultivating the S. cerevisiae is growing comparably to the brewing yeast strain, though we seem to have entered a phase of substrate limitation. Autoclaved media lack solved protein and is thus for neither of the yeasts a satisfying medium; added hop also represses their growth (S. cerevisiae shows a higher susceptability - which seems logic, since the brewing yeast is selected to survive and prosper in beer brewing environment).
The next step would be to brew with our "laboratory strain" S. cerevisiae.
Preparative digest of PCR1 and PCR2 and P50 and P123
Investigator: Andrea
Digestion of PCR1 and PCR2 with XbaI and AgeI
| volume | reagent |
| 10 µl | PCR-Product |
| 2 µl | NEB Buffer |
| 0.2 µl | BSA |
| 1 µl | XbaI (10 U/µl) |
| 1 µl | AgeI (10 U/µl) |
| 7 µl | ddH2O |
Digestion of P50 with XbaI and NgoMIV
| volume | reagent |
| 10 µl | P50 |
| 4 µl | NEB Buffer |
| 0.4 µl | BSA |
| 1 µl | XbaI (10 U/µl) |
| 2 µl | NgoMIV (10 U/µl) |
| 22.6 µl | ddH2O |
Digestion of P123 with XbaI and AgeI
| volume | reagent |
| 20 µl | P123 |
| 4 µl | NEB Buffer |
| 0.4 µl | BSA |
| 1 µl | XbaI (10 U/µl) |
| 1 µl | AgeI (10 U/µl) |
| 14 µl | ddH2O |
Incubation: 37 °C, 3 h
Preparative gel
Investigator: Andrea
- 40 µl pSB1C3 + 4 µl loading dye
- 40 µl pYES2 + 4 µl loading dye
- 20 µl PCR1 + 2 µl loading dye
Ligation of digested PCR1 (digested with XbaI and AgeI) with pSB1C3 (digested with XbaI and AgeI)
Investigator: Andrea
Concentration (Nano Drop:
LIMS Citrus (PCR 1) = 18.6 ng/µl
P50 (digested with XbaI and NgoMIV) = 24.9 ng/µl
P123 (digested with XbaI and AgeI) = 44.5 ng/µl
PCR1 with P123 digested with XbaI and AgeI/NgoMIV
| volume | reagent |
| 1.19µl | P123 |
| 6.81 µl | PCR 1 |
| 1 µl | T4 DNA Ligase |
| 1 µl | T4-ligase buffer (10x) |
- water bath 16 °C over night
Miniprep of 4CL, CHS and OMT in pYES (P71 and P72)
Investigator: Daniela
Aim of the experiment: Extract plasmids (pYES) that contain 4CL, CHS and OMT
The day before 2 clones were picked for each enzyme.
QIAprepS Spin Miniprep Kit
- step 3: invert 2-3 times (don't shake to avoid destruction of genomic DNA)
Concentration:
- 4CL+ in pYES clone 1: c = 172.5 ng/µl
- 4CL- in pYES clone 1: c = 192.1 ng/µl
- CHS+ in pYES clone 1: c = 136.6 ng/µl
- CHS- in pYES clone 1: c = 85.7 ng/µl
- OMT+ in pYES clone 1: c = 116.0 ng/µl
- OMT- in pYES clone 1: c = 129.7 ng/µl
- 4CL+ in pYES clone 2: c = 160.0 ng/µl
- 4CL- in pYES clone 2: c = 144.5 ng/µl
- CHS+ in pYES clone 2: c = 128.5 ng/µl
- CHS- in pYES clone 2: c = 116.2 ng/µl
- OMT+ in pYES clone 2: c = 142.8 ng/µl
- OMT- in pYES clone 2: c = 134.5 ng/µl
Control digest of 4CL, CHS and OMT in pYES
Investigator: Daniela
Aim of the experiment: Test whether the ligation of 4CL, CHS and OMT in pYES was successful
| volume | reagent |
| 2.5 µl | 4CL+ in pYES clone 1 / 4CL- in pYES clone 1 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 14.8 µl | ddH2O |
| volume | reagent |
| 3 µl | 4CL+ in pYES clone 2 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 14.3 µl | ddH2O |
| volume | reagent |
| 3.5 µl | 4CL- in pYES clone 2 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 13.8 µl | ddH2O |
| volume | reagent |
| 3.5 µl | CHS+ in pYES clone 1 / OMT+ in pYES clone 1 / OMT- in pYES clone 1 / CHS+ in pYES clone 2 / OMT+ in pYES clone 2 / OMT- in pYES clone 2 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 13.8 µl | ddH2O |
| volume | reagent |
| 4 µl | CHS- in pYES clone 2 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI(20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 13.3 µl | ddH2O |
| volume | reagent |
| 5 µl | CHS- in pYES clone 1 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI(20U/µl) |
| 2 µl | NEB4 (10x) |
| 0.2 µl | BSA (100x) |
| 12.3 µl | ddH2O |
expected bands:
- pYES: about 5800bp
- 4CL: 1685bp
- CHS: 1173bp
- OMT: 1222bp
Transformation of ligationsproducts of 4CL and PAL in pSB1C3-RFC25 and of PAL in pYES in E.coli (see July, 18th)
Investigator: Daniela
- 4CL+ (PCR15) in pSB1C3-RFC25 (P132) ->Chlp
- 4CL- (PCR16) in pSB1C3-RFC25 (P132) ->Chlp
- PAL+ (PCR32) in pSB1C3-RFC25 (P133) ->Chlp
- PAL- (PCR33) in pSB1C3-RFC25 (P133) ->Chlp
- PAL+ (PCR32) in pYES(P72) ->Amp
- PAL- (PCR33) in pYES(P72) ->Amp
- negative control: P72 ->Amp
- negative control: P132 ->Chlp
- negative control: P133 ->Chlp
- adding 5µl ligation product in 100 µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 30 min for ampicillin and 45 min for chloramphenicol, 180 rpm
- plate on agar with ampicillin or chloramphenicol over night
Friday, July 20th
Gel purification of Xba1/ Age1 digested P79
Investigator: Roman
Aim of the Experiment: Aim of the experiment is the purification of the double digested vector p79 (pSB1C3_RFC25_Linker), in which the N- terminal SS is to be cloned.
Operational sequence:
- The mixture was seperated by gel electrophoresis (LMP agarose)
- The DNA was cut out of the gel (2 pieces) and weight ==> 200 mg each piece.
- Afterwards the plasmid DNA was extracted as described in the Quiagen Gel Purification protocol.Elution was performed in two steps (à 25 µl) with elution buffer (heated up to 40°). To improve the yeald of plasmid DNA during the elution, the column was incubated at RT for about 4 minutes before centrifugation
Miniprep of transformed E. coli XL1-Blue with pYES2 new from P50 tube (4x), BBa_K365005 (4x), BBa_K207000 (1x)and BBa_K207001 (1x)
Investigator: Jeff
Aim of the study:: Miniprep
Operational sequence:
- Miniprep has been performed after manufacturer's protocol. QIAGEN - QIAprep Spin Miniprep Kit.
Analytical digestion and gelelectrophoresis of P152, P155, P156, P157, P158, P159, P160, P161
Investigator: Jeff
Aim of the study: Analytical digestion and gelelectrophoresis of P152, P155, P156, P157, P158, P159, P160, P161, to prove the insert and backbone size.
Operational sequence:
- Analytical digestion and gelelectrophoresis were performed after lab's standard protocol.
- Digestion with XbaI and PstI-HF in Buffer 4.
Picking from the overnight transformation of the ligated P126+PCR31
Investigator: Jeff
Aim of the study: Picking from the overnight transformation of the ligated P126+PCR31, to see whether ligation is successful or not.
Operational sequence:
- Picking and overnight culture after standard laboratory's protocol. (AmpR LB-medium)
PCR of P23 (LexA, BBa_K105005) to indroduce RFC25 pre- and suffix because last ligation was not successful
Investigator: Jeff
Aim of the experiment: Last ligation with LexA was not successful and the tube with the PCR product was empty from the ligation, so one has to do another PCR for another ligation.
Operational sequence:
- Clone 3 (tube P23) of BBa_K105005 (LexA) has beed choosen for the PCR.
PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer |
| 1 µl | 10 µM Reverse Primer |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA (BBa_K105005) from P23 (Clone 3) |
| 35.75 µL | ELGA Water |
| =50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 55 °C | 60 s | |
| 68 °C | 60 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | overnight |
- Freezed at -20 °C. Stil has to be purified with PCR purification kit on the next day
Control digest of pYES (P153 and P154)
Investigator: Mary, Ingmar, Daniela
Aim of the experiment: Test whether the vector pYES is right.
| volume | reagent |
| 1.5 µl | P153 |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 16.25 µl | ddH2O |
| volume | reagent |
| 1.5 µl | P153 |
| 0.25 µl | Xbal (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 14.25 µl | ddH2O |
| volume | reagent |
| 1.5 µl | P153 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 14 µl | ddH2O |
| volume | reagent |
| 1.5 µl | P153 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 14 µl | ddH2O |
| volume | reagent |
| 1.5 µl | P153 |
| 0.25 µl | NgoMIV (10U/µl) |
| 2 µl | NEB4 (10x) |
| 16.25 µl | ddH2O |
| volume | reagent |
| 1.5 µl | P153 |
| 0.25 µl | NgoMIV (10U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 0.25 µl | SpeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13.75 µl | ddH2O |
| volume | reagent |
| 2.8 µl | P154 |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 14.95 µl | ddH2O |
| volume | reagent |
| 2.8 µl | P154 |
| 0.25 µl | Xbal (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 12.95 µl | ddH2O |
| volume | reagent |
| 2.8 µl | P154 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 12.7 µl | ddH2O |
| volume | reagent |
| 2.8 µl | P154 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 12.7 µl | ddH2O |
| volume | reagent |
| 2.8 µl | P154 |
| 0.25 µl | NgoMIV (10U/µl) |
| 2 µl | NEB4 (10x) |
| 14.95 µl | ddH2O |
| volume | reagent |
| 2.8 µl | P154 |
| 0.25 µl | NgoMIV (10U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 0.25 µl | SpeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 12.45 µl | ddH2O |
- P153
- P154
The vector seems to be correct. However NgoMIV did not digest the plasmid properly.
Saturday, July 21st
Miniprep of transformed E. coli XL1-Blue with ligation product of P126+PCR31
Investigator: Jeff
Aim of the study: Miniprep
Operational sequence:
- Miniprep has been performed after manufacturer's protocol. QIAGEN - QIAprep Spin Miniprep Kit.
Analytical digestion and gelelectrophoresis of the minipreps of transformed E. coli XL1-Blue with ligation product of P126+PCR31
Investigator: Jeff
Aim of the experiment: To see whether ligation ist successful.
- Reaction batch for each plasmid:
| Reagent | Volume in µl |
| Tango Buffer 10x | 4 µl |
| NdeI (Fermentas) | 0.5 µl |
| BamHI (Fermentas) | 0.5 µl |
| Plasmid DNA | 2.5 µl |
| ddH2O | 12.5 µl |
| TOTAL | 20 µl |
- Incubation at 37 °C for 1 h 30 min.
- Analytical gelelectrophoresis at 90 V for 1 h.
- Order of gel-pockets:
| 100 bp ladder | P172 | P173 | P174 | 1000 bp ladder |
| corrupt | corrupt | corrupt |
- Ligation was not successful!
PCR purification of PCR products from P23
Investigator: Jeff
Aim of the experiment Purification of PCR products from P23.
Operational sequence:
- The 4 tubes with PCR products of P23 were purificated with the PCR purification kit from Qiagen after manufacturer'S protocol.
Sunday, July 22nd
Preparative digest of pYes2 RFC 25 and ligation with PCR products PCR 15 - 20 and 32+33
Investigator: Ingmar
Aim of the experiment: Intsert the genes of PAL, 4CL, CHS and OMT in pYEs2 RFC 25 in order to test their expression in yeast.
Preparative digest of P 154 with XbaI and NgoMIV and of P 153 with XbaI and PstI
| volume | reagent |
| 7 µl | ddH2O |
| 2.5 µl | NEB 4 buffer |
| 2.5 µl | Tango buffer |
| 4 µl | XbaI (10 U/µl) |
| 4 µl | NgoMIV (10 U/µl) |
| 30 µl | P 153 |
| volume | reagent |
| 10 µl | ddH2O |
| 4 µl | Tango buffer |
| 2 µl | XbaI (10 U/µl) |
| 4 µl | PstI (10 U/µl) |
| 20 µl | P 154 |
Incubation: 37 °C, 3 h
DNA preparative gel electrophoresis
- gel: 1% with LMP-agarose
- Each digest product was mixed with an adequate volume of DNA loading buffer and loaded into the gel
- The separation process lasted 90 min at 70 V.
Gel picture of control digest with PstI
- All digest products show the expected bonds at 5849 bp (digest with XbaI and NgoMIV) and 5785 bp (digest with XbaI and PstI) respectively.
Gelextration
- cut the bands
- QIAquick Gel Extractrion Kit was used
- step 6 was left out
- step 9: 30µl buffer, 4 min incubation
Products were named as follows:
- P154 digested with XbaI and NgoMIV: P175
- P153 digested with XbaI and PstI: P176
Ligation
| PCR product | Volume | Vector | Volume | Volume T4 DNA Ligase | Volume T4 DNA Ligase buffer | Volume ddH20 | New Plasimd Number |
| PCR15 | 5.25 | P 176 | 2.75 | 1 µl | 2 µl | 9 µl | P 177 |
| PCR16 | 5.25 | P 176 | 2.75 | 1 µl | 2 µl | 9 µl | P 178 |
| PCR 17 | 7.2 | P 175 | 0.8 | 1 µl | 2 µl | 9 µl | P 179 |
| PCR 18 | 7.2 | P 175 | 0.8 | 1 µl | 2 µl | 9 µl | P 180 |
| PCR 19 | 7.1 | P 175 | 0.9 | 1 µl | 2 µl | 9 µl | P 181 |
| PCR 20 | 7.1 | P 175 | 0.9 | 1 µl | 2 µl | 9 µl | P 182 |
| PCR 32 | 7.25 | P 175 | 0.75 | 1 µl | 2 µl | 9 µl | P 183 |
| PCR 33 | 7.5 | P 175 | 0.5 | 1 µl | 2 µl | 9 µl | P 184 |
Monday, July 23rd
Miniprep of PAL, 4CL in pSB1C3 and PAL in pYES
Investigator: Mary
Aim of the experiment: Extract plasmids (pYES and pSB1C3-RFC25) that contain 4CL and PAL´
The day before one clone were picked for each enzyme from plates from 19th July 2012.
QIAprepS Spin Miniprep Kit
- step 3: invert 2-3 times (don't shake to avoid destruction of genomic DNA)
Concentration:
- 4CL+ (PCR15) in pSB1C3-RF25 (P132): c = 340 ng/µl -> new name of Ligationproduct after Miniprep: P185
- 4CL- (PCR16) in pSB1C3-RF25 (P132): c = 385 ng/µl -> new name of Ligationproduct after Miniprep: P186
- PAL+ (PCR32) in pSB1C3-RF25 (P133): c = 395 ng/µl -> new name of Ligationproduct after Miniprep: P187
- PAL- (PCR33) in pSB1C3-RF25 (P133): c = 255 ng/µl -> new name of Ligationproduct after Miniprep: P188
- PAL+ (PCR32) in pYES (P72): c = 190 ng/µl
- PAL- (PCR33) in pYES (P72): c = 238 ng/µl
Control digest of 4CL, PAL in pSB1C3-RFC25 and PAL in pYES
Investigator: Mary
Aim of the experiment: Test whether the ligation of 4CL, PAL in pSB1C3-RFC25 and PAL in pYES was successful
Plasmids are taken from miniprep from 23.07.2012
| volume | reagent |
| 2.5 µl | 4CL+ in pSB1C3-RFC25 / 4CL- in pSB1C3-RFC25 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
| volume | reagent |
| 2.5 µl | PAL+ in pSB1C3-RFC25 / PAL- in pSB1C3-RFC25 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
| volume | reagent |
| 2.5 µl | PAL+ in pSB1C3-RFC25 / PAL- in pSB1C3-RFC25 |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | AgeI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
expected bands:
- pYES: about 5800bp
- pSB1C3-RFC25: 2070bp
- 4CL: 1685bp
- PAL: 2151bp
Ligation of PAL+/- in pYES was not succesful Ligation of PAL+/- and 4CL+/- in pSB1C3-RFC25 were succesful! (pSB1C3 is overlapping with PAL)
APT Solubilisation and Transformation
Investigator: Mary
Aim of the experiment: solubilisation of the synthetic gene of APT and transformation of sent plasmid (including APT) in E.coli to copy it if necessary
short zentrifugation of sent product and adding 50µl bidest. H2O to it (5µg in 50µl = concentration of 0.1 µg/µl) the dissolved product was named as G1 (as Geneproduct number 1) and stored at -20°C
2µl of solved product used for transformation in Ecoli (Kit of Quiagen) and Amp-resistance (said GeneArt) Plating cells on Agar with Amp, 37°C over night
Preparative digest of APT
Investigator: Mary
Aim of the experiment: Extraction of the sequence of APT out of the sent plasmid.
| volume | reagent |
| 10µl | ddH2O |
| 4 µl | NEB 4 buffer |
| 4 µl | BSA (10x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | AgeI (20 U/µl) |
| 20 µl | solubilised APT-product (see Solubilisation APT) |
Incubation: 37°C, 2.5h
Bond at 1244bp as expected:
The bond was cutted out of the gel and stored at -20°C (P189)
Transformation of ligationsproducts of 4CL, CHS, OMT and PAL in pYES in E.Coli (Ligation see 22th of July)
Investigator: Mary
Aim of the experiment: ligation of enzymes in pYES to transform it into yeast if this was successful.
- 4CL+ (PCR15) in pYES (P176) -> new name: P177
- 4CL- (PCR16) in pYES (P176) -> new name: P178
- CHS+ (PCR17) in pYES (P175) -> new name: P179
- CHS- (PCR18) in pYES (P175) -> new name: P180
- OMT+ (PCR19) in pYES (P175) -> new name: P181
- OMT- (PCR20) in pYES (P175) -> new name: P182
- PAL+ (PCR32) in pYES (P175) -> new name: P183
- PAL- (PCR33) in pYES (P175) -> new name: P184
- adding 5µl ligation product in 100 µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 30 min 180 rpm
- plate on agar with ampicillin and incubate over night at 37°C
Gel- purification of hybridized oligos: SV40 SS (Tube PCR34)
Investigator: Roman Aim of the experiment: Purification of the hybridized oligos in order to make them ready for ligation in pSB1C3 Operational Sequence:
- oligos were seperated by gel electrophoresis (LMP agarose)
- picture:
- afterwards the DNA was cut out and extracted with the Quiagen Gel- purification kit as described in the manufacturer's protocol. Elution was performed in two steps á 25 µl elution buffer. DNA was collected in one tube, which was annotated with PCR34 purif.
- determined concentration (NanoDrop): ca. 385 ng/µl
Tuesday, July 24nd
Gelextraction of APT digested with Xbal and AgeI (P189)
Investigator: Daniela
Gelextration
- QIAquick Gel Extractrion Kit was used
- the product was named P189
- concentration: c = 9.1 ng/µl
Ligation of APT (P189) in pSB1C3-RFC25 (P133)
Investigator: Daniela
APT (P189) in pSB1C3-RFC25 (P133)
| volume | reagent |
| 0.86 µl | P133 |
| 7.14 µl | APT (P189) |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 9 µl | ddH2O |
Negative control:
| volume | reagent |
| 0.86 µl | P133 |
| 16.14 µl | ddH2O |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
- water bath 16 °C
NanoDrop determination of PCR34- PCR38
Investigator:Roman
Aim: Determination of the concentration of the samples previously to the ligation
Determined values:
| PCR Product | Concentration in ng/µl |
| PCR34 | 385 (hybridized oligos) |
| PCR35 | 41,2 |
| PCR36 | 39,5 |
| PCR37 | 41,9 |
| PCR38 | 57,4 |
- PCR38 will be used in following restriction digest
Restriction digest of p123 and PCR38 with NgoMIV and Pst1
Investigator: Roman Aim of the experiment: To prepare samples of p123 (pSB1C3 RFC25) and PCR38 for a following ligation Operational sequence: p123 was digested in a 40 µl preparation (miniprep product):
| Substance | Volume |
| Plasmid- DNA | 20 µl |
| Enzyme NgoMIV | 2 µl |
| Enzyme Pst1 | 2 µl |
| Buffer 4 | 4 µl |
| ddH2O | 12 µl |
PCR38 was digested in a 50 µl preparation (PCR product)
| Substance | Volume |
| Purified PCR product | 25 |
| Enzyme NgoMIV | 2 µl |
| Enzyme Pst1 | 2 µl |
| Buffer 4 | 5 µl |
| ddH2O | 16 µl |
The preparations were incubated at 37°C for 3h and then stored at -20°C in box The tubes were annotated with "p123_doubledigest_NgoMIV+Pst1_unpurified_20120724" and "PCR38_doubledigest_NgoMIV+Pst1_unpurified_20120724" Afterwards, the samples were load on a 1% universal agarose gel and separated at 100 V for ca. 45 min. Corresponding gel- bands were cut out and stored at -20°C over night until extraction.
Ligation of p151 and PCR34
Investigator: Roman
Aim: Ligation of fragments previously to a transformation
Operational Sequence: Length of fragments:
- p151: 2070 bp; c = 9,5 ng/µl
- PCR34: 67 bp; c = 385 ng/µl
20 µl preparation:
| Substance | Volume |
| p151- DNA | 11 µl |
| PCR34- DNA | 3 µl (out of a 1/100 dilution) |
| Ligase | 0,5 |
| T4 Ligase Buffer | 2 µl |
| ddH2O | 3,5 µl |
Negative controls were prepared the same way, using 3 µl of ddH2O instead of PCR34- DNA. The preparation was incubated 30 min at room temperature and then over night at 16°C (water bath). Afterwards, the samples were stored at 4°C until the transformation.
Ligation of p126 and PCR31
Investigator: Roman
Aim:Ligation of fragments previously to a transformation
Operational Sequence: Length of fragments:
- p126: 7961 bp; c = 46 ng/µl
- PCR34: 306 bp; c = 11,3 ng/µl
10 µl preparation:
| Substance | Volume |
| p126- DNA | 2 µl |
| PCR31- DNA | 1 µl |
| Ligase | 0,5 |
| T4 Ligase Buffer | 1 µl |
| ddH2O | 5,5 µl |
Negative controls were prepared the same way, using 1 µl of ddH2O instead of PCR31- DNA. The preparation was incubated 30 min at room temperature and then over night at 16°C (water bath).
Ligation of p175 with "PCR1, 19.7.2012" and p144
Investigator: Roman
Aim: Ligation of fragments previously to a transformation
Operational Sequence: Length of fragments:
- p175: 5900 bp; c = 107,3 ng/µl
- PCR1: 1680 bp; c = 18 ng/µl
- p144: ??? bp; c = ???
10 µl preparation:
| Substance | Volume |
| p175- DNA | 1 µl |
| PCR1/ p144- DNA | 3 µl |
| Ligase | 0,5 |
| T4 Ligase Buffer | 1 µl |
| ddH2O | 4,5 µl |
Note: This preparation was not made optimal, due to use of a wrong fragment lengths (PCR1 and p144, respectively) during the calculation of the volumes. Negative controls were prepared the same way, using 3 µl of ddH2O instead of PCR fragment. The preparation was incubated 30 min at room temperature and then over night at 16°C (water bath).
Preparation of yeast SCU Minimal Medium for Plates
Investigator: Katrin, Daniela
- The recipe was taken from the pYES2 manual.
- The ingredients were dissolved in 900 ml dest. water (ELGA) corresponding to the recipe. Lysine was only available as lysine-dihydrochloride, therefore 0.149 g instead of 0.1 g were used. Uracil was omited.
- The medium was divided in 2 x 450 ml. One will later be used as the induction medium through the addition of galactose the other one will be used as the non-induction medium (addition of glucose). Sugar solutions will be added after autoclaving to prevent maillard-reaction.
- 10 g agar and a magnetic stir bar were added to each preparation
- Autoclaving.
- Glucose solution: 10 g glucose were dissolved in 50 ml ELGA.
- Galactose solution: 10 g galactose were dissolved in 50 ml ELGA.
- No raffinose will be used.
- Autoclaving
Transformation of APT in pSB1C3-RFC25 (P190)
Investigator: Daniela
- APT (P189) in pSB1C3 (P133) -> new name: P190
- Negative control P133
- adding 5µl ligation product in 100 µl competent XL blue E.coli cells
- incubation 30 min on ice
- 5 min at 37°C
- adding cells in 1 ml LB (without antibiotica) and incubate at 37°C, 45 min 180 rpm
- plate on agar with chloramphenicol and incubate over night at 37°C
Picking Clones of 4CL, CHS, OMT and PAL in pYES and APT in original plasmid
investigator: Daniela
Aim of the experiment: preculture over night for the miniprep next day
Wednesday, 25th
Restriction digest of p123 with Xba1 and Age1
Investigator: Roman
Aim of the experiment: Aim of the experiment is the preparation of the vector pSB1C3 RFC25 (p123) for a ligation with the PCR fragment PCR34 (as repetition for the ligation of p151 with PCR34, which has probably not worked, due to low vector concentration.
Operational sequence: 30 µl preparation for restriction digest of p123
| Substance | Volume |
| p123- DNA | 10 µl (results in ca. 5µg DNA) |
| Xba1- RE | 1 µl |
| Age1- RE | 1 µl |
| NEBuffer 4 | 3 µl |
| ddH2O | 15 µl |
The preparation was incubated at 37°C for 3,5 h. Afterwards the fragments were purificated by means of a preparative gel electrophoresis (see below).
Restriction digest of p123 with Age1 and Pst1
Investigator: Roman
Aim of the experiment: Aim of the experiment is the preparation of the pSB1C3- Vector (p123) for a ligation with PCR38 (which has been digested with NgoMIV and Pst1).
Operational sequence: 30 µl preparation for restriction digest of p123
| Substance | Volume |
| p123- DNA | 10 µl (results in ca. 5µg DNA) |
| Pst1- RE | 1 µl |
| Age1- RE | 1 µl |
| NEBuffer 4 | 3 µl |
| ddH2O | 15 µl |
The preparation was incubated at 37°C for 3,5 h. Afterwards the fragments were purificated by means of a preparative gel electrophoresis (see below).
Gel extraction of PCR38 and p123 (both digested with NgoMIV and Pst1)
Investigator: Roman
Aim of the experiment: The separated PCR38 and p123 (pSB1C3 RFC25) DNA- fragments from yesterday's restriction digest (both NgoMIV and Pst1) are to be extracted from agarose- gel (which has been stored at -20°C over night), to make them ready for further usage.
Operational sequence: The extraction was performed as described in the manual of the Quiagen Gel- extraction kit. Elution was performed with 30 µl of elution- buffer EB in two steps (à 15 µl), temperated at 50°C. Concentration was determined with NanoDrop:
- PCR38: 17,3 ng/µl
- p123: 37,2 ng/µl
Tubes were annotated with p191 (p123 NgoMIV/Pst1) and p192 (PCR38 NgoMIV/Pst1).
Gel purification of p123 (Xba1/Age1 double digest) and p123 (Age1/Pst1 double digest)
Investigator: Roman
Aim of the experiment: Aim of the experiment is the purification of two different double digested p123 samples, to make them ready for ligation with the inserts PCR38 and PCR34, respectively.
Operational sequence: The samples were loaded on a 1% universal agarose gel and separated at 100 V for about 45 min. Afterwards, the corresponding gel- bands were cut out and weight. Extraction from gel was performed as described in the manufacturers protocoll (Quiagen gel extraction kit). Changes: Elution was performed in one step with 30µl ddH2O (autoclaved) temperated at 50°C. Concentrations were determined with NanoDrop:
- p123 (Xba1/Age1): 70,8 ng/µl
- p123 (Pst1/Age1): 59,1 ng/µl
The tubes were annotated with p206 (p123 Age1/Pst1) and p207 (p123 Age1/Xba1).
Ligation of p123 (Age1/Pst1) and PCR38
Investigator: Roman
Aim of the experiment: Ligation of the fragments p123 and PCR38
Operational sequence: Length of fragments:
- p123: 2070 bp; c = 59,1 ng/µl
- PCR38: 617 bp; c = 17,3 ng/µl
20 µl preparation:
| Substance | Volume |
| p123- DNA | 3 µl (ca. 180 ng vector- dna) |
| PCR38- DNA | 10 µl |
| Ligase | 0,5 |
| T4 Ligase Buffer | 2 µl |
| ddH2O | 4,5 µl |
Negative controls were prepared the same way, using 10 µl of ddH2O instead of PCR38- DNA. The preparation was incubated 30 min at room temperature and then over night at 16°C (water bath). Afterwards, the samples were stored at 4°C in the fridge until transformation.
Ligation of p123 (Age1/Xba1) and PCR34
Investigator: Roman
Aim of the experiment: Ligation of fragments p123 and PCR34 Operational sequence: Length of fragments:
- p123: 2070 bp; c = 70,8 ng/µl
- PCR34: 67 bp; c = 3,85 ng/µl (1:100 dilution of original sample)
20 µl preparation:
| Substance | Volume |
| p123- DNA | 3 µl (ca. 210 ng vector- dna) |
| PCR34- DNA | 6 µl (out of 1:100 dilution) |
| Ligase | 0,5 |
| T4 Ligase Buffer | 2 µl |
| ddH2O | 8,5 µl |
Negative controls were prepared the same way, using 6 µl of ddH2O instead of PCR34- DNA. The preparation was incubated 30 min at room temperature and then over night at 16°C (water bath). Afterwards, the samples were stored at 4°C in the fridge until transformation.
Miniprep of 4CL, CHS, OMT, PAL in pYes and APT in original plasmid from GeneArt
Investigator: Katrin
Aim of the experiment: extraction of plasmids (pYes2) that contain 4CL, CHS, OMT, PAL; extraction of original plasmid from GeneArt with APT
QIAprepS Spin Miniprep Kit
- step 3: invert 2-3 times (don't shake to avoid destruction of genomic DNA)
the Minipreps were named as follows:
- P193: 4CHL+ in pSB1C3-RFC25 (c = 110,7 ng/µl)
- P194: 4CL- in pYEs (c = 342,8 ng/µl)
- P195: CHS+ in pYEs (c = 423,6 ng/µl)
- P196: CHS- in pYEs (c = 92,1 ng/µl)
- P197: OMT+ in pYEs (c = 394,6 ng/µl)
- P198: OMT- in pYEs (c = 183,0 ng/µl)
- P199: PAL+ in pYEs (c = 138,4 ng/µl)
- P200: PAL- in pYEs (c = 464,5 ng/µl)
- P201: APT in original plasmid (c = 260,0 ng/µl)
Transformation of Ligation-products into E.coli XL-1 Blue
Investigator: Andrea
- for each Biobrick 100 µl cells were used and pooled together with 5 µl of ligation product from the 24th of july
(3 hours at 16°C and over night at 4 °C) and from the 19th of july (5 days at 16°C)
- Incubation on ice for 30 min
- 5 min heat shock at 37 °C
- cells were prefilled with 1 ml of LB-medium and incubated in a cell-culture shaker at 37 °C for 40 min
- 100 µl of these cell suspension were plated on antibiotic selection plates (ligations in pYES: Ampicillin; ligations in pSB1C3: Chloramphenicol)
- cell suspension was centrifuged at 13000 rpm for 90 s for resuspending the pellet with 100 µl LB and plating also
- incubation at 37 °C overnight
Thursday, 26th
Transformation of ligation products of P126+PCR31, P151+PCR34, P123+PCR38 and P123+PCR34 in E. coli XL1-Blue
Investigator: Jeff
Aim of the experiment:Transformation of ligation products of P126+PCR31 (AmpR), P151+PCR34 (CamR), P123+PCR38 (CamR) and P123+PCR34 (CamR) in E. coli XL1-Blue
Operational sequence:
- Performed after lab's standard protocol.
Miniprep of APT in pSB1C3-RFC25 and control digestion with Xba1 and Pst1
Investigator: Mary
Aim of the experiment: extraction of plasmid (pSB1C3-RFC25) that contains APT and testing if ligation was succesful; three clones were picked the day before
Miniprep
QIAprepS Spin Miniprep Kit
- step 3: invert 2-3 times (don't shake to avoid destruction of genomic DNA)
the Minipreps were named as follows:
- APT in pSB1C3-RFC25 clone 1: c=96 ng/µl
- APT in pSB1C3-RFC25 clone 2: c=108 ng/µl
- APT in pSB1C3-RFC25 clone 3: c=97 ng/µl
Control digest
| volume | reagent |
| 2.5 µl | 4CL+/-, PAL+/-, CHS+/-, OMT+/-, APT+/- |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
expected bands:
- pSB1C3-RF25: 2070bp
- APT: 1244bp
Control digest of 4CL, PAL, CHS, OMT, APT in pYES
Investigator: Mary
Aim of the experiment: Test whether the ligation of 4CL, PAL, CHS, OMT, APT in pYES was successful
Plasmids are taken from miniprep from 25.07.2012
| volume | reagent |
| 2.5 µl | 4CL+/-, PAL+/-, CHS+/-, OMT+/-, APT+/- |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
expected bands:
- pYES: about 5800bp
- 4CL: 1685bp
- PAL: 2151bp
- CHS: 1173bp
- OMT: 1222bp
- APT: 1244bp
pour on yeast agarplates
Investigator: Mary
Aim of the experiment: prepare selection-agarplates without Uracil (Some plates including glucose, some including galactose)
it was strange that the agar was still pretty liquid after waiting for one hour.
see pYES-manual
Picking of clones of transformation from 25.07.
Investigator: Lara
Aim: Amplification of clones for extraction of plasmids and subsequent proof of functional ligation.
6 clones were picked for each ligation (PCR1 in pYESnew, PCR2 (p144) in pYESnew and PCR in pSB1C3. Clones containing pYESnew were put into 4 ml LB with 4 µl of ampillicin stock solution, clones containing pSB1C3 were put into chloramphenicol-containing media. Incubation at 37 °C over night.
Transformation of plasmids P40&P42 into E.coli XL1 blue
Investigator: Lara
Aim:
Transformation of plasmids P40 and P42 into E.coli XL1 blue to get plasmids of higher quality. P40&P42 are directly extracted from the expression strain BL21(DE). The quality of the plasmids might not be sufficient for Quickchange site-directed mutagenesis. Therefore transformation into E.coli XL1 blue and subsequent plasmid preparation to get plasmids with sufficient quality for SDM.
Procedure:
100 μl of competent XL1 blue cells were thawed on ice. 1 µl plasmid DNA (P40 or P42) was added. Incubation for 30 min on ice. 5 min heatshock at 37°C. 1 ml of LB medium without antibiotic was added, incubation for 45 min at 180 rpm/37°C. After incubation, 100 µl of the cell suspension were plated on antibiotic containing plates (P40: Kan, P42: Amp). The remaining solution was centrifuged for 60 sec, resuspended in 100 µl LB and plated, as well. Incubation at 37 °C over night.
Friday, July 27th
Picking from the overnight transformation of ligated P126+PCR31, P151+PCR34, P123+PCR38 and P123+PCR34 in E. coli XL1-Blue
Investigator: Jeff
Aim of the experiment Picking of transformated E. coliXL1-Blue with ligation product of P126+PCR31 (AmpR), P151+PCR34 (CamR), P123+PCR38 (CamR) and P123+PCR34 (CamR).
Procedure
- Colonies were picked with pipette tips and transformed into a new cell-culture tube with 4 ml of LB-medium and antibiotics and were put overnight in a 180 rpm cell culture shaker at 37 °C. P126+PCR31 (AmpR), P151+PCR34 (CamR), P123+PCR38 (CamR) and P123+PCR34 (CamR)
Transferring E. coliXL1-Blue colonies, transformed with BBa_I15008 (heme oxygenase) and BBa_K105005 (LexA), from old antibiotic plates on new plates
Investigator: Jeff
Aim of the experiment With a sterile incolation loop the colonies were transferred on a new antibiotic plate, because the plates are already 4 weeks old.
Procedure
- Incolation loop was put for few second into the flame of a Bunsen burner
- Colonies from plate with BBa_I15008 (heme oxygenase, KanR) and BBa_K105005 (LexA, AmpR) were transferred on a new antibiotic plate
- Overnight-culture at 37 °C
Miniprep and control digest of PAL in pYES picked on thursday 26th July
Investigator: Ingmar
Aim of the experiment: Test whether the ligation of PAL in pYES was successful
Miniprep Using a Quiagen Miniprep kit a plasmid isolation of two picked clones of the PAL+ in pYes2 RFC25 ligation was done. The miniprep product of the first clone was named P219, the one of the second clone P220. Both were used for the following control digest.
control digest
| volume | reagent |
| 2.5 µl | PAL+ |
| 0.25 µl | Xbal (20U/µl) |
| 0.25 µl | PstI (20U/µl) |
| 2 µl | NEB4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
expected bands:
- pYES:5819bp
- PAL: 2151bp
As the picture only shows one band at the expected length of the vecotr backbone the ligation was not successfull and will be repeated with a reduced quotient of insert to vector of 3.
Ligation of PAL+ (PCR32) and APT (P189) with pYes2 RFC25 (P175)
Investigator: Ingmar
Aim of the experiment: Insert the genes of PAL and APT in pYEs2 RFC 25 in order to test their expression in yeast.
Ligation
| PCR product | Volume | Vector | Volume | Volume T4 DNA Ligase | Volume T4 DNA Ligase buffer | Volume ddH20 | New Plasimd Number |
| PAL+ (PCR32) | 6.6 | P 175 | 1.4 | 1 µl | 2 µl | 9 µl | P 221 |
| APT (P 189) | 7.06 | P 175 | 0.94 | 1 µl | 2 µl | 9 µl | P 222 |
| control: ddH2O | 7 | P 175 | 1 | 1 µl | 2 µl | 9 µl |
The Ligation product of PAL+ and pYes was labeled P233, the one of APT and pYes P234.
Plasmid extraction of plasmids PCR1/pYes, P144/pYES, PCR1/pSB1C3
Investigator: Lara
Aim of the experiment: Extract plasmids from ligation of PCR1 in pYES, P144 in pYES and PCR1 in pSB1C3 for further analytical restriction digest. (6 clones were picked for each ligation and plasmids subsequently extracted.)
Prodedure: Plasmids were extraced by using Qiagen plasmid miniprep kit.
Following concentrations were obtained:
1_1 - 1_6
- PCR1 in pYESnew clone 1: c=66 ng/µl
- PCR1 in pYESnew clone 2: c=182 ng/µl
- PCR1 in pYESnew clone 3: c=98 ng/µl
- PCR1 in pYESnew clone 4: c=326 ng/µl
- PCR1 in pYESnew clone 5: c=87 ng/µl
- PCR1 in pYESnew clone 6: c=156 ng/µl
2_1 - 2_6
- P144(PCR2) in pYESnew clone 1: c=195 ng/µl
- P144(PCR2) in pYESnew clone 2: c=202 ng/µl
- P144(PCR2) in pYESnew clone 3: c=198 ng/µl
- P144(PCR2) in pYESnew clone 4: c=77 ng/µl
- P144(PCR2) in pYESnew clone 5: c=95 ng/µl
- P144(PCR2) in pYESnew clone 6: c=200 ng/µl
3_1 -3_6
- PCR1 in pSB1C3-RFC25 clone 1: c=116 ng/µl
- PCR1 in pSB1C3-RFC25 clone 2: c=110 ng/µl
- PCR1 in pSB1C3-RFC25 clone 3: c=116 ng/µl
- PCR1 in pSB1C3-RFC25 clone 4: c=112 ng/µl
- PCR1 in pSB1C3-RFC25 clone 5: c=139 ng/µl
- PCR1 in pSB1C3-RFC25 clone 6: c=80 ng/µl
Restriction digest of plasmids from 6 clones each of PCR1/pYESnew, P144(PCR2)/pYESnew, PCR1/pSB1C3
Investigator: Lara
Aim of the experiment: Analytical restriction digest of plasmids to check for insert.
| volume | reagent |
| 3 µl | DNA |
| 2 µl | Buffer 4 |
| 0,25 µl | Xba1 |
| 0,25 µl | Spe1-HF |
| 0,2 µl | BSA(100x) |
| 14,3 µl | ddH2O |
Incubation: 37 °C; 1,5 h
A mastermix for 19 samples was made.
Analytical gel electrophoresis
Investigator: Lara
Aim of the experiment: Analytical gel electrophoresis of products from restriction digest.
Small-Scale Yeast Transformation of 4CL, CHS, OMT and PAL- in pYES
Investigator: Daniela
Aim of the experiment:Transformation of P193 - P198 and P200 in Yeast
The protocol on page 13 of the pYES2 manual was used.
Only modifications are noted:
step 1: inoculate in 4 ml YPD medium
step 2: OD600 = 10 -> to determine a OD600 of 0.4 in 50 ml, 2.5 ml yeast suspension and 47.5 ml YPD medium were used (1:20 dilution)
steps 3 and 4: centrifugation for 5 min, 4 °C
step 5: room temperature was about 35 °C
step 6:
1 µg plasmid DNA
- P193: c = 110.7 ng/µl -> 9 µl
- P194: c = 342.8 ng/µl -> 3 µl
- P195: c = 423.6 ng/µl -> 2.4 µl
- P196: c = 92.1 ng/µl -> 11 µl
- P197: c = 394.6 ng/µl -> 2.5 µl
- P198: c = 183.4 ng/µl -> 5.5 µl
- P200: c = 464.5 ng/µl -> 2.2 µl
100 µg denatured sheared salmon sperm: the one from Simon was used -> 10 µl
step 8: incubation was carried out at 35 °C because the thermoblock did not achieve a lower temperature
The SCU- plates are incubated at 30°C over the weekend.
Colonies were grown on 2 plates (31.07.2012): CHS+ : 1 colony OMT+ : 3 colonies
-> repetition of transformation on plates with glucose!
Saturday, July 28th
Miniprep of picked E. coli XL1-Blue transformated with ligation products of P126+PCR31, P151+PCR34, P123+PCR38 and P123+PCR34
Sunday, July 29th
Picking of clones from PAL+ for a overnight culture
Investigator: Ingmar
Aim of the experiment:Inoculation of LB medium with clones picked from the transfomation of P183 (PAL+ (PCR32) in pYES (P175)) done on Monday, 23rd July in order to test on monday wheather the Ligation was successfull. Operationale sequence:
- Two clones were picked an transferred to 6 ml LB medium containing 1:1000 Ampicillin.
- Incubation overnight at 37°C, 180 rpm.
Monday, July 30th
Analytical digestion and gelelectrophoresis of the plasmids P221-P232
Investigator: Saskia
Aim of the experiment: Analytical digest of P221-P223 with NdeI and BamHI and of P224-P232 with XbaI and AgeI-HF and analytical gelelectrophoresis .
Procedure:
- Analytical restriction digest and gelelectrophoresis of P221-P223 with NdeI and BamHI
- Reaction batch for each plasmid:
| Reagent | Volume in µl |
| Tango Buffer 10x | 4 µl |
| BamHI (NEB) | 0.5 µl |
| NdeI (NEB) | 0.5 µl |
| Plasmid DNA | 2.5 µl |
| ddH2O | 12.5 µl |
| TOTAL | 20 µl |
- Incubation at 37 °C for 1 h 20 min.
- Analytical gelelectrophoresis (1%) at 90 V for 40-45 min.
- Analytical restriction digest and gelelectrophoresis of P224-P232 with XbaI and AgeI-HF
- Reaction batch for each plasmid:
| Reagent | Volume in µl |
| NEBuffer4 10x | 2 µl |
| BSA 100x | 0.2 µl |
| XbaI (Fermentas) | 0.25 µl |
| AgeI-HF (Fermentas) | 0.25 µl |
| Plasmid DNA | 2.5 µl |
| ddH2O | 14.8 µl |
| TOTAL | 20 µl |
- Incubation at 37 °C for 1 h 20 min.
- Analytical gelelectrophoresis (1%) at 90 V for 40-45 min.
- Order of gel-pockets:
| 1 kbp ladder | P221 | P222 | P223 | 100 bp ladder |
| Corrupt | Corrupt | Corrupt |
- Order of gel-pockets:
| 1 kbp ladder | P224 | P225 | P226 | P227 | P228 | P229 | P230 | P231 | P232 | 100 bp ladder |
| Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | ?Questionable? | ?Questionable? | ?Questionable? |
Transformation of ligation products of PAL+ (PCR32) and APT(P189) in pYES2 RFC 25 (P175)into E.coli Xl1-Blue
Investigator: Ingmar
Aim of the experiment:Transformation of the ligation products of PAL+ (PCR32), APT (P189) and the negative control in pYes in XL1 Blue. Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 5 µl of the ligation products
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Miniprep and control digest of PAL+ in pYES picked on sunday 29th July
Investigator: Ingmar
Aim of the experiment: Test whether the ligation of PAL in pYES was successful
Miniprep
Using a Quiagen Miniprep kit a plasmid isolation of two picked clones of the PAL+ in pYes2 RFC25 ligation was done. The miniprep product of the first clone was named P235, the one of the second clone P236. Both were used for the following control digest.
control digest
| volume | reagent |
| 2.5 µl | PAL+ |
| 0.25 µl | Xbal (10U/µl) |
| 0.25 µl | PstI (10U/µl) |
| 2 µl | Tango (10x) |
| 15 µl | ddH2O |
expected bands:
- pYES:5795bp
- PAL: 2201bp
The picture shows in both cases the two expected bands at 5795 bp an 2201 bp. Therefore the ligation was successfull.
Miniprep of Schwab plasmids P40 & P42 from E.coli XL1 blue
Investigator: Lara
Aim of experiment: Get plasmids carrying lavendula limonene synthase gene for subsequent site-directed mutagenesis.
Plasmid concentrations:
- P40 clone 1: 155 ng/µl
- P40 clone 2: 114 ng/µl
- P42 clone 1: 40 ng/µl
- P42 clone 2: 32 ng/µl
Analytical restriction digest of extracted Schwab plasmids carrying Lavendula LS
Investigator: Lara
Aim of experiment: Check whether insert is OK.
- Digest of P40 clone 1 and clone 2
| volume | reagent |
| 2.5 µl | plasmid DNA |
| 0.25 µl | NcoI |
| 0.25 µl | HindIII |
| 2 µl | Buffer Tango (10x) |
| 15 µl | ddH2O |
- Digest of P42 clone 1 and clone 2
| volume | reagent |
| 4.5 µl | plasmid DNA |
| 0.25 µl | EcoRI |
| 0.25 µl | NotI |
| 2 µl | Buffer Orange (10x) |
| 13 µl | ddH2O |
Incubation for 1,5 h at 37°C.
Clones renamed:
- P40 clone 1: now P241
- P40 clone 2: now P242
- P42 clone 1: now P243
- P42 clone 2: now P244
Analytical restriction digest of plasmids containing citrus LS (repetition of digest from July, 27th)
Investigator: Lara
Aim of experiment: Check whether ligations PCR1/pYESnew, P144(PCR2)/pYESnew and PCR1/pSB1C3 have worked. This time, a standard protocol with 2,5 µl of DNA was used. Furthermore, Pst-1 HF was used instead of Spe1 (as on Friday, July 27th).
- Digest of plasmids PCR1/pYESnew clone 1,3,5 and 6; P144(PCR2)/pYESnew clone 1,3,5 and 6; PCR1/pSB1C3 clone 1,5 and 6.
| volume | reagent |
| 2.5 µl | plasmid DNA |
| 0.25 µl | Pst1-HF |
| 0.25 µl | Xba1 |
| 2 µl | NEB Buffer 4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
Incubation for 1,5 h at 37 °C.
Tuesday, July 31st
Picking from the overnight transformation of ligated P123+PCR38 in E. coli XL1-Blue from July, 27th
Investigator: Jeff, Georg
Aim of the experiment: Repeat picking from the overnight transformation of ligated P123+PCR38 in E. coli XL1-Blue because the last 3 picked colonies are negative.
Procedure:
- Colonies were picked with pipette tips and transformed into a new cell-culture tube with 4 ml of LB-medium and antibiotics and were put overnight in a 180 rpm cell culture shaker at 37 °C. P123+PCR38 (CamR).
Quickchange of Schwab plasmid P40 clone 1
Investigator: Lara
Aim of experiment: Remove Age1 restriction site in gene coding for lavendula limonene synthase.
PCR
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 2 µl | Plasmid P241 |
| 1 µl | 1:10 dilution of O60 (10 pmol/µL) |
| 1 µl | 1:10 dilution of O61 (10 pmol/µL) |
| 18 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 12 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 7 min |
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation into E.coli Xl1-Blue
- thawing of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the SDM PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Kan-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Kan-LB-plate
Preparative digest of PCR1, PCR2 with XbaI and AgeI (additional positive control P155)
Investigator: Andrea
Digestion of PCR1, PCR2 and P155 (pYES) with XbaI and AgeI
| volume | reagent |
| 8 µl | PCR1 |
| 4 µl | NEB Buffer |
| 0.4 µl | BSA |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | AgeI (20 U/µl) |
| 25,6 µl | ddH2O |
| volume | reagent |
| 10 µl | PCR2 |
| 4 µl | NEB Buffer |
| 0.4 µl | BSA |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | AgeI (20 U/µl) |
| 23,6 µl | ddH2O |
| volume | reagent |
| 10 µl | P155 |
| 4 µl | NEB Buffer |
| 0.4 µl | BSA |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | AgeI (20 U/µl) |
| 23,6 µl | ddH2O |
Mastermix of Buffer, BSA and Enzymes was added to the DNA and water.
Incubation: 37 °C, 3 h
Ligation of digested PCR1 with P133 (pSB1C3) and P175 (pYES) (XbaI and AgeI)
Investigator: Andrea
Concentration (Nano Drop:
LIMS Citrus (PCR 1) = ? ng/µl
LIMS Citrus (PCR 2) = ? ng/µl
P175 = 107,3 ng/µl
P133 = 41.8 ng/µl
PCR1 and P175
| volume | reagent |
| 1 µl | P175 |
| 14,27 µl | PCR 1 |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 1,73 µl | ddH2O |
PCR1 and P133
| volume | reagent |
| 2,4 µl | P133 |
| 14,27 µl | PCR 1 |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 0,33 µl | ddH2O |
PCR2 and P175
| volume | reagent |
| 1 µl | P175 |
| 11,99 µl | PCR 2 |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 4,01 µl | ddH2O |
PCR2 and P133
| volume | reagent |
| 2,4 µl | P133 |
| 11,99 µl | PCR 2 |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 2,61 µl | ddH2O |
Negativ control P175
| volume | reagent |
| 1 µl | P175 |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 16 µl | ddH2O |
Negativ control P133
| volume | reagent |
| 2,4 µl | P175 |
| 1 µl | T4 DNA Ligase |
| 2 µl | T4-ligase buffer (10x) |
| 14,4 µl | ddH2O |
- water bath 16 °C over night
Media and plates for yeast transformation
Investigator: Martin, Alois
Aim of the experiment: Producing media and plates for yeast transformation (according to pYes_manual_kommentiert)
- YPD-Medium, YPD-plates, 10X TE, 10X LiAc, 1x LiAc/0.5x TE
August
Wednesday, August 1st
Miniprep of picked E. coli XL1-Blue transformated with ligation products of P123+PCR38
Investigator: Jeff, Dennis
Aim of the experiment: Miniprep of picked E. coli XL1-Blue transformated with ligation products of P123+PCR38.
Procedure:
- Miniprep was performed after manufacturer's protocol. (P251-P260)
Analytical digestion and gelelectrophoresis of Miniprep E. coli XL1-Blue transformated with ligation products of P123+PCR38 and of pGBKT7 plasmid
Investigator: Dennis, Jeff
Aim of the experiment: Analytical digestion and gelelectrophoresis of Miniprep E. coli XL1-Blue transformated with ligation products of P123+PCR38 to see whether the ligation was successful and of pGBKT7 plasmid to see if the MfeI and BamHI restriction enzymes work.
Procedure:
- Mastermix for digestion with XbaI+AgeI-Hf of ligation products of P123+P38:
| Reagent | Volume in µl |
| Buffer 4 (10x) | 22 µl |
| BSA (100x) (NEB) | 2.2 µl |
| XbaI (NEB) | 1.75 µl |
| AgeI-Hf (NEB) | 1.75 µl |
| ddH2O | 162.8 µl |
| TOTAL MASTERMIX | 192.5 µl |
- 17.5 µl of Mastermix + 2.5 µl of the plasmid DNA (P251-P260)
- Composition for analytical restriction digestion of P99 with MfeI+BamHI
| Reagent | Volume in µl |
| Buffer G (10x) | 2 µl |
| MfeI (NEB) | 0.25 µl |
| BamHI (NEB) | 0.25 µl |
| Plasmid DNA (P99; pGBKT7) | 2.5 µl |
| ddH2O | 15 µl |
| TOTAL | 20 µl |
- Analytic Gelelectrophoresis for 1 h at 90 V.
- Order of gel-pockets:
| 1 kbp ladder | P251 | P252 | P253 | P254 | P255 | P256 | P257 | P258 | P259 | P260 | 100 bp ladder |
| 100 ladder | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | Corrupt | 1 kb ladder |
- Order of gel-pockets:
| 100 bp ladder | P99 control digestion with MfeI+BamHI | P99 control undigested | P123 control undigested | 100 bp ladder (accidently) |
| Okay | Okay | Okay |
→MfeI and BamHI are working! (Expected DNA strands for P99 digestion at 731 bp, 2443 bp and 4129 bp; that's what we got.)
Preperative digestion and gelelectrophoresis of PCR37 with XbaI+AgeI-HF
Investigator: Jeff
Aim of the experiment:
Procedure:
| volume | reagent |
| 25 µl | PCR37 |
| 5 µl | NEBuffer 4 (10x) |
| 0.5 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | AgeI-HF (20 U/µl) |
| 17.5 µl | ddH2O |
| =50 µl | TOTAL |
Gel extraction of preperative gelelectrohphoresis of PCR37
Investigator: Jeff
Aim of the experiment:
Ligation of PCR39+P207 for cloning LexA with new RFC25 pre- and suffix into a plasmid
Investigator: Jeff
Aim of the experiment:
Procedure
| Substance | Volume |
| P207 (digested plasmid DNA (P123) with XbaI and AgeI-HF) | 1.42 µl (~100 ng vector dna) |
| PCR39 (digested insert DNA (PCR37) with XbaI and AgeI-HF) | 10.17 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 5.91 µl |
| TOTAL | =20 µl |
PCR of P80 to introduce RFC25 pre- and suffix to Pif3
Investigator: Jeff
Aim of the experiment:
Procedure:
- PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer O31 1:10 dilution (TUM12-LSPS-fw) |
| 1 µl | 10 µM Reverse Primer O32 1:10 dilution (TUM12-LSPS-rv ) |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA P80 |
| 35.75 µL | ELGA Water |
| =50 µL | TOTAL |
- The PCR program was performed after following scheme with following conditions
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 55 °C | 60 s | |
| 68 °C | 30 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | infinite |
PCR purification of PCR of P80
Investigator: Jeff
Aim of the experiment:
Transformation of ligation products and quickchange products
Investigator: Lara
Aim: Transformation of ligation products (ligation July 31st) and quickchange products (repetition, quickchange of July 31st).
Transformation into E.coli Xl1-Blue
- thawing of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the SDM PCR product or 5 µl of ligation product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Kan-LB-plate (Quickchange product), Amp-plate (pYES) or Chloramphenicol (pSB1C3)
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an antibiotic containing LB-plate
Miniprep and control digest of APT in pYES
Investigator: Katrin
Aim of the experiment: Test whether the ligation of APT in pYES was successful
Miniprep
Using a Quiagen Miniprep kit, a plasmid isolation of three picked clones of a APT in pYes2 RFC25 ligation was done. The miniprep products were named P261 (1st clone), P262 (2nd clone) and P263 (3rd clone).All three of them were used for the following control digest.
The resulting plamid-concentrations (measured with nanodrop) were clone 1 (P261): 149,0 ng/µl clone 2 (P262): 153,5 ng/µl clone 3 (P263): 158,4 ng/µl
control digest
| volume | reagent |
| 2.5 µl | APT (clones 1,2,3) |
| 0.25 µl | Xbal (10U/µl) |
| 0.25 µl | AgeI (10U/µl) |
| 2 µl | NEB 4 (10x) |
| 2 µl | BSA (10x) |
| 13 µl | ddH2O |
expected bands (pYes has 2 AgeI restriction site):
- pYES fragment 1:5398 bp
- pYes fragment 2: 451
- APT: 1244 bp
Media and plates for yeast transformation
Investigator: Martin, Alois
Aim of the experiment: Producing media and plates for yeast transformation (according to pYes_manual_kommentiert)
- 20% glucose, 20x SC stock (without uracil and without yeast nitrogen base), SC plates with glucose
Thursday, August 2nd
Transformation of ligation product P207/PCR39
Investigator: Lara
Aim: Transformation of ligation products P207/PCR39 and P207/NK.
Transformation into E.coli Xl1-Blue
Unfortunately the lab assistents went home so that the incubation times had to be shortened.
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 5 µl of the ligation products
- incubation on ice for 25 min
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubation at 37°C and 180 rpm for 25 min
- plating of 100 µl on an Chlp-LB-plate
- sedimenting the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspension of the sediment in 100 µl LB-medium and plating it as well on an Chlp-LB-plate
Repetition of Quickchange mutagenesis of lavendula limonene synthase
Investigator: Lara
Aim: No colonies were observable after second transformation of quickchange PCR products. Therefore the quickchange pcr reaction was repeated with a modified protocol:
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P242 |
| 0.5 µl | 1:10 dilution of O60 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P242 |
| 0.5 µl | 1:10 dilution of O61 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Turbo DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 68°C | 7 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were once more applied.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
! Unfortunately someone turned off the heat block, so that the reaction tube was below 37 °C (approx. 25 °C) for about 45 min. After recognition (after one hour of incubation), the heat block was turned on again and the reaction tube was incubated for another 30 min.
Transformation into E.coli Xl1-Blue
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the PCR product
- incubation on ice for 25 min
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubation at 37°C and 180 rpm for 25 min
- plating of 100 µl on an Kan-LB-plate
- sedimenting the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspension of the sediment in 100 µl LB-medium and plating it as well on an Kan-LB-plate
PCR of C-LS3 (BBa_I742111, Trafo 16.6)
Investigator: Lara
Aim: To get more PCR product in case another preparative digest is needed.
Primer with consensus sequence
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer O27 |
| 1 µl | 10 µM Reverse Primer O30 |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µl) |
| 1 µl | Plasmid DNA (BBa_I742111) Clone 3 |
| 35.75 µl | dd water |
| 50 µL | TOTAL |
Primer without consensus sequence
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer O28 |
| 1 µl | 10 µM Reverse Primer O30 |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µl) |
| 1 µl | Plasmid DNA (BBa_I742111) Clone 3 |
| 35.75 µl | dd water |
| 50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 47 °C | 30 s | |
| 68 °C | 1,75 min | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | 1 h |
Analytical gel electrophoresis
Preparation of YPD Medium and Picking of S. cerevisiae clones
Investigator: Lara, Katrin
YPD Yeast Extract Peptone Dextrose Medium (1 liter)
the following substances were dissolved in 1000 ml of water and then autoclaved
- 1% yeast extract
- 2% peptone
- 2% dextrose (D-glucose)
Picking of two clones of S. cerevisiae DD (17.17.2012): The clones were put into YPD medium and incubated at 30 °C over night. Additional 0,8 g Glucose were added to one of the cell culture tubes.
Friday, August 3rd
Transformation of E.coli XL1 blue with ligation product P207/PCR39
Investigator: Lara
Aim: Incubation times of transformation of August 2nd had to be shortened because the lab employees left. Therefore the transformation didn't work properly. --> Repetition of the transformation of E.coli XL1 blue with ligation products P207/PCR39 and P207/NK.
Transformation into E.coli Xl1-Blue
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 5 µl of the ligation products
- incubation on ice for 30 min
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubation at 37°C and 180 rpm for 45 min
- plating of 100 µl on an Chlp-LB-plate
- sedimenting the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspension of the sediment in 100 µl LB-medium and plating it as well on an Chlp-LB-plate
Analytical restriction digest of quickchange products
Investigator: Lara
Aim: Check whether site directed mutagenesis to eliminate Age1 restriction site has worked.
Quickchange PCR product purification:
The Quickchange PCR products were purified with Qiagen PCR Purification Kit before analytical restriction digest:
- P241 SDM: 7 ng/µl
- P242 SDM: 63 ng/µl
Restriction digest of p241 SDM (31.7.), p242 SDM (2.8.) and p242 (as positive control)
| volume | reagent |
| 2.5 µl | plasmid DNA |
| 0.25 µl | Age1-HF |
| 2 µl | NEBuffer 4 |
| 2 µl | BSA (10x) |
| 13.25 µl | ddH2O |
Analytical Gel
1. Standard (1 kb Gene Ruler)
2. p242, digested with Age1 (positive control)
3. p241 SDM
4. p241 SDM, digested with Age1
5. p242 SDM
6. p242 SDM, digested with Age1
7. PCR 42
8. PCR 43
Transformation of E.coli XL1 blue with P242 SDM
Investigator: Lara
Aim: Incubation times of transformation of August 2nd had to be shortened because the lab employees left. Therefore the transformation didn't work properly. --> Repetition of the transformation with P242 SDM.
Transformation into E.coli Xl1-Blue
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- two transformations: 1. addition of 1 µl of the purified Quickchange PCR product; 2. addition of 4 µl of the purified Quickchange PCR product
- incubation on ice for 30 min
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubation at 37°C and 180 rpm for 45 min
- plating of 100 µl on an Kan-LB-plate
- sedimenting the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspension of the sediment in 100 µl LB-medium and plating it as well on an Kan-LB-plate
Repetition of small-Scale Yeast Transformation of 4CL, CHS, OMT, PAL and APT in pYES
Investigator: Mary
Aim of the experiment:Transformation of P193 - P198, P200, P236, P263 and GFP (as a positive control - from simon) in Yeast
The protocol on page 13 of the pYES2 manual was used.
Only modifications are noted:
step 1: inoculate in 4 ml YPD medium
step 2: OD600 = 9 -> to determine a OD600 of 0.4 in 50 ml, 2.2 ml yeast suspension and 42.5 ml YPD medium were used, 5ml 20% Glucose was added (wrong information about the YPD-medium: we suggested that Glucose caramelizes when autoclaved together with medium - but it is not the case! Only agar and glucose will caramelize).
steps 3 and 4: centrifugation for 5 min, 4 °C
step 6:
1 µg plasmid DNA
- P193: c = 110 ng/µl -> 9.1 µl
- P194: c = 342 ng/µl -> 2.9 µl
- P195: c = 423 ng/µl -> 2.4 µl
- P196: c = 92 ng/µl -> 10.8 µl
- P197: c = 394 ng/µl -> 2.5 µl
- P198: c = 183 ng/µl -> 5.5 µl
- P200: c = 464 ng/µl -> 2.2 µl
- P236: c = 291 ng/µl -> 3.4 µl
- P263: c = 158 ng/µl -> 6.3 µl
- Positive control: 2.19/4 (pYES2+EGFP) c = 200 ng/µl -> 5µl
100 µg denatured sheared salmon sperm: the one from Simon was used -> 10 µl
step 7:
7 ml 50% PEG3350 was synthesized (3.5 g in 7 ml bidest.water) 6.4 ml 50% PEG3350 + 800 µl 10x LiAc + 800 µl 10x TE were mixed
The cells were plated out on SC-U plates with Glucose and incubated the weekend at 30°C
Transformation of E.coli XL1 blue with ligation products
Investigator: Katrin
Aim: Transformation of ligation products PCR2/P133, PCR1/P133, MK/P133, PCR2/P175, MK/P175
Transformation into E.coli Xl1-Blue
- thawing of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- of 1 µl of the SDM PCR product or 5 µl of ligation product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 45 min
- plate 100 µl on an Amp-plate (pYES (P175) or Chloramphenicol-plate (pSB1C3, P133)
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an antibiotic containing LB-plate
Sunday, August 5th
PCR of pGBKT7 (P99) to introduce RFC pre- and suffix to Gal4 DNA binding domain (Gal4DBD/Gal4BD)
Investigator: Jeff
Aim of the experiment:PCR of pGBKT7 (P99) to introduce RFC pre- and suffix to Gal4 DNA binding domain (Gal4DBD/Gal4BD) for fusion protein construction of N'-PhyB(N-Terminus)-20aaLinker-GalDBD-C'. (If there is time one can direcly clone PhyB in the C-Terminus of Gal4DBD directly.)
Procedure:
- PCR with primer: TUM12-Gal4BD-fw and TUM12-Gal4BD-rv
- Reaction batch:
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 100 µM Forward Primer O73 (TUM12-Gal4BD-fw) |
| 1 µl | 100 µM Reverse Primer O74 (TUM12-Gal4BD-rv) |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA P99 |
| 35.75 µL | ddH2O |
| =50 µL | TOTAL |
- The PCR program was performed after following scheme with following conditions
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 57 °C | 30 s | |
| 68 °C | 30 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | infinite |
Transfer of old transformed E. coli XL-1 blue from old antibiotic plates to new antibiotic plates and of old Y190 S. cerevisiae strain to a new YPD plates
Investigator: Jeff
Aim of the experiment: Transfer of old transformed E. coli XL-1 blue from old antibiotic plates to new antibiotic plates and of old Y190 S. cerevisiae strain to a new YPD plates.
Procedure:
- Transformed E. coli XL1-Blue: With an incolution loop under sterile conditions the bacteria colonies from the plates were transferred by dilution plating on new antibiotic plates. Attempts were made to take all the colonies to have a multiclonal plate.
- The E. coli XL1-Blue plates from these transformations were taken:
| Parts for Phycocyanobilin synthesis: | |||
|---|---|---|---|
| BBa_I15008 in pSB2K3 (KanR): Heme oxygenase | |||
| Parts for N'-SV40NLS-PhyB(NT)-20aaLinker-LexA-C' and N'-PhyB(NT)-20aaLinker-Gal4DBD-C'/N'-SV40NLS-PhyB(NT)-20aaLinker-Gal4DBD-C' construction: | |||
| BBa_K207000 in pSB3K3 (KanR): PhyB(621NT)-GalDBD | BBa_K207001 in pSB1A2 (AmpR): PhyB(621NT) | BBa_K105005 in pSB2K3 (KanR): LexA DNA binding protein | pGBKT7 (KanR): Yeast-two-hybrid vector with C-terminal MCS after Gal4 DNA binding domain (Gal4DBD/Gal4BD) |
| Parts for N'-Gal4AD-linker-Pif3 construction: | |||
| BBa_K365000 in pSB1C3 (CamR): Pif3(100NT) | pGADT7 AD (AmpR): Yeast-two-hybrid vector with C-terminal MCS after Gal4 transcription activation domain (Gal4AD) | ||
| Oligopeptide linkers for linkage of two functional independent protein domains: | |||
| BBa_K365005 in pSB1C3 (CamR): 20 amino acids long linker with RFC25 pre- and suffix | BBa_K243006 in BBa_K157000 (AmpR): 12 amino acids long linker (Gly-Gly-Ser-Gly)x3 | ||
| Synthetical promoter constructs for LexA based Y2H: | |||
| BBa_K165055 in BBa_J63009 (AmpR): LexA binding sites + mCYC + Kozak + YFPx2 + ADH1 terminator | BBa_K165031 in pSB1AK3 (AmpR, KanR): LexA binding sites + mCYC | ||
- Y190 S. cerevisiae: With an incolution loop under sterile conditions the yeast colonies from the plates were transferred by dilution plating on new YPD plates. Attempts were made to take all the colonies to have a multiclonal plate.
| Genotype: | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MATa | ura3-52 | his3-200 | ade2-101 | lys2-801 | trp1-901 | leu2-3, 112 | gal4Δ | gal80Δ | cyhR2 | LYS2 : : GAL1(UAS)-HIS3(TATA)-HIS3 | MEL1 | URA3 : : GAL1UAS-GAL1TATA-lacZ |
| Reporters: | ||||||||||||
| HIS3 | lacZ | MEL1 | ||||||||||
| Transformation markers | ||||||||||||
| trp1 | leu2 | cyhR2 | ||||||||||
Picking from the overnight transformation of ligated P207+PCR39 product in E. coli XL1-Blue
Investigator: Jeff
Aim of the experiment: Picking from the overnight transformation of ligated P207+PCR39 product in E. coli XL1-Blue from the August, 2nd and August, 3rd.
Procedure:
- 4 colonies from August, 2nd and 4 colonies from August, 3rd were taken. Total: 8 colonies.
- Colonies were picked with pipette tips and transferred into a new cell-culture tube with 4 ml of LB-medium and antibiotics. These tubes were put overnight in a 180 rpm cell culture shaker at 37 °C. P207+PCR39 (CamR).
Picking of E. coli XL1-Blue transformed with BBa_K165055
Investigator: Jeff
Aim of the experiment: Picking of E. coli XL1-Blue transformed with BBa_K165055.
Procedure:
- 2 colonies were taken.
- Colonies were picked with pipette tips and transferred into a new cell-culture tube with 4 ml of LB-medium and antibiotics. These tubes were put overnight in a 180 rpm cell culture shaker at 37 °C. BBa_K165055 in BBa_J63009 (AmpR).
Picking of E. coli XL1-Blue transformed with pGADT7 AD
Investigator: Jeff
Aim of the experiment: Picking of E. coli XL1-Blue transformed with pGADT7 AD.
Procedure:
- 2 colonies were taken.
- Colonies were picked with pipette tips and transferred into a new cell-culture tube with 4 ml of LB-medium and antibiotics. These tubes were put overnight in a 180 rpm cell culture shaker at 37 °C. pGADT7 AD (AmpR).
Picking of clones of transformation August, 1st, 2nd and 3rd
Investigator: Jeff
Aim: Picking.
Procedure:
- 3 or 4 clones were picked for each ligation. Clones containing pYES2-new were put into 4 ml LB with 4 µl of ampillicin stock solution, clones containing pSB1C3 were put into chloramphenicol containing media. Incubation at 37 °C overnight.
Monday, August 6th
Miniprep of overnight culture of transformated E. coli XL1-Blue with with ligated P207+PCR39
Investigator: Jeff, Dennis
Aim of the experiment: Miniprep of overnight culture of transformated E. coli XL1-Blue with with ligated P207+PCR39
Procedure:
- Miniprep was done after manufacturer's protocol for 8 picked colonies. (P285-P292)
Analytical digestion and gelelectrophoresis of miniprep P285-P292
Investigator: Jeff, Dennis
Aim of the experiment: Analytical digestion with NgoMIV+PstI-HF and gelelectrophoresis of miniprep P285-P292.
Procedure:
- Mastermix for NgoMIV+PstI-HF
| volume | reagent |
| 22 µl | NEBuffer 4 (10x) |
| 2.75 µl | NgoMIV |
| 2.75 µl | PstI-HF |
| 165 µl | ddH2O |
| =192.5 µl | TOTAL |
- Reaction batch
| volume | reagent |
| 2,5 µl | Plasmid-DNA (P285-P292) |
| 17,5 µl | Mastermix for NgoMIV+PstI-HF |
| =20 µl | TOTAL |
| 100 bp ladder DNA ladder | P285 | P286 | P287 | P288 | P289 | P290 | P291 | P292 | P295 | P296 | 1 kbp ladder DNA ladder |
| Corrupt | Ligation successful | Ligation successful | Ligation successful | Ligation successful | Ligation successful | Ligation successful | Ligation successful | please see next subsection | plase see next subsection |
Analytical digestion and gelelectrophoresis of miniprep P295, P296
Investigator: Jeff, Dennis
Aim of the experiment: Analytical digestion with XbaI+PstI-HF and gelelectrophoresis of miniprep P295 and P296.
Procedure:
- Digestion reaction batch
| volume | reagent |
| 2.5 µl | Plasmid-DNA (P295, P296) |
| 0.25 µl | XbaI |
| 0.25 µl | PstI-HF |
| 2 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 14.8 µl | ddH2O |
| =20 µl | TOTAL |
| 100 bp ladder DNA ladder | P285 | P286 | P287 | P288 | P289 | P290 | P291 | P292 | P295 | P296 | 1 kbp ladder DNA ladder |
| Please see last subsection | Please see last subsection | Please see last subsection | Please see last subsection | Please see last subsection | Please see last subsection | Please see last subsection | Please see last subsection | corrupt | corrupt |
Analytical digestion and gelelectrophoresis of miniprep P293, P294
Investigator: Jeff, Dennis
Aim of the experiment: Analytical digestion and gelelectrophoresis of miniprep P2P93, P294 to see whether the enzymes EcoRI and BamHI are working. Because the size of the insert is too short, the backbone is additionally cut with Eco91I (cuts only 1x in the backbone). So digestion were done with EcoRI+Eco91I and BamHI+Eco91I.
Procedure
- Reaction batch for P293:
| volume | reagent |
| 2.5 µl | Plasmid-DNA (P293) |
| 0.25 µl | EcoRI (Fermentas) |
| 0.25 µl | Eco91I (Fermentas) |
| 2 µl | Buffer 0 (Fermentas) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Reaction batch for P294:
| volume | reagent |
| 2.5 µl | Plasmid-DNA (P293) |
| 0.5 µl | BamHI (Fermentas) |
| 0.25 µl | Eco91I (Fermentas) |
| 4 µl | Tango-Buffer (10x) (Fermentas) |
| 12.75 µl | ddH2O |
| =20 µl | TOTAL |
| 100 bp ladder DNA ladder | P293 (digested with EcoRI+Eco91I) | P294 (digested with BamHI+Eco91I) | PCR48 | PCR49 | P230 | P231 | 1 kbp ladder DNA ladder |
| 2 DNA strands are visible and have expected length | 2 DNA strands are visible and have expected length | Please see next subsection | Please see next subsection | Please see next subsection | Please see next subsection |
Analytical digestion and gelelectrophoresis of miniprep P230 and P232
Investigator: Jeff, Dennis
Aim of the experiment: Analytical digestion and gelelectrophoresis of miniprep P230 and P232 to check whether the oligohybrdization product PCR34 has been successful ligated to pSB1C3 with RFC25 pre- and suffix.
Procedure:
- Reaction batch for P230 and P231:
| volume | reagent |
| 2,5 µl | Plasmid-DNA (P285-P292) |
| 17,5 µl | Mastermix for NgoMIV+PstI-HF (please see subsection: "Analytical digestion and gelelectrophoresis of miniprep P285-P292 ") |
| =20 µl | TOTAL |
| 100 bp ladder DNA ladder | P293 | P294 | PCR48 | PCR49 | P230 | P231 | 1 kbp ladder DNA ladder |
| Please see next subsection | Please see next subsection | Please see next subsection | Please see next subsection | 91 bp long insert seems to be visible after manual gain of signal | 91 bp long insert seems to be visible after manual gain of signal |
- To be sure, one has to sequence this with the iGEM primer VF2 (O68) and VR (O69).
Analytical gelelectrophoresis of PCR48 and PCR49
Investigator: Jeff, Dennis
Aim of the experiment: Analytical gelelectrophoresis of PCR48 and PCR49 to check whether the PCR has been successful.
Procedure:
| 100 bp ladder DNA ladder | P293 | P294 | PCR48 | PCR49 | P230 | P231 | 1 kbp ladder DNA ladder |
| Please see next subsection | Please see next subsection | PCR was successful | PCR was successful | Please see next subsection | Please see next subsection |
Preperative digestion of P289
Investigator: Jeff
Aim of the experiment: Preperative digestion and gelelectrophoresis of P289 in order to fuse insert (LexA with RFC25 pre- and suffix) into the N-terminus of 20aa linker in pSB1C3 (P206). Intermediate construction: N'-20aaLinker-LexA-C'. Final construction should be N'-SV40NLS-PhyB(642NT/908NT)-20aaLinker-LexA-C'.
Procedure:
- Reaction batch:
| volume | reagent |
| 20 µl | Plasmid 289 |
| 4 µl | NEBuffer 4 (10x) |
| 2 µl | NgoMIV (10 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 11 µl | ddH2O |
| =40 µl | TOTAL |
- Preperative gelelectrophoresis was done the next day
Extraction of ADH1-Promoter, CYC1-Terminator, TEF1-Promoter
Investigator: Georg
Aim of the experiment: Miniprep of overnight cultures of transformed E. coli XL1-Blue
Procedure:
- Miniprep was done after manufactuer's protocol for 3x3 colonies.
PCR of TEF1-Terminator, TEF2-Promoter and ADH1-Terminator
Investigator: Georg
Aim of the experiment: Amplification of TEF1-Terminator, TEF2-Promoter from genomic DNA of Saccharomyces cerevisiae and ADH1-Terminator from yeast Vector PGADT7 AD
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer |
| 1 µl | 10 µM Reverse Primer |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Final: 1.25 units/50 µl) |
| 1,3µl P237/ 1µl P98 | |
| 33.75 µl | dd water |
| 50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 56 °C | 1 min | |
| 68 °C | 2 min | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | 1 h |
Preparative digestion of PCR products and TEF1-,ADH1-Promoter with XbaI and PstI
Investigator: Georg
- Mastermix for XbaI+PstI-HF
| volume | reagent |
| 12 µl | NEBuffer 4 (10x) |
| 3 µl | XbaI |
| 3 µl | PstI-HF |
| 1,2 µl | BSA 100x |
| 20,8 µl | ddH2O |
| =40 µl | TOTAL |
- Mastermix for XbaI+PstI-HF
| volume | reagent |
| 15 µl | NEBuffer 4 (10x) |
| 3 µl | XbaI |
| 3 µl | PstI-HF |
| 1,5 µl | BSA 100x |
| 27,5 µl | ddH2O |
| =50 µl | TOTAL |
Ligation of lavendula LS and citrus LS with pSB1C3 and pYES
Procedure
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| PCR 42 RL | 0.8 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 15.7 µl |
| TOTAL | =20 µl |
Miniprep of Ligation of PCR1 and PCR2 in P133 or P175
Investigator: Andrea
Using a Quiagen Miniprep kit. The miniprep products were named P266-P284.
The resulting plamid-concentrations (measured with nanodrop) were
- P 266: 1,3 ng/µl
- P 267: 0,2 ng/µl
- P 268: 1,0 ng/µl
- P 269: 13,5 ng/µl
- P 270: 10,3 ng/µl
- P 271: 12,1 ng/µl
- P 272: 14,7 ng/µl
- P 273: 12,8 ng/µl
- P 274: 13,2 ng/µl
- P 275: 9,4 ng/µl
- P 276: 8,5 ng/µl
- P 277: 12,4 ng/µl
- P 278: 10,2 ng/µl
- P 279: 7,1 ng/µl
- P 280: 5,7 ng/µl
- P 281: 8,0 ng/µl
- P 282: 8,6 ng/µl
- P 283: 31,4 ng/µl
- P 284: 12,6 ng/µl
PCR of Schwab plasmid Quickchange-DNA to amplify gene for lavendula LS
Instructor: Andrea
Aim: PCR of Schwab plasmids after Quickchange-Mutagenesis with primers which were designed to amplify the lavendula LS gene and to add RFC25 restriction sites.
3 different primer combinations were used:
1. O33/O37
2. O34/O37
3. O35/O37
Each primer combination was used for plasmid DNA amplification of P283.
PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer |
| 1 µl | 10 µM Reverse Primer |
| 0.25 µl | OneTaq Hot Start DNA Polymerase (Final: 1.25 units/50 µl) |
| 3 µl | Plasmid DNA P283 |
| 33.75 µl | dd water |
| 50 µL | TOTAL |
- The PCR program was performed after following scheme:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 50 °C | 30 s | |
| 68 °C | 1,75 min | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | 1 h |
Preparation of SC-U medium with 2% glucose (300ml) or galactose (700 ml) for expression in Saccharomyces cerevisiae
Investigator: Daniela
1 l medium:
- One alliquot (50 ml) of amino acis prepared by Alois Bräuer was used
- 6.7 g yeast nitrogen base
- 850 ml ELGA water
- after dissolving, the solution was divided into 270 ml and 630 ml
- autoclaving
Sugar solution were prepared separately:
- 6 g glucose were dissolved in 30 ml ELGA water
- 14 g galactose were dissolved in 70 ml ELGA water
- autoclaving
After autoclaving:
- 30 ml of glucose solution were added to 270 ml SC-U medium
- 70 ml of galactose solution were added to 630 ml SC-U medium
Inoculation of Saccharomyces cerevisiae colonies of transformation products from August, 3rd
Investigator: Daniela
Aim of the experiment: First step of the expression protocol in Saccharomyces cerevisiae
Inoculation of a single colony of Saccaromyces cerevisiae transformed with enzymes PAL+/-, 4CL+/-, CHS+/-, OMT+/-, APT and the positive control GFP in pYES, respectively in 15 ml of SC-U medium with 2% glucose. Incubation at 30°C over night.
Tuesday, 7th august
Plating of received E. coli containing biobricks BBa_K181000, BBa_K365002 and BBa_K365003
Investigator: Jeff
Aim of the experiment: The received biobricks were already transformed in E. coli and were in an agar stabs. These E. coli cells were transferred with an inoculation loop on antibiotic selection plates and were incubated over night. The received biobricks were: BBa_K181000 (This is the PcyA gene, known to convert biliverdin to phycocyanobilin, codon optimized for S. cerevisiae (baker's yeast). ), BBa_K365002 (This part consists in the first 908 amino-acids of Phytochrome B from A. thaliana.) and BBa_K365003 (This part consists in the first 642 amino-acids of Phytochrome B from A. thaliana.).
Procedure:
- Bacterias containing plasmids with biobricks were transferred with a sterile inoculation loop on antibiotic plates and were incubated at 37 °C overnight.
- The biobricks were: BBa_K181000 in BBa_J63010 (AmpR), BBa_K365002 in pSB1C3 (CamR) and BBa_K365003 in pSB1C3.
Preperative gelelectrophrosis of digested P289
Investigator: Jeff, Dennis
Aim of the experiment: Preperative digestion and gelelectrophoresis of P289 in order to fuse insert (LexA with RFC25 pre- and suffix) into the N-terminus of 20aa linker in pSB1C3 (P206). Intermediate construction: N'-20aaLinker-LexA-C'. Final construction should be N'-SV40NLS-PhyB(642NT/908NT)-20aaLinker-LexA-C'.
Procedure:
- Gelelectorphoresis was performed on a 1% low-melting agarose gel at 70 for 1.5 h.
- DNA strand at about 600-700 bp was cut out
- Cut out band was purified with gel extraction kit from Qiagen after manufacturer's protocol.
Ligation of P206+P310
Investigator: Jeff, Dennis
Aim of the experiment: Ligation of P206+P310
Procuedure: Reaction batch
| Substance | Volume |
| P206 | 1.7 µl (~100 ng vector DNA) |
| P310 | 8.5 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 7.3 µl |
| TOTAL | =20 µl |
Preperative digestion and gelelectrophrosis of P123 and P80
Investigator: Jeff
Aim of the experiment: Preperative digestion and gelelectrophoresis of P80 and P123 in order to fuse insert (Pif3 with RFC25 pre- and suffix) of P80 into the C-terminus of 20aa linker in pSB1C3 (P123). Intermediate construction: N'-Pif3-20aaLinker-C'. Final construction should be N'-Pif3-20aaLinker-Gal4AD-C'.
Procedure:
- Reaction batch for P123:
| volume | reagent |
| 20 µl | Plasmid-DNA P123 |
| 4 µl | NEBuffer 4 (10x) |
| 1 µl | EcoRI-HF (20 U/µl) |
| 2 µl | NgoMIV (10 U/µl) |
| 13 µl | ddH2O |
| =40 µl | TOTAL |
- Reaction batch for P80:
| volume | reagent |
| 20 µl | Plasmid-DNA P80 |
| 4 µl | NEBuffer 4 (10x) |
| 0.44 µl | BSA (100x) |
| 1 µl | EcoRI-HF (20 U/µl) |
| 1 µl | AgeI (20 U/µl) |
| 13.6 µl | ddH2O |
| =40 µl | TOTAL |
- Gelelectorphoresis was performed on a 1% low-melting agarose gel at 70 for 1.5 h.
- DNA strand at about 300 bp from P80 was cut out. And the band from P123 at about 2.1 kbp was cut out.
- Cut out bands were purified with gel extraction kit from Qiagen after manufacturer's protocol.
| P80 digested with EcoRI-HF+AgeI-HF | 100 bp ladder DNA ladder | P123 digested with EcoRI-HF+NgoMIV | 1 kbp ladder DNA ladder |
| Gel OK | Gel OK |
Ligation of P311+P312
Investigator: Jeff
Aim of the experiment: Ligation of P311+P312
Procuedure: Reaction batch
| Substance | Volume |
| P312 | 1.92 µl (~100 ng vector DNA) |
| P311 | 6.38 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 9.2 µl |
| TOTAL | =20 µl |
Picking of received E. coli containing biobricks BBa_K181000, BBa_K365002 and BBa_K365003
Investigator: Jeff
Aim of the experiment: Picking of received E. coli containing biobricks BBa_K181000, BBa_K365002 and BBa_K365003.
Procedure:
- For each biobrick 1 coloniy was taken with a pipette tip and transferred into a cell-culture tube with 4 of LB-medium and antibiotics (CamR for BBa_K365002 and BBa_K365003, AmpR for BBa_K181000).
- These 3 tubes were put overnight in a 180 rpm cell culture shaker at 37 °C.
Preparative restriction digest of PCR 56,57,58 (lavendula LS) and PCR 42,43 (citrus LS)
Investigator: Lara
Aim: Prepare digested PCR fragments for further ligation into pYES and pSB1C3.
| volume | reagent |
| 25 µl | PCR products |
| 5 µl | NEBuffer 4 (10x) |
| 0.5 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | AgeI-HF (20 U/µl) |
| 17.5 µl | ddH2O |
| =50 µl | TOTAL |
- A mastermix for 16 reactions was produced.
- PCR products were digested at 37 °C for 3 hours.
P187 was used as positive control for restriction digest:
- 3 µl p187 + 15 µl of Mastermix (see above)
Gelelectrophoresis:
Gel 1:
1. Gene ruler 1 kb
2. PCR 42
3. PCR 43
5. P187
Gel 2:
1. Gene ruler 1 kb
2. PCR 56
3. PCR 57
4. PCR 58
Digested PCR products (PCR 42 RL, PCR 43 RL, PCR 56 RL, PCR 57 RL, PCR 58 RL) are stored in the iGEM PCR product box at -20°C.
- RL = ready for ligation
Analytical restriction digest of ligation products after miniprep
Investigator: Lara
Aim: Check ligation of PCR 1 and PCR 2 into pYES and pSB1C3.
| volume | reagent |
| 20 µl | miniprep products |
| 2.3 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 0.25 µl | XbaI (20 U/µl) |
| 0.25 µl | Pst1-HF (20 U/µl) |
| =23 µl | TOTAL |
- Incubation at 37°C for 2 h.
Gelelectrophoresis:
Ligation of lavendula LS and citrus LS with pSB1C3 and pYES
Investigator: Lara
Aim: Ligate both lavendula and citrus limonene synthase (with all primer combinations -> with or without consensus sequence) with pSB1C3 and pYES.
Procedure
PCR 42 RL in pYES (p175)
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| PCR 42 RL | 0.8 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 15.7 µl |
| TOTAL | =20 µl |
PCR 42 RL in pSB1C3 (p133)
| Substance | Volume |
| P133 | 2.5 µl (~100 ng vector dna) |
| PCR 42 RL | 2.5 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 12.5 µl |
| TOTAL | =20 µl |
PCR 43 RL in pYES (p175)
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| PCR 43 RL | 1.5 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 15.0 µl |
| TOTAL | =20 µl |
PCR 43 RL in pSB1C3 (p133)
| Substance | Volume |
| P133 | 2.5 µl (~100 ng vector dna) |
| PCR 43 RL | 4 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 11 µl |
| TOTAL | =20 µl |
PCR 56 RL in pYES (p175)
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| PCR 56 RL | 2 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 14.5 µl |
| TOTAL | =20 µl |
PCR 56 RL in pSB1C3 (p133)
| Substance | Volume |
| P133 | 2.5 µl (~100 ng vector dna) |
| PCR 56 RL | 6 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 9 µl |
| TOTAL | =20 µl |
PCR 57 RL in pYES (p175)
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| PCR 57 RL | 2 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 14.5 µl |
| TOTAL | =20 µl |
PCR 57 RL in pSB1C3 (p133)
| Substance | Volume |
| P133 | 2.5 µl (~100 ng vector dna) |
| PCR 57 RL | 6 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 9 µl |
| TOTAL | =20 µl |
PCR 58 RL in pYES (p175)
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| PCR 58 RL | 2 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 14.5 µl |
| TOTAL | =20 µl |
PCR 58 RL in pSB1C3 (p133)
| Substance | Volume |
| P133 | 2.5 µl (~100 ng vector dna) |
| PCR 58 RL | 6 µl (~3x of n(vector)) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 9 µl |
| TOTAL | =20 µl |
Negative control/pYES (p175)
| Substance | Volume |
| P175 | 1 µl (~100 ng vector dna) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 16.5 µl |
| TOTAL | =20 µl |
Negative control/pSB1C3 (p133)
| Substance | Volume |
| P133 | 2.5 µl (~100 ng vector dna) |
| T4 ligase | 0.5 |
| T4 ligase buffer | 2 µl |
| ddH2O | 15 µl |
| TOTAL | =20 µl |
- ligation over night at 16 °C (waterbath).
Eppis with Ligation products are stored in two 50 ml Falcon tubes at -20°C (lowest drawer)!
Induction of PAL,4CL,CHS,OMT,APT and GFP in pYES in S. cerevisiae in SC-U medium with 2% galactose
Investigator: Mary, Daniela
Aim of the experiment: Expression of all enzymes in Saccharomyces cerevisiae.
| enzyme | OD600 | volume to be pelleted [ml] |
| PAL+ | 3.246 | 6.2 |
| PAL- | 3.604 | 5.5 |
| 4CL+ | 3.875 | 5.2 |
| 4CL- | 4.0 | 5.0 |
| CHS+ | 3.685 | 5.43 |
| CHS- | 6.68 | 5.43 |
| OMT+ | 3.795 | 5.27 |
| OMT- | 4.085 | 4.896 |
| APT | 3.415 | 5.86 |
| GFP | 3.855 | 5.188 |
Incubation at 30 °C over night.
Preparation of solutions for generating chemo competent E.Coli- cells
Investigator: Roman
Procedure:
Required solutions (for a 24ml suspension of competent cells)
- 0,1 M MgCl2
- 0,05 M CaCl2
- 0,05 M CaCl2, 15% v/v glycerine
Prepared volumes:
- 30 ml CaCl2, 15% v/v glycerine; 0,22g CaCl2 were mixed with 4,5 ml glycerine and 25,5 ml bidestillated water and autoclaved
- 500 ml MgCl2; 10,17g MgCl2 were mixed with 500 ml bidestillated water and autoclaved
- 250 ml CaCl2; 1,84g CaCl2 were mixed with 250 ml bidestillated water and autoclaved
Preparation of competent cells see next day.
Wednesday, August 8th
Miniprep of overnight culture of E. coli containing biobricks BBa_K181000, BBa_K365002 and BBa_K365003
Investigator: Jeff, Dennis
Aim of the experiment: Miniprep of overnight culture of E. coli containing biobricks BBa_K181000, BBa_K365002 and BBa_K365003.
Procedure:
- Miniprep with Kit from Qiagen after manufacturer's protocol
Preperative digestion and gelelectrophoresis of P27 and P315
Investigator: Jeff, Dennis
Aim of the experiment:
Procedure:
- Reaction batch for P27:
| volume | reagent |
| 25 µl | Plasmid-DNA P27 |
| 4 µl | NEBuffer 4 (10x) |
| 0.4 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 8.6 µl | ddH2O |
| =40 µl | TOTAL |
- Reaction batch for P315:
| volume | reagent |
| 25 µl | Plasmid-DNA P27 |
| 4 µl | NEBuffer 4 (10x) |
| 0.4 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 8.6 µl | ddH2O |
| =40 µl | TOTAL |
| P27 digested with XbaI+PstI-HF BBa_I15008: Heme oxygenase 1 (HO1) | 100 bp DNA ladder | P315 digested with XbaI+PstI-HF BBa_K181000: PcyA | 1 kbp DNA ladder |
| Backbone and insert correct | Backbone and insert correct |
Analytical digestion and gelelectrophoresis of P313 BBa_K365002: PhyB(908NT)and P314 BBa_K365003 : PhyB(642NT)
Investigator: Jeff, Dennis
Aim of the experiment:
Procedure:
- Reaction batch for P313 (BBa_K365002):
| volume | reagent |
| 2.5 µl | Plasmid-DNA P313 |
| 2 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 0.25 µl | XbaI (20 U/µl) |
| 0.25 µl | PstI-HF (20 U/µl) |
| 14.8 µl | ddH2O |
| =20 µl | TOTAL |
- Reaction batch for P314 (BBa_K365003):
| volume | reagent |
| 2.5 µl | Plasmid-DNA P313 |
| 2 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 0.25 µl | XbaI (20 U/µl) |
| 0.25 µl | PstI-HF (20 U/µl) |
| 14.8 µl | ddH2O |
| =20 µl | TOTAL |
| 100 bp DNA ladder | P313 digested with XbaI+PstI-HF (BBa_K365002: PhyB(908NT) | P314 digested with XbaI+PstI-HF (BBa_K365003: PhyB(642NT) | 1 kbp DNA ladder |
| Backbone and insert correct | Backbone and insert correct |
Transformation of E. coli XL1-Blue with ligation products P206+P310 and P312+P311
Investigator: Jeff
Aim of the experiment: Transformation of E. coli XL1-Blue with ligation products P206+P310 (N'-20aaLinker-LexA-C' construction) and P312+P311 (N'-Pif3-20aaLinker-C')
Procedure:
- Transformation was performed after standard transformation protocol from the lab.
- Transformated E. coli XL1-Blue cells were plated on antibiotic plates and were put at 37 °C overnight.
- P206+P310 (CamR), P312+P311 (CamR)
Picking of E. coli XL1-Blue colonies containing BBa_K365000
Investigator: Jeff
Aim of the experiment: Picking of E. coli XL1-Blue colonies containing BBa_K365000 to a miniprep on the nextday from the overnight culture.
Procedure:
- 3 colonies were picked with pipette tips and transferred into a new cell-culture tube with 4 ml of LB-medium and chloramphenicol. These tubes were put overnight in a 180 rpm cell culture shaker at 37 °C.
Miniprep of ligation products of transformation from August 3rd
Investigator: Lara
Aim: Get transformed plasmids for further analytical restriction digest to check whether ligation worked.
Procedure: Miniprep was done using Qiagen miniprep kit.
- PCR1/p175 clone 1: 200 ng/µl
- PCR1/p175 clone 2: 260 ng/µl
- PCR1/p175 clone 3: 460 ng/µl
- PCR1/p175 clone 4: 450 ng/µl
- PCR2/p175: 222 ng/µl
- PCR2/p133 clone 1: 145 ng/µl
- PCR2/p133 clone 2: 150 ng/µl
- PCR2/p133 clone 3: 140 ng/µl
- PCR2/p133 clone 4: 510 ng/µl
Eppis containing miniprep products were stored in 50 ml Falcon tubes at the lowest drawer of -20°C.
--> will be thrown away in case ligation hasn't worked.
Analytical restriction digest of miniprep products
Investigator: Lara
Aim: Analytical restriction digest to check whether ligation of Tuesday, July 31st has worked.
| volume | reagent |
| 2.5 µl | plasmid DNA |
| 0.25 µl | Xba1 |
| 0.25 µl | Spe1 |
| 2 µl | NEBuffer 4 |
| 0,2 µl | BSA (100x) |
| 14.8 µl | ddH2O |
- Incubation for 1.5 h at 37°C.
Analytical Gel
1. Standard (1 kb Gene Ruler)
2. PCR1/p175 clone 1
3. PCR1/p175 clone 2
4. PCR1/p175 clone 4
5. PCR1/p175 clone 3
6. PCR2/p175
7. PCR2/p133 clone 1
8. PCR2/p133 clone 2
9. PCR2/p133 clone 3
10.PCR2/p133 clone 4 --> successful!
Transformation of ligation products from ligation of August 7th
Investigator: Lara
Aim: Transform E.coli XL 1 blue with ligation products to produce colonies containing plasmid DNA.
Transformation into E.coli Xl1-Blue
- thawing of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- adding of 5 µl of ligation products
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- adding of 1 ml LB-medium without antibiotics to the cells and incubation at 37°C and 180 rpm for 60 min
- plating of 100 µl of the cells on an Amp-plate (pYES (P175) or Chloramphenicol-plate (pSB1C3, P133)
- sedimentation of the leftover in a centrifuge (30 - 60 sec, 13 000 rpm), resuspension of the sediment in 100 µl LB-medium and plating it as well on an antibiotic containing LB-plate
Cell lysis of S. cerevisiae
Investigator: Mary,Daniela
Aim of the experiment:Receive (hopefully) expressed enzymes.
The protocol 'Expression Hefe' (Dropbox) was used. If you want to repeat the experiment please read comments before you start.
You should make a glycerol stock of clones picked. You should as well take a sample of the uninduced yeast and do a cell lysis to be able to compare the induced and uninduced status.
Preparation of sodium phosphate buffer (50 mM):
- 1 M Na2HPO4x2H2O (M = 177.99 g/mol): 1.7799 g dissolved in 10 ml ELGA water
- 1 M NaH2PO4x2H2O (M = 156.01 g/mol): 0.468 g dissolved in 3 ml ELGA water
- 8.095 ml Na2HPO4 and 1.905 ml NaH2PO4 and 190 ml ELGA water were mixed -> pH should be 7.5 but in our case it wasn't. Therefore we used all of the Na2HPO4, but could only achieve an pH of 7.47.
Preparation of PMSF (100 mM)->solution is in the fridge:
- 0.0174 g were dissolved in 1 ml isopropanol (techn.)
Preparation of breaking buffer:
- 11.25 ml sodium phosphate buffer (50 mM)
- 750 µl Glycerol (80%)
- 24 µl EDTA
Preparation of breaking buffer with PMSF:
- 6 ml of breaking buffer
- 60 µl of PMSF (100 mM)
Preparations for SDS gel the next day:
The absorption A280 was measured with Nano Drop. The SDS gel should be loaded with 30 µg protein, each assay was filled up with buffer to a total volume of 10 µl. 2.5 µl 5xLaemmlired were added and all assays were denatured at 95 °C for 5 min.
| enzyme | concentration [mg/ml] (A280) | concentration of 1:10 dilution [mg/ml] (A280) | volume of 1:10 dilution with 30 µg protein [µl] |
| PAL+ | 64.024 | 7.946 | 3.78 |
| PAL- | 61.140 | 8.605 | 3.486 |
| 4CL+ | 85.65 | 9.459 | 3.17 |
| 4CL- | 54.166 | 7.443 | not applied on the gel |
| CHS+ | 65.338 | 6.045 | 4.96 |
| CHS- | 51.167 | 6.135 | 4.89 |
| OMT+ | 49.128 | 5.453 | 5.5 |
| OMT- | 62.074 | 4.193 | 7.15 |
| APT | 42.554 | 4.787 | 6.27 |
| GFP | 36.443 | 4.247 | 7.06 |
Preparation chemocompetent E.Coli- cells
Investigator: Roman, Simon
Operational sequence:
The competent cells were prepared as described in the protocoll:
- 600ml LB medium were inoculated with 6ml of an E.Coli XL1 blue overnight culture and grown to an OD550 of 0,67
- centrifugation and washing of the cells as described in the protocol
- cells were aliquoted as 100µl suspensions in the cold room
- storage at -80°C
Thursday, August 9th
PCR amplification of a fragment of P313
Investigator: Roman
Aim of experiment: Amplification of wanted fragment
Operational Sequence:
PCR- reaction batch was prepared as follows:
| substrate | volume in µl |
| ddH2O | 35,75 |
| One Taq Standard Reaction Buffer (5x) | 10 |
| Plasmid template (p313) | 1 |
| Primer O70 (10 µM) | 1 |
| Primer O72 (10 µM) | 1 |
| dNTP (10 mM) | 1 |
| Polymerase One Taq | 0,25 |
Negative control was prepared the same way, with 1 µl ddH2O instead of plasmid template.
PCR temperature program:
- initial heating: 94°C for 30s
- 30 cycles
- denaturation: 94°C for 30s
- annealing: 67°C for 60s
- elongation: 68°C for 3 min
- final elongation: 68°C for 5 min
- 4°C
Afterwards, the PCR reaction tubes were stored over night in the fridge at 4 °C
Miniprep of picked E. coli XL1-Blue containing BBa_K365000
Investigator: Dennis
Aim of the experiment: Miniprep of picked E. coli XL1-Blue containing BBa_K365000. From those minipreps PCRs should be done to introduce RFC25 pre- and suffixes and MfeI and BamHI restriction sites.
Procedure:
- Miniprep was performed with Qiaprep Spin miniprep kit from Qiagen.
- Miniprep was performed after manufacturer's protocol.
Preperative digestion and gelelectrophoresis of P324
Investigator: Dennis
Aim of the experiment: Preperative digestion and gelelectrophoresis of P324 for fusion protein construction the plasmid containing the phytochrome interacting factor 3 including the RFC25 pre- and suffix was digested with NgoMIV and PstI-HF.
Procedure:
- Reaction batch:
| volume | reagent |
| 20 µl | Plasmid-DNA Pxxx |
| 4 µl | NEBuffer 4 (10x) |
| 2 µl | NgoMIV (10 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 13 µl | ddH2O |
| =40 µl | TOTAL |
- After 2-3 h of incubation time at 37 °C the reaction mix was mixed with 4.44 µl DNA running buffer (10x) and was pipetted into a gelpocket of an 1% low-melting agarose gel.
- The gel was running for about 1 at 80 V
- Then the gel extraction kit from Qiagen was used for extracting the linearized insert from the agarose gel.
Picking from the overnight transformation of ligated P206+P310 product and P312+P311 product in E. coli XL1-Blue
Investigator: Dennis
Aim of the experiment: Picking from the overnight transformation of ligated P206+P310 product and P312+P311 product in E. coli XL1-Blue from August, the 8th.
Procedure:
- 5 colonies from P206+P310 transformated cells and 5 colonies from P312+P311 transformated cells were picked.
- Colonies were picked with pipette tips and transferred into a new cell-culture tube with 4 ml of LB-medium and chloramphenicol. These tubes were put overnight in a 180 rpm cell culture shaker at 37 °C.
Picking of colonies of transformation (August 8th)
Investigator: Lara
Aim: Get cells for miniprep for further analytical restriction digest to check whether ligation has worked.
Procedure: Picking of 3 clones each of PCR42/p175, PCR43/p175, PCR56/p175, PCR57/p175, PCR58/p175 into 4 ml LB with Amp and PCR42/p133, PCR43/p133, PCR56/p133, PCR57/p133, PCR58/p133 into 4 ml with Chlp. Inbucation at 37°C (shaker) over night.
SDS-PAGE of crude protein extract after induced expression of PAL, 4CL, CHS, OMT, APT and GFP in yeast
Investigator: Daniela
Aim of the experiment: Getting the right concentration of protein for the western blot.
The preparation of the samples was already described on August, 8th.
- 12 % gel
- 6 µl Unstained Protein MW Marker (Thermo Scientific)
- 12.5 µl sample
Mass of enzymes:
- PAL = 76.880 kDa
- 4CL = 61.053 kDa
- CHS = 42.713 kDa
- OMT = 39.265 kDa
- APT = 45.481 kDa
The enzymes of the coumaroyl pathway were not detectable within the other proteins. However the gel looks good, therefore the same concentrations were used for the western blot.
Western Blot of PAL, 4CL, CHS, OMT, APT and GFP expressed in yeast
Investigator: Daniela
Aim of the experiment: Detecion of enzymes via Strep-tag
SDS-PAGE:
The supernatant of the lysis reaction (see August, 8th) was diluted 1:10 with ELGA water. The absorption A280 was measured with Nano Drop. The SDS gel should be loaded with 30 µg protein, each assay was filled up with buffer to a total volume of 10 µl. 2.5 µl 5xLaemmlired were added and all assays were denatured at 95 °C for 5 min.
| enzyme | concentration of 1:10 dilution [mg/ml] (A280) | volume of 1:10 dilution with 30 µg protein [µl] |
| PAL+ | 8.585 | 3.49 |
| PAL- | 7.824 | 3.83 |
| 4CL+ | 10.218 | 2.94 |
| CHS+ | 12.875 | 2.33 |
| CHS- | 7.245 | 4.14 |
| OMT+ | 8.158 | 3.68 |
| OMT- | 5.002 | 6 |
| APT | 5.15 | 5.82 |
| GFP | 4.805 | 6.24 |
- 12 % gel
- 120 V, 1.5 h
Blotting:
- blotting of proteins on a nitrocellulose membrane (Whatman)
- 50mA, 1 h
Blocking:
- wash 3x5min with PBS-T0.1
- the membrane is blocked over night at 4 °C on a shaking device
Retransformations with pSB1C3
Investigator: Roman
Aim of experiment: Aim of the experiment was on the one hand to fill up the stock of usable pSB1C3 for both, the caffeine and the whole team. On the other hand, the experiment was used as a test for previously prepared competent E.Coli cells
Operational sequence: The used vectors for the transformations were pSB1C3 RFC25 (p123), and pSB1C3 containing CHS- from the coumaroyl group (p136). Vector lengths are 2070 bps and xxx bps, respectively).
Transformations:
- Transformation of (old) chemo competent e.coli cells, used up to now, and the newly prepared with p123
- Transformation of the new chemo competent e.coli cells with p136 with two slightly different methods:
- as described in previous protocolls, with an heat shock at 37°C for 5 minutes
- with an heat shock at 42°C for 30 seconds, followed by an incubation on ice for 30 minutes (before the cells being incubated with LB- medium)
The reaction batches were plated out on chloramphenicol containing LB- agar plates (both, unconcentrated and concentrated, i.e. resuspended in ca. 100 µl LB medium). Plates were incubated over night at 37°C.
Friday, August 10th
Analytical gelelectrophoresis of PCR product of P313 to introduce RFC25 pre- and suffix
Investigator: Roman
Aim of the experiment: Analytical gelelectrophoresis of PCR product of P313 to introduce RFC25 pre- and suffix to see wether the PCR was successful.
Procedure:
- The PCR product was first purificated with PCR purification kit from Qiagen
- Analytical gelelectrophoresis was performed at 90 V for ~1 h.
- PCR was successful!
Miniprep of overnight culture of transformated E. coli XL1-Blue with with ligated P206+P310 and ligated P312+P311
Investigator: Dennis, Roman
Aim of the experiment: Miniprep of overnight culture of transformated E. coli XL1-Blue with with ligated P206+P310 and ligated P312+P311.
Procedure:
- Miniprep was done with the Qiagen Qiaprep spin miniprep kit and was performed after manufacturer's protocol.
Transformation analysis and preparation of over- night cultures
Investigator:Roman
Aim of the experiment: Analysis of the transformation with old and new chemocompetent E.Coli- cells and preparation of over night (over weekend, respectively) cultures, to amplificate the vector pSB1C3
Results:
- All plates showed a lot of colonies
- The concentrated plates were quite overgrown, the unconcentrated ones showed single colonies
- no difference could be detected between transformation with 42°C heat shock (followed by 30 min incubation on ice, see previous day) and 37°C heat shock
Newly prepared competent cells seem to work, further investigations (e.g. plating out with different antibiotics), see below.
Operational sequence: Liquid cultures of the following transformants were prepared:
- XL1-blue with p123 (old competent cells used)
- XL1-blue with p123 (new competent cells used)
- XL1- blue with p136
To prepare the over night cultures, 4ml of LB medium containing chloramphenicol (4µl) were inoculated with a pipet- tip and incubated at room temperature over the weekend with 180 rpm.
Plating of newly generated chemocompetent cells with different antibiotics
Investigator: Roman
Aim: Test, if prepared competent cells (untransformed) are sensitive to different antibiotics
Operational sequence:
Used antibiotics:
- ampicillin
- kanamycin
- tetracyclin
- chloramphenicol
100µl of untransformed, melted E.Coli- cells were plated and incubated over the weekend at room temperature. No growth should occure.
Western Blot of PAL, 4CL, CHS, OMT, APT and GFP expressed in yeast - continued
Investigator: Daniela
Aim of the experiment: Detecion of enzymes via Strep-tag
Detection:
- via Streptavidin-AP (1:4000) ->7ml, Streptavidin-AP diluted in PBST0.1
- 1h incubation, be sure that the whole membrane is covered with Streptavidin-AP solution.
Developing:
- 15 ml AP-buffer mixed with 45 µl BCIP (50 mg/ml in DMF) and 7.5 µl NBT (75 mg/ml in 70 % DMF)
- incubation: about 30 min
Expected bands:
PAL: length: 716, mass = 76,880 kDa
4CL: no strep-tag ->negative control (length: 561, mass = 61.053 kDa)
CHS: length: 390, mass = 42.713 kDa
OMT: length: 352, mass = 39.265 kDa
APT: length: 411, mass = 45.481 kDa
GFP: mass: 26.9 kDa ->positive control
Saturday, August 11th
Repetition of SDS-PAGE and Western Blot of crude protein extract after induced expression of PAL, 4CL, CHS, OMT, APT and GFP in yeast
Investigator: Ingmar
Aim of the experiment: Execute the Western Blot using the StrepMab Classic antibody in order to test whether the immunospecifity is better. Furthermore one of the first SDS-PAGEs was done with an increased protein concentration.
Operational sequence
- To test wheather the increased protein concentrations lead to overloaded bonds, two SDS-PAGE gels which have been stained with Coomassie brilliant blue afterwards were run parallely to the ones for the Western blots.
- The preparation of the samples was executed as describes on August, 8th.
- 12 % SDS-PAGE gel, width of the collection gel: 2 mmm
- 6 µl Unstained Protein MW Marker (Thermo Scientific) for the staining with Coomassie brilliant blue and 6 µl Prestained Protein MW Marker (Thermo Scientific)for the western blots.
- 15 µl sample
Mass of enzymes:
- PAL = 76.880 kDa
- 4CL = 61.053 kDa
- CHS = 42.713 kDa
- OMT = 39.265 kDa
- APT = 45.481 kDa
The enzymes of the coumaroyl pathway were not detectable within the other proteins. However the gel looks good, therefore the same concentrations were used for the western blot.
Western Blot of PAL, 4CL, CHS, OMT, APT and GFP expressed in yeast
Investigator: Daniela
Aim of the experiment: Detecion of enzymes via Strep-tag
SDS-PAGE:
The supernatant of the lysis reaction (see August, 8th) was diluted 1:10 with ELGA water. The absorption A280 was measured with Nano Drop. The SDS gel should be loaded with 30 µg protein, each assay was filled up with buffer to a total volume of 10 µl. 2.5 µl 5xLaemmlired were added and all assays were denatured at 95 °C for 5 min.
| enzyme | concentration of 1:10 dilution [mg/ml] (A280) | volume of 1:10 dilution with 30 µg protein [µl] |
| PAL+ | 8.585 | 3.49 |
| PAL- | 7.824 | 3.83 |
| 4CL+ | 10.218 | 2.94 |
| CHS+ | 12.875 | 2.33 |
| CHS- | 7.245 | 4.14 |
| OMT+ | 8.158 | 3.68 |
| OMT- | 5.002 | 6 |
| APT | 5.15 | 5.82 |
| GFP | 4.805 | 6.24 |
- 12 % gel
- 120 V, 1.5 h
Blotting:
- blotting of proteins on a nitrocellulose membrane (Whatman)
- 50mA, 1 h
Blocking:
- wash 3x5min with PBS-T0.1
- the membrane is blocked over night at 4 °C on a shaking device
Sunday, August 12th
Western Blot of PAL, 4CL, CHS, OMT, APT and GFP expressed in yeast - continued
Investigator: Daniela
Aim of the experiment: Detecion of enzymes via Strep-tag
Detection:
- via Streptavidin-AP (1:4000) ->7ml, Streptavidin-AP diluted in PBST0.1
- 1h incubation, be sure that the whole membrane is covered with Streptavidin-AP solution.
Developing:
- 15 ml AP-buffer mixed with 45 µl BCIP (50 mg/ml in DMF) and 7.5 µl NBT (75 mg/ml in 70 % DMF)
- incubation: about 30 min
Expected bands:
PAL: length: 716, mass = 76,880 kDa
4CL: no strep-tag ->negative control (length: 561, mass = 61.053 kDa)
CHS: length: 390, mass = 42.713 kDa
OMT: length: 352, mass = 39.265 kDa
APT: length: 411, mass = 45.481 kDa
GFP: mass: 26.9 kDa ->positive control
Monday, August 13th
Analytical digestion and gelelectrophoresis of the minipreps of transformated E. coli XL1-Blue with ligated P206+P310 products (P327-P331)
Investigator: Jeff
Aim of the experiment: Analytical digestion and gelelectrophoresis of the minipreps of transformated E. coli XL1-Blue with ligated P206+P310. P327-P331 (N'-20aaLinker-LexA-C').
Procedure:
- Mastermix for analytical digestion with NgoMIV+PstI-HF
| volume | reagent |
| 12 µl | NEBuffer 4 (10x) |
| 1.5 µl | NgoMIV |
| 1.5 µl | PstI-HF |
| 90 µl | ddH2O |
| =105 µl | TOTAL |
- 17.5 µl of the mastermix for NgoMIV+PstI-HF was added to 2.5 µl of plasmid DNA of P327, P328, P329, P330, P331.
- Incubation for 2 h at 37 °C.
Order of gel-pockets:
| 100 bp ladder DNA ladder | P327 (digested with NgoMIV+PstI-HF) | P328 (digested with NgoMIV+PstI-HF) | P329 (digested with NgoMIV+PstI-HF) | P330 (digested with NgoMIV+PstI-HF) | P331 (digested with NgoMIV+PstI-HF) | P310 control (before ligation and already digested with NgoMIV+PstI-HF) | 1 kbp ladder DNA ladder |
| Ligation/Fusion was successful! | Ligation/Fusion was successful! | Ligation/Fusion was successful! | Ligation/Fusion was successful! | Ligation/Fusion was successful! | Control OK! |
Analytical digestion and gelelectrophoresis of the minipreps of transformated E. coli XL1-Blue with ligated P312+P311 products (P332-P336)
Investigator: Jeff
Aim of the experiment: Analytical digestion and gelelectrophoresis of the minipreps of transformated E. coli XL1-Blue with ligated P206+P310. P327-P331 (N'-20aaLinker-LexA-C').
Procedure:
- Mastermix for analytical digestion with EcoRI-HF+AgeI-HF
| volume | reagent |
| 12 µl | NEBuffer 4 (10x) |
| 1.2 µl | BSA (100x) |
| 1.5 µl | EcoRI-HF |
| 1.5 µl | AgeI-HF |
| 88.8 µl | ddH2O |
| =105 µl | TOTAL |
- 17.5 µl of the mastermix for EcoRI-HF+AgeI-HF was added to 2.5 µl of plasmid DNA of P332, P333, P334, P335, P336.
- Incubation for 2 h at 37 °C.
Order of gel-pockets:
| 100 bp ladder DNA ladder | P332 (digested with EcoRI-HF+AgeI-HF) | P333 (digested with EcoRI-HF+AgeI-HF) | P334 (digested with EcoRI-HF+AgeI-HF) | P335 (digested with EcoRI-HF+AgeI-HF) | P336 (digested with EcoRI-HF+AgeI-HF) | P311 control (before ligation and already digested with EcoRI-HF+AgeI-HF) | 1 kbp ladder DNA ladder |
| Ligation/Fusion was successful! | Ligation/Fusion was successful! | Ligation/Fusion was successful! | Ligation/Fusion was successful! | Ligation/Fusion was successful! | Control OK! |
Preperative digestion of PCR41 (Pif3 100NT)
Investigator: Jeff
Aim of the experiment: Contruction of N'-SV40NLS-Gal4AD-pGADT7 AD linker-Pif3(100NT)-C'.
Procedure:
- Reaction batch:
| volume | reagent |
| 25 µl | PCR41 |
| 5 µl | Buffer G (10x) |
| 2 µl | BamHI (10 U/µl) |
| 2 µl | MfeI (MunI) (10 U/µl) |
| 16 µl | ddH2O |
| =50 µl | TOTAL |
Preperative digestion of PCR48 (Gal4DBD)
Investigator: Jeff
Aim of the experiment: To backup amplificated Gal4DBD with RFC25 pre- and suffix in pSB1C3.
Procedure:
- Reaction batch:
| volume | reagent |
| 25 µl | PCR48 |
| 5 µl | NEBuffer 4 (10x) |
| 0.5 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 17.5 µl | ddH2O |
| =50 µl | TOTAL |
- WRONG DIGESTION, THERE IS NO PstI-HF SITE IN THE PCR PRODUCT! IT SHOULD HAVE BEEN XbaI+AgeI-HF
Preperative digestion of PCR65 (PhyB 908NT)
Investigator: Jeff
Aim of the experiment: To backup amplificated PhyB (908NT) with RFC25 pre- and suffix in pSB1C3.
Procedure:
- Reaction batch:
| volume | reagent |
| 25 µl | PCR65 |
| 5 µl | NEBuffer 4 (10x) |
| 0.5 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 17.5 µl | ddH2O |
| =50 µl | TOTAL |
- WRONG DIGESTION, THERE IS NO PstI-HF SITE IN THE PCR PRODUCT! IT SHOULD HAVE BEEN XbaI+AgeI-HF
Preperative digestion of P292
Investigator: Jeff
Aim of the experiment: Only the backbone of this part is needed; this part including LexA is used because to see if the double digest work. If it doesn't work, there won't be 2 DNA strands. Backbone was cut out for ligation with SV40NLS (PCR34).
Procudure:
- Reaction batch:
| volume | reagent |
| 20 µl | P292 |
| 4 µl | NEBuffer 4 (10x) |
| 0.4 µl | BSA (100x) |
| 1 µl | XbaI (20 U/µl) |
| 1 µl | PstI-HF (20 U/µl) |
| 13.6 µl | ddH2O |
| =40 µl | TOTAL |
- WRONG DIGESTION! IT SHOULD HAS BEEN XbaI+AgeI-HF! BUT IT CAN BE USED FOR OTHER PURPOSES
Preperative digestion of P293 (Gal4AD)
Investigator: Jeff
Aim of the experiment: Contruction of N'-SV40NLS-Gal4AD-pGADT7 AD linker-Pif3(100NT)-C'.
Procedure:
- Reaction batch:
| volume | reagent |
| 20 µl | P293 |
| 8 µl | Buffer Tango (10x) |
| 4 µl | EcoRI (10 U/µl) |
| 2 µl | BamHI (10 U/µl) |
| 6 µl | ddH2O |
| =40 µl | TOTAL |
Preperative gelelectrophoresis of digested PCR41, PCR48, PCR65
Investigator: Jeff
Aim of the experiment: Preperative gelelectrophoresis of digested PCR41, PCR48, PCR65.
Procedure:
- Preperative gelelectrophoresis was performed at 70 V for 90 min.
- 5.55 µl of DNA loading buffer (10x) was added to each of the digested PCR products.
Order of gel-pockets:
| PCR41 digested MfeI+BamHI | 100 bp ladder DNA ladder | PCR48 digested with XbaI+PstI-HF | 1 kbp ladder DNA ladder | PCR65 digested with XbaI+PstI-HF |
| Okay! | Length OK but wrong digestion, should have been XbaI+AgeI-HF | Length OK but wrong digestion, should have been XbaI+AgeI-HF |
Preperative gelelectrophoresis of digested P292 and P293
Investigator: Jeff
Aim of the experiment: Preperative gelelectrophoresis of digested P292 and P293
Procedure:
- Preperative gelelectrophoresis was performed at 70 V for 90 min.
- 4.44 µl of DNA loading buffer (10x) was added to each of the digested plasmid products.
Order of gel-pockets:
| P292 digested XbaI+PstI-HF | 100 bp ladder DNA ladder | P293 digested with EcoRII+BamHI | 1 kbp ladder DNA ladder |
| Length OK but wrong digestion, should have been XbaI+AgeI-HF | Okay! |
Ligation of P71+P322
Investigator: Jeff
Aim of the experiment: Ligation of P71+P322.
Procedure:
- Reaction batch:
| volume | reagent |
| 4.18 µl | P71 |
| 3.63 µl | P322 |
| 2 µl | T4 ligase buffer (10x) |
| 0.25 µl | T4 ligase (5 U/µl) |
| 9.69 µl | ddH2O |
| =20 µl | TOTAL |
- No negative control because there was no sufficient amount of P71 anymore.
- The ligation was performed at room temperature for 1 h after manufacturer's protocol.
Ligation of P71+P323
Investigator: Jeff
Aim of the experiment: Ligation of P71+P323.
Procedure:
- Reaction batch:
| volume | reagent |
| 4.18 µl | P71 |
| 5.01 µl | P323 |
| 2 µl | T4 ligase buffer (10x) |
| 0.25 µl | T4 ligase (5 U/µl) |
| 8.31 µl | ddH2O |
| =20 µl | TOTAL |
- No negative control because there was no sufficient amount of P71 anymore.
- The ligation was performed at room temperature for 1 h after manufacturer's protocol.
Ligation of P206+P337
Investigator: Jeff
Aim of the experiment: Ligation of P206+P337.
Procedure:
- Reaction batch:
| volume | reagent |
| 1.69 µl | P206 |
| 7.62 µl | P337 |
| 2 µl | T4 ligase buffer (10x) |
| 0.25 µl | T4 ligase (5 U/µl) |
| 8.19 µl | ddH2O |
| =20 µl | TOTAL |
- Negative control was performed with P206 only wihtout insert, i. e. the only difference is that ddH2O was used instead of P337 in the reaction batch.
- The ligation was performed at room temperature for 1 h after manufacturer's protocol.
Ligation of P342+PCR66
Investigator: Jeff
Aim of the experiment: Ligation of P342+PCR66.
Procedure:
- Reaction batch:
| volume | reagent |
| 1.04 µl | P342 |
| 1.92 µl | PCR66 |
| 2 µl | T4 ligase buffer (10x) |
| 0.25 µl | T4 ligase (5 U/µl) |
| 14.54 µl | ddH2O |
| =20 µl | TOTAL |
- Negative control was performed with P342 only wihtout insert, i. e. the only difference is that ddH2O was used instead of PCR66 in the reaction batch.
- The ligation was performed at room temperature for 1 h after manufacturer's protocol.
Transformation of E. coli XL1-Blue with ligation products of P71+P322, P71+P323, P206+P337 and P342+PCR66
Investigator: Jeff
Aim of the experiment: Transformation of E. coli XL1-Blue with ligation products of P71+P322, P71+P323, P206+P337 and P342+PCR66.
Procedure:
- 5 µl of the ligation products (P71+P322, P71+P323, P206+P337 and P342+PCR66) and their negative controls (P206 NK, P324 NK) were added to 100 µl of CaCl2 competent E. coli XL1-Blue cells on ice.
- 30 min incubation on ice.
- 5 min heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of those cell suspension were plated on suitable antiotic plates.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on suitable antbiotic plates.
Analysis of plated, untransformed chemocompetent E.Coli- cells
Investigator: Roman
Aim: To test the prepared chemocompetent E.Coli- cells.
Results:
- Plate with kanamycin: no colonys (as expected)
- Plate with ampicillin: no colonys (as expected)
- Plate with tetracyclin: full (as expected, due to natural resistance)
- Plate with chloramphenicol: 12 colonies (not expected, should be empty)
The plating on a chloramphenicol containing plate will be repeated, at 37°C over night instead of room temperature over the weekend, because perhaps chloramphenicol is degraded by the bacteria after two days
small-Scale Yeast Transformation of pYES without insert
Investigator: Katrin, Ingmar
Aim of the experiment:Transformation of P152 (pYes without insert) in Yeast
The protocol on page 13 of the pYES2 manual was used. To ensure a positive result, two transformations were made.
Only modifications are noted:
step 1: inoculate in 4 ml YPD medium
step 2: OD600 = 5,6 -> to determine a OD600 of 0.4 in 50 ml, 3 ml yeast suspension and 50 ml YPD medium were used.
steps 3 and 4: centrifugation for 5 min, 4 °C
step 6:
1 µg plasmid DNA
- P152: c = 156.8 ng/µl -> 6,4 µl
100 µg denatured sheared salmon sperm: the one from Simon was used -> 10 µl
The cells were plated out on SC-U plates with Glucose and incubated at 30°C
Tuesday, August 14th
Picking from the overnight transformation of ligated P71+P322, P71+P323, P206+P337 and P342+PCR66 product in E. coli XL1-Blue
Investigator: Jeff
Aim of the experiment: Picking from the overnight transformation of ligated P71+P322, P71+P323, P206+P337 and P342+PCR66 product in E. coli XL1-Blue.
Procedure:
- 5 colonies of each transformation from P71+P322 (AmpR), P71+P323 (AmpR), P206+P337 (CamR) and P342+PCR66 (AmpR) transformated cells were picked.
- Colonies were picked with pipette tips and transferred into a new cell-culture tube with 4 ml of LB-medium and antibiotics (P71+P322 (AmpR), P71+P323 (AmpR), P206+P337 (CamR) and P342+PCR66 (AmpR)). These tubes were put overnight in a 180 rpm cell culture shaker at 37 °C.
Reapeat of transformation of E. coli XL1-Blue with ligation products of P71+P322, P71+P323, P206+P337 and P342+PCR66 from August, the 13th
Investigator: Jeff
Aim of the experiment: Reapeat of transformation of E. coli XL1-Blue with ligation products of P71+P322, P71+P323, P206+P337 and P342+PCR66 from August, the 13th. The reason is that the freshly prepared competent cells are contaminated with a foreign plasmid.
Procedure:
- 5 µl of the ligation products (P71+P322, P71+P323, P206+P337 and P342+PCR66) and their negative controls (P206 NK, P324 NK) were added to 100 µl of CaCl2 competent E. coli XL1-Blue cells on ice.
- 30 min incubation on ice.
- 5 min heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of those cell suspension were plated on suitable antiotic plates.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on suitable antbiotic plates. Accidently, only the pellet of P342+PCR66 and P342 NK was plated.
Analytical restriction digest of ligation products of 10.08. after miniprep
Investigator: Andrea
Aim: Check ligation of PCR 42 and PCR43 (Citrus) & PCR 56, PCR 57 and PCR 58 (Lavendula) into pYES and pSB1C3.
| volume | reagent |
| 20 µl | miniprep products |
| 2 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 0.25 µl | XbaI (20 U/µl) |
| 0.25 µl | Pst1-HF (20 U/µl) |
| =20 µl | TOTAL |
- Master-Mix: 7 µl Enzyme, 56 µl NEB-Buffer, 5,6 µl BSA, 414,4 µl ddH2O
- 2,5 µl DNA added to 17,5 µl Master-Mix
- Incubation at 37°C for 1,5 h.
Gelelectrophoresis:
Preparation of material for protein expression in yeast
Investigator: Ingmar, Katrin
Aim of the experiment:Preparation of media for protein expression in yeast (expression will be done on thursday, august 16th)
the following media were made:
SC-U medium (synthetic complete medium, without uracil)
- 0.67% yeast nitrogen base (without amino acids)
- 2% carbon source (i.e. glucose or raffinose)
- 0.01% (adenine, arginine, cysteine, leuine, lysine, threonine, tryptophan)
- 0.005% (aspartic acid, histidine, isoleucine, methionine, phenylalanine, proline, serine, tyrosine, valine)
Breaking buffer
- 50 mM sodium phosphate buffer, pH 7,5
- 1 mM EDTA
- 5% (v/v) glycerol
- 1 mM PMSF
Tuesday, August 15th
Miniprep of overnight culture of transformated E. coli XL1-Blue with with ligated P71+P322, P71+P323, P206+P337 and P342+PCR66
Investigator: Jeff
Aim of the experiment: Miniprep of overnight culture of transformated E. coli XL1-Blue with with ligated P71+P292, P71+P293, P206+P337 and P342+PCR66.
Procedure:
- Miniprep was done with the Qiagen Qiaprep spin miniprep kit and was performed after manufacturer's protocol.
Analytical digestion and gelelectrophoresis of the minipreps of transformated E. coli XL1-Blue with ligated P71+P322 and P71+P323 products
Investigator: Jeff
Aim of the experiment: Analytical digestion and gelelectrophoresis of the minipreps of transformated E. coli XL1-Blue with ligated P71+P322 and P71+P323 products (P352-P356).
Procedure:
- Mastermix for analytical digestion with XbaI+PstI-HF
| volume | reagent |
| 22 µl | NEBuffer 4 (10x) |
| 2.2 µl | BSA (100x) |
| 2.75 µl | XbaI (20 U/µl) |
| 2.75 µl | PstI-HF (20 U/µl) |
| 162.8 µl | ddH2O |
| =192.5 µl | TOTAL |
- 17.5 µl of the mastermix for XbaI+PstI-HF was added to 2.5 µl of plasmid DNA of P352-P356.
- Incubation for 2 h at 37 °C.
- Analytical gelelectrophoresis was performed at 90 V for 60 min.
repetition of analytical restriction digest of ligation products of 10.08. after miniprep
Investigator: Andrea
Aim: Check ligation of PCR 42 and PCR43 (Citrus) & PCR 56, PCR 57 and PCR 58 (Lavendula) into pYES and pSB1C3.
| volume | reagent |
| 20 µl | miniprep products |
| 2 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 0.25 µl | XbaI (20 U/µl) |
| 0.25 µl | SpeI (20 U/µl) |
| =20 µl | TOTAL |
- Master-Mix: 7 µl Enzyme, 56 µl NEB-Buffer, 5,6 µl BSA, 414,4 µl ddH2O
- 2,5 µl DNA added to 17,5 µl Master-Mix
- Incubation at 37°C for 2 h.
Gelelectrophoresis:
Preparation of cells for protein expression in yeast
Investigator: Katrin
Aim of the experiment:Picking transformed Saccharomyces cerevisae cells for protein expression in yeast (expression will be done on Thursday, August 16th)
- 900 ml of SCU medium was mixed with 100 ml glucose
- one clone from each plates with PAL+/-, CHS+/-, 4CL+/-, OMT1+/-, APT, GFP was picked,resuspended in 15 ml SCU medium with glucose and incubated at 30 °C overnight
 "
"