Team:Cambridge/Ratiometrica/Results
From 2012.igem.org
(→Ratiometrica Results) |
(→Ratiometrica Results) |
||
| Line 85: | Line 85: | ||
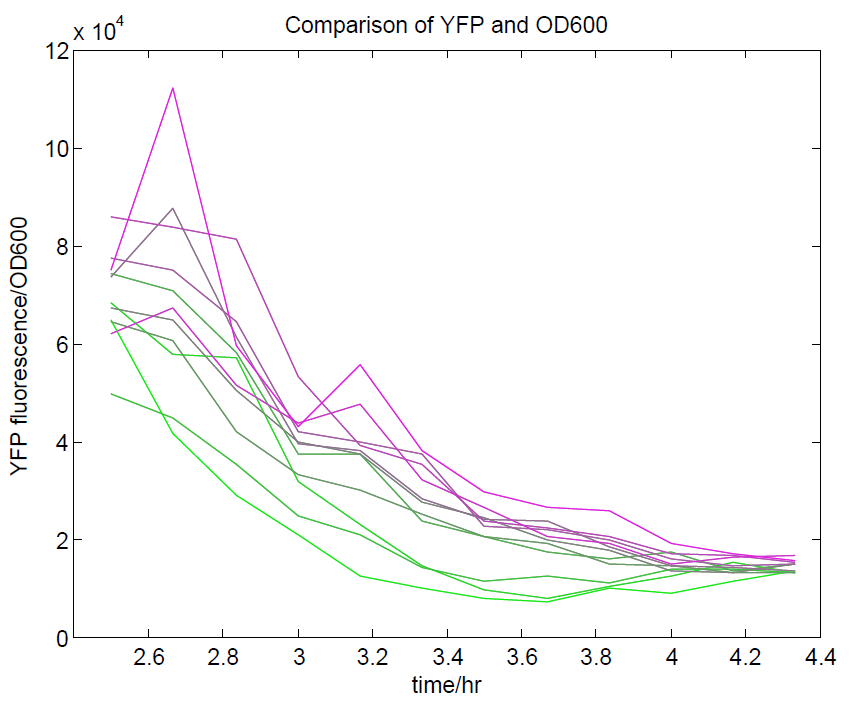
[[File:YFPdividedbyOD600.png|500px|center|thumb|YFP levels divided by OD600. Because YFP should be a measure of cell density along with OD600, we expected that this ratio should approach a constant value. This appears to be the case, although only at higher cell densities.]] | [[File:YFPdividedbyOD600.png|500px|center|thumb|YFP levels divided by OD600. Because YFP should be a measure of cell density along with OD600, we expected that this ratio should approach a constant value. This appears to be the case, although only at higher cell densities.]] | ||
| - | |||
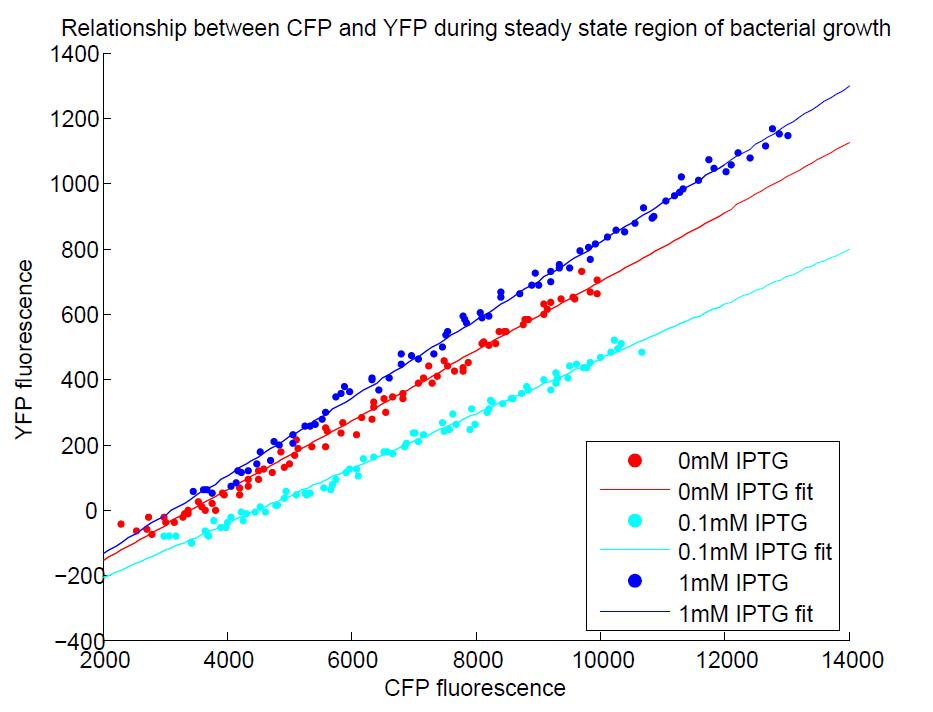
| - | [[File:GradientsComparisonFull.png| | + | Additionally, in another series of experiments where we induced with IPTG, we found that the CFP:YFP ratio for individual wells was steady upon reaching steady state, indicating that it acts as a good internal control. The construct as a whole did not respond to IPTG as we expected, but we think this is a problem with our assay, as the internal reference seems to be working well! |
| + | |||
| + | [[File:GradientsComparisonFull.png|500px|center|thumb|The CFP:YFP ratio is remarkably constant during steady state]] | ||
A difference in CFP has also been observed between uninduced and induced (1mM IPTG) plates under the fluorescence microscope. Unfortunately, we were unable to test this induction (and the consequences of ratiometric measurement on the reliability of resulting data) with the plate reader before the deadline (possibly due to difficulties with culturing the cells in M9 minimal medium). We will reattempt this assay before the jamboree and are confident that results should be as expected. | A difference in CFP has also been observed between uninduced and induced (1mM IPTG) plates under the fluorescence microscope. Unfortunately, we were unable to test this induction (and the consequences of ratiometric measurement on the reliability of resulting data) with the plate reader before the deadline (possibly due to difficulties with culturing the cells in M9 minimal medium). We will reattempt this assay before the jamboree and are confident that results should be as expected. | ||
| Line 93: | Line 94: | ||
Transformation of B. subtilis with the plasmid has also been attempted- unfortunately we are still finetuning our B. subtilis transformation protocol. Again, we hope to show that this works before the Jamboree. | Transformation of B. subtilis with the plasmid has also been attempted- unfortunately we are still finetuning our B. subtilis transformation protocol. Again, we hope to show that this works before the Jamboree. | ||
| - | + | = Luciferase Construct = | |
Our ratiometric luciferase construct arrived with full sequence coverage. Unfortunately, during construction it was found to be unexpectedly toxic. This hampered characterisation, as it the construct tended to be lost, and necessitated its submission to the registry in a low copy number backbone (with permission from HQ). | Our ratiometric luciferase construct arrived with full sequence coverage. Unfortunately, during construction it was found to be unexpectedly toxic. This hampered characterisation, as it the construct tended to be lost, and necessitated its submission to the registry in a low copy number backbone (with permission from HQ). | ||
| Line 117: | Line 118: | ||
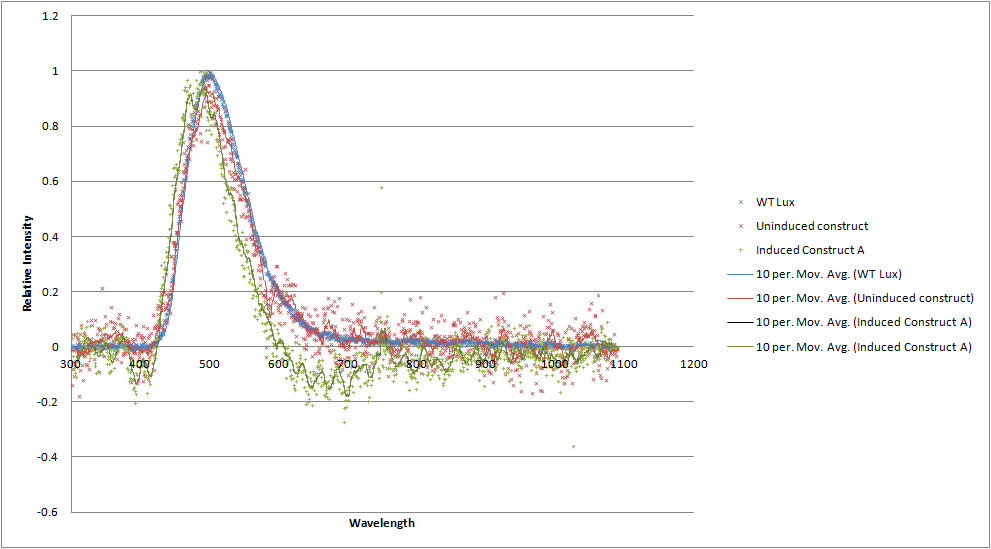
| - | UPDATE: We have obtained emission spectra for the construct when uninduced and induced. The | + | '''UPDATE:''' We have obtained emission spectra for the construct when uninduced and induced. The spectra are a little noisy due to the difficulties in growing and expressing the toxic construct in culture, meaning that the emission is not very bright. Both in the uninduced and induced state, there '''does not''' seem to be a spectral shift. This is not what we had hoped for, but we have a number of hypotheses as to why. The rest of the lux genes are arranged in a cotranscribed operon, so it's possible the wildtype luxA subunit has a huge advantage over the modified one in terms of binding luxB as it is made. Alternatively, the spectral shift produced by the mOrange - luxA fusion is simply not as the original paper described. |
| - | [[File:Luciferases_graph.png| | + | [[File:Luciferases_graph.png|700px|center|thumb| A graph comparing the emission spectrum of our construct (red and green) to wildtype luciferase (blue). No shoulder at 560nm is apparent.]] |
Revision as of 00:11, 27 October 2012
Contents |
Judging Form
- Please help the judges by filling out this form. Tell them what medal you think you deserve and why. Tell them which special prizes you should win. Help them find your best parts. Show them how you thought about the safety of your project. Helping the judges will help you too.
- Team: Cambridge
- Region: Europe
- iGEM Year:2012
- Track:Foundational Advance
- Project Name:Parts for a reliable and field ready biosensing platform
- Project Abstract: Implementation of biosensors in real world situations has been made difficult by the unpredictable and non-quantified outputs of existing solutions, as well as a lack of appropriate storage, distribution and utilization systems. This leaves a large gap between a simple, functional sensing mechanism and a fully realised product that can be used in the field.
We aim to bridge this gap at all points by developing a standardised ratiometric luciferase output in a Bacillus chassis. This output can be linked up with prototyped instrumentation and software for obtaining reliable quantified results. Additionally, we have reduced the specialized requirements for the storage and distribution of our bacteria by using Bacillus' sporulation system. To improve the performance of our biosensing platform we have genetically modified Bacillus’ germination speed. Lastly, we demonstrated the robustness of our system by testing it with a new fluoride riboswitch, providing the opportunity to tackle real life problems.
iGEM Medals for non-software teams
- We believe our team deserves the following medal:
- Bronze
- Silver
- √Gold
Because we met the following criteria (check all that apply and provide details where needed)
Requirements for a Bronze Medal
- √Register the team, have a great summer, and plan to have fun at the Regional Jamboree.
- √Successfully complete and submit this iGEM 2012 Judging form.
- √Create and share a Description of the team's project using the iGEM wiki and the team's parts using the Registry of Standard Biological Parts.
- √Plan to present a Poster and Talk at the iGEM Jamboree.
- √Enter information detailing at least one new standard BioBrick Part or Device in the Registry of Standard Biological Parts. Including:
- √Primary nucleaic acid sequence
- √Description of function
- √Authorship
- Safety notes, if relevant.
- √Acknowedgment of sources and references
- √Submit DNA for at least one new BioBrick Part or Device to the Registry.
Additional Requirements for a Silver Medal
- √Demonstrate that at least one new BioBrick Part or Device of your own design and construction works as expected; characterize the operation of your new part/device.
- √Enter this information and other documentation on the part's 'Main Page' section of the Registry
Part Number(s): [http://partsregistry.org/Part:BBa_K911004 BBa_K911004]
Additional Requirements for a Gold Medal: (one OR more)
- Improve an existing BioBrick Part or Device and enter this information back on the Experience Page of the Registry.
Part Number(s): None - √Help another iGEM team by, for example, characterizing a part, debugging a construct, or modeling or simulating their system.
Link to this information on your wiki. Page name: Team:Cambridge/Outreach/Collaboration - √Outline and detail a new approach to an issue of Human Practice in synthetic biology as it relates to your project, such as safety, security, ethics, or ownership, sharing, and innovation.
Link to this information on your wiki.
Page name: Team:Cambridge/HumanPractices/Overview,Team:Cambridge/HumanPractices/MarketResearch,Team:Cambridge/HumanPractices/FutureDirections
iGEM Prizes
All teams are eligible for special prizes at the Jamborees. more... To help the judges, please indicate if you feel you should be evaluated for any of the following special prizes:
- √Best Human Practice Advance
- √Best Experimental Measurement
- Best Model
Please explain briefly why you should receive any of these special prizes:
Best Human Practice Advance:
We feel that we deserve this prize for three reasons:
- We explored the impacts, *both positive and negative*, of synthetic biology as a solution to real world problems, through interviewing professionals working in a relevant field, namely the impact of arsenic water contamination in Bangladesh.
- We recognized existing problems with the way the current direction of synthetic. On going through the registry we found that most of the characterization data for biosensing parts is often neither comparable nor replicable. We have worked to solve this issue, for example with our ratiometric dual channel output.
- *Our project doesn’t stop here*, in Chanel number 6 (Team:Cambridge/HumanPractices/FutureDirections) we considered the future implications and technological applications of our project, as well as the means by which it could be improved by subsequent users. We feel that the end to an iGEM project should not be the conclusion of an idea, but the start of it.
Best BioBrick Measurement Approach:
It is absolutely vital that a quantitative, numerical, robust, and flexible measurement approach exists to relay information to a user that is an accurate representation of the input processed by a biological device. Working from these principles, the following was done:
- We designed and built Biologger, a *cheap, arduino-based, fully functional automatic rotary device* that has an incorporated ratiolumnometer
- Our project is entirely open-sourced and open-platform. We have published source code for the two applications which serve to operate the device, one for PCs and the other for Android devices, as well as the open source circuit design that provides this ratiometric reading. Furthermore, the Android app is able to receive its data wirelessly, which we feel is a great advance in BioBrick measurement.
- Our dual-channel luciferase reporter was successfully tested with a dilution series of E.coli transformed with the Lux Operon (under pBAD) biobrick (Part BBa_K325909) of the Cambridge iGEM 2010 team. It can detect, with good accuracy, both different light intensities, as well as the percentages of blue or orange frequencies in a sample.
- Our device was successfully tested using artificial light to detect different frequencies (colours) as well.
Having done all the above, we believe that this fully open-sourced instrumentation kit (mechanical) chassis, electronics, software code), estimated at *$35.00* (or $85.00 if a Bluetooth modem is required), is a complete BioBrick measurement solution for any and all BioBricks with a light output.
Team_Parts
To help the judges evaluate your parts, please identify 3 of your parts that you feel are best documented and are of the highest quality.
- Best new BioBrick part (natural)
- [http://partsregistry.org/Part:BBa_K911003 BBa_K911003]
- Best new BioBrick part (engineered)
- [http://partsregistry.org/Part:BBa_K911004 BBa_K911004]
- Best improved part(s): None
List any other parts you would like the judges to examine:[http://partsregistry.org/Part:BBa_K911001 BBa_K911001], [http://partsregistry.org/Part:BBa_K911008 BBa_K911009], [http://partsregistry.org/Part:BBa_K911008 BBa_K911008]
Please explain briefly why the judges should examine these other parts:
- Magnesium Sensitive Riboswitch [http://partsregistry.org/Part:BBa_K911001 BBa_K911001]
As a riboswitch sensing construct, this part is an entirely new type of biosensor (along with the fluoride construct) that could potentially change the way we think about designing input genetic circuits. Unlike the fluoride riboswitch, it is a derepression system and therefore serves to demonstrate the principle that riboswitches can be used regardless of whether they turn on or off their reporter. - Fluorescent ratiometric construct for standardizing promoter output [http://partsregistry.org/Part:BBa_K911009 BBa_K911009]
Fluorescence is a major cornerstone for biosensors in the registry, however, most parts do not involve the use of a ratiometric output, which has been shown in the literature to provide much more reliable and meaningful data. This part not only furthers the development of ratiometric measurements in molecular biology but due to the choice of promoters and terminators it can be used to characterize the difference in activity between E. coli and B. Subtilis - Fast Germination (B. subtilis) [http://partsregistry.org/Part:BBa_K911008 BBa_K911008]
This part is entirely novel for the registry and fully utilizes the recombination machinery inherent in the Bacillus chassis. Have spores that can germinate at a faster rate is certainly a worthy achievement and could help with experiments with B. Subtilis that any future iGEM teams may wish to perform.
iGEM Safety
For iGEM 2012 teams are asked to detail how they approached any issues of biological safety associated with their projects.
The iGEM judges expect that you have answered the four safety questions Safety page on your iGEM 2012 wiki.
Please provide the link to that page: Page name: Team:Cambridge/Safety
Attribution and Contributions
For iGEM 2012 the description of each project must clearly attribute work done by the team and distinguish it from work done by others, including the host labs, advisors, and instructors.
Please provide the link to that page, or comments in the box below: Page name: Team:Cambridge/Attributions
Comments
If there is any other information about your project you would like to highlight for the judges, please provide a link to your wiki page here: Team:Cambridge/Overview/DesignProcess
Ratiometrica Results
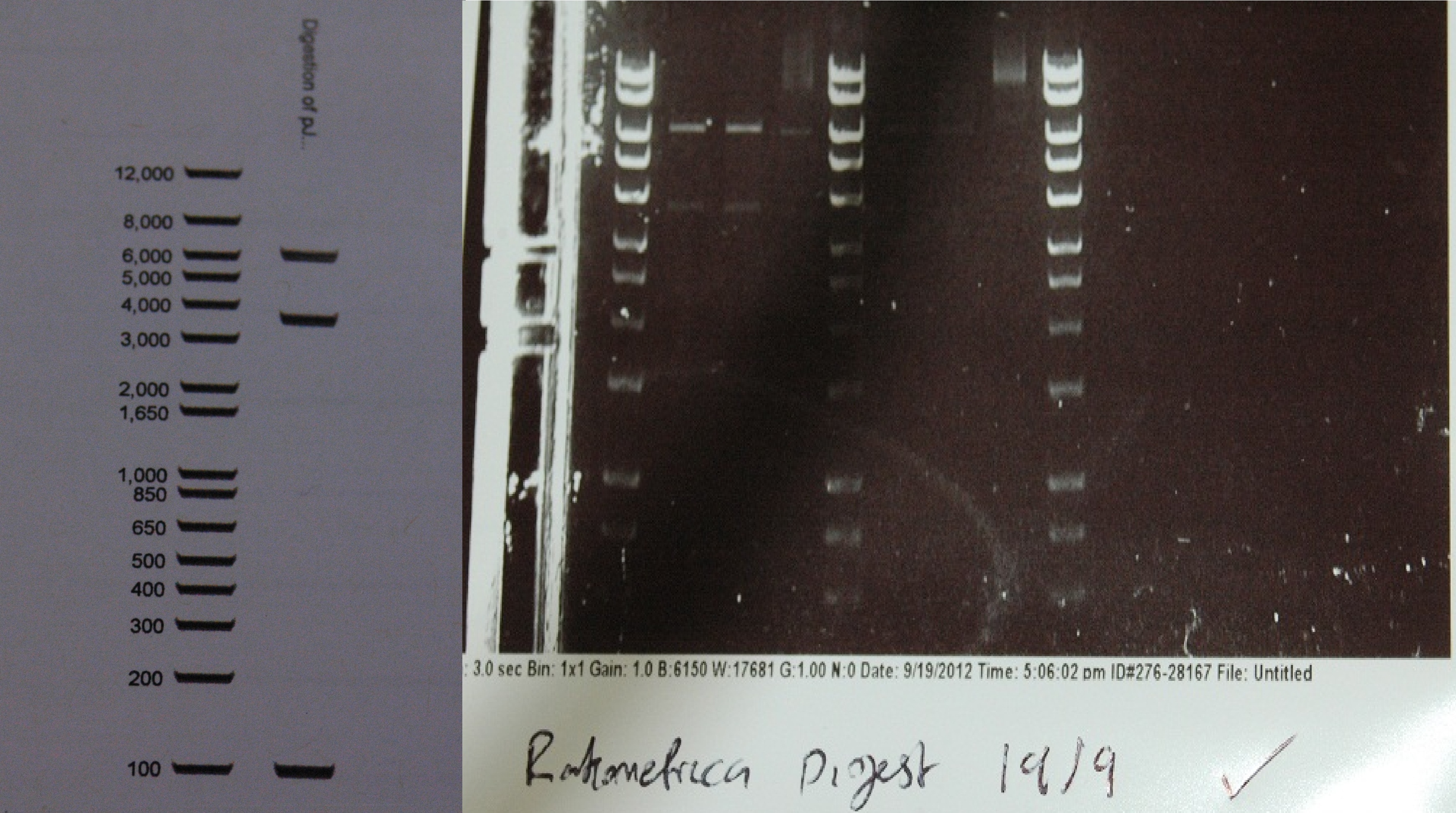
After a lot of technical difficulties, we were able to assemble our fluorescent construct using Gibson assembly. The photo below shows the predicted and obtained digest pattern using HINDIII.
The biobrick DNA has been sequenced and sent to the registry.
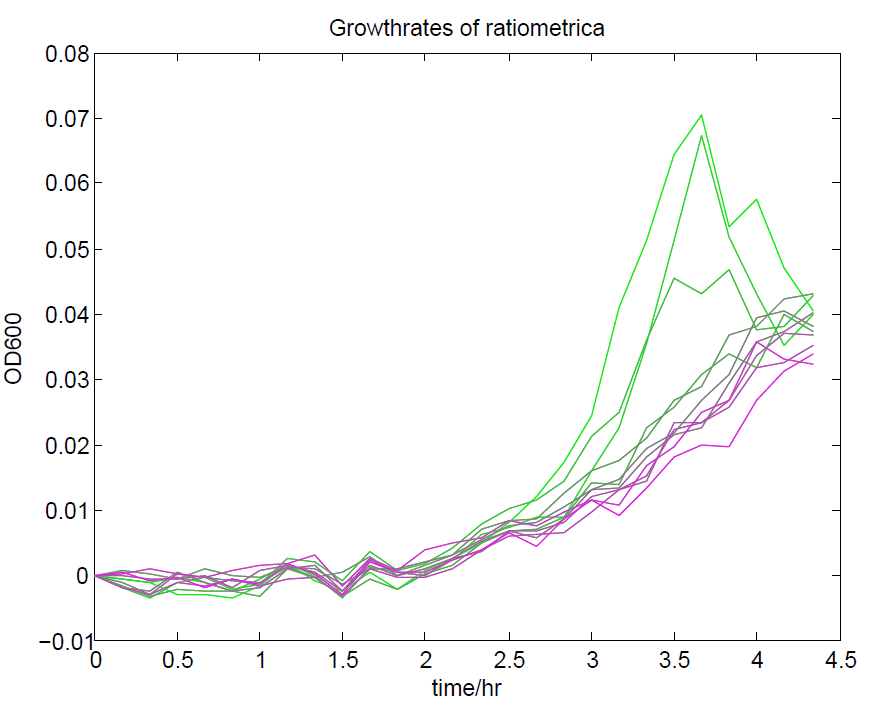
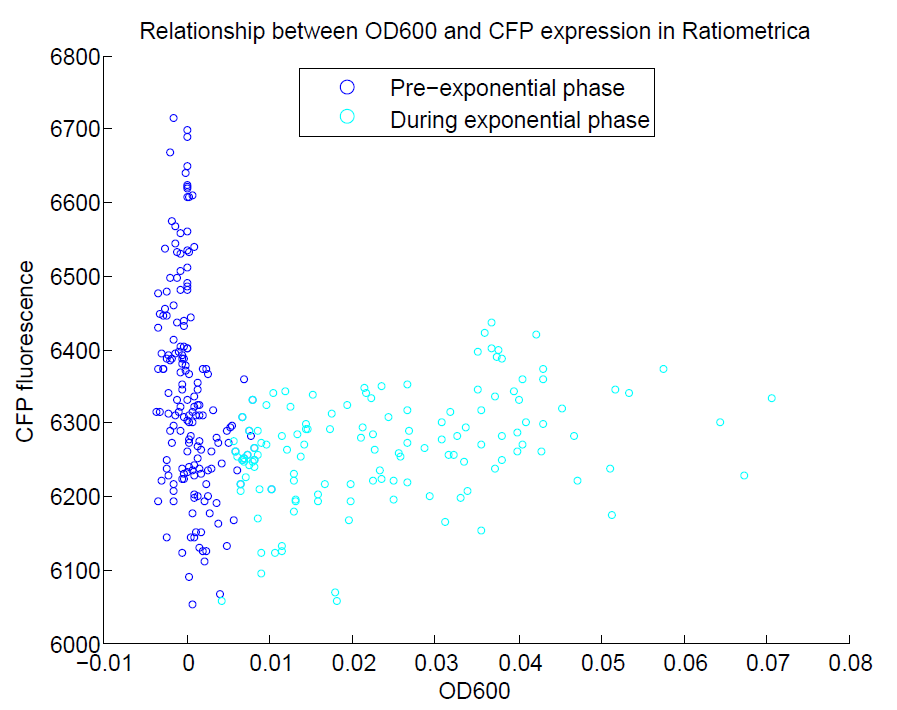
We ran a plate reader assay with ratiometrica with no induction of the reporter CFP. The results of this assay can be seen. In summary:
- No change in CFP is observed with changing OD600.
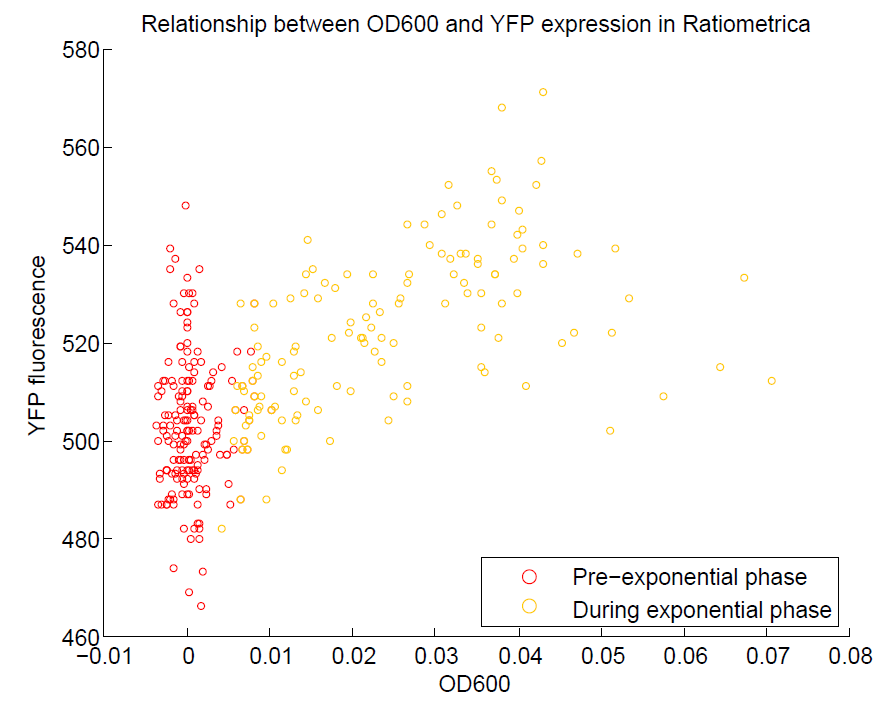
- YFP change is approximately proportional to OD600.
- The ratio of YFP to OD600 approaches a constant value at higher cell densities.
These properties are exactly what we expected them to be. This assay also tells us that we should aim to take our ratiometric measurements only during late exponential phase, as this is where the data is most reliable.
Additionally, in another series of experiments where we induced with IPTG, we found that the CFP:YFP ratio for individual wells was steady upon reaching steady state, indicating that it acts as a good internal control. The construct as a whole did not respond to IPTG as we expected, but we think this is a problem with our assay, as the internal reference seems to be working well!
A difference in CFP has also been observed between uninduced and induced (1mM IPTG) plates under the fluorescence microscope. Unfortunately, we were unable to test this induction (and the consequences of ratiometric measurement on the reliability of resulting data) with the plate reader before the deadline (possibly due to difficulties with culturing the cells in M9 minimal medium). We will reattempt this assay before the jamboree and are confident that results should be as expected.
Transformation of B. subtilis with the plasmid has also been attempted- unfortunately we are still finetuning our B. subtilis transformation protocol. Again, we hope to show that this works before the Jamboree.
Luciferase Construct
Our ratiometric luciferase construct arrived with full sequence coverage. Unfortunately, during construction it was found to be unexpectedly toxic. This hampered characterisation, as it the construct tended to be lost, and necessitated its submission to the registry in a low copy number backbone (with permission from HQ).
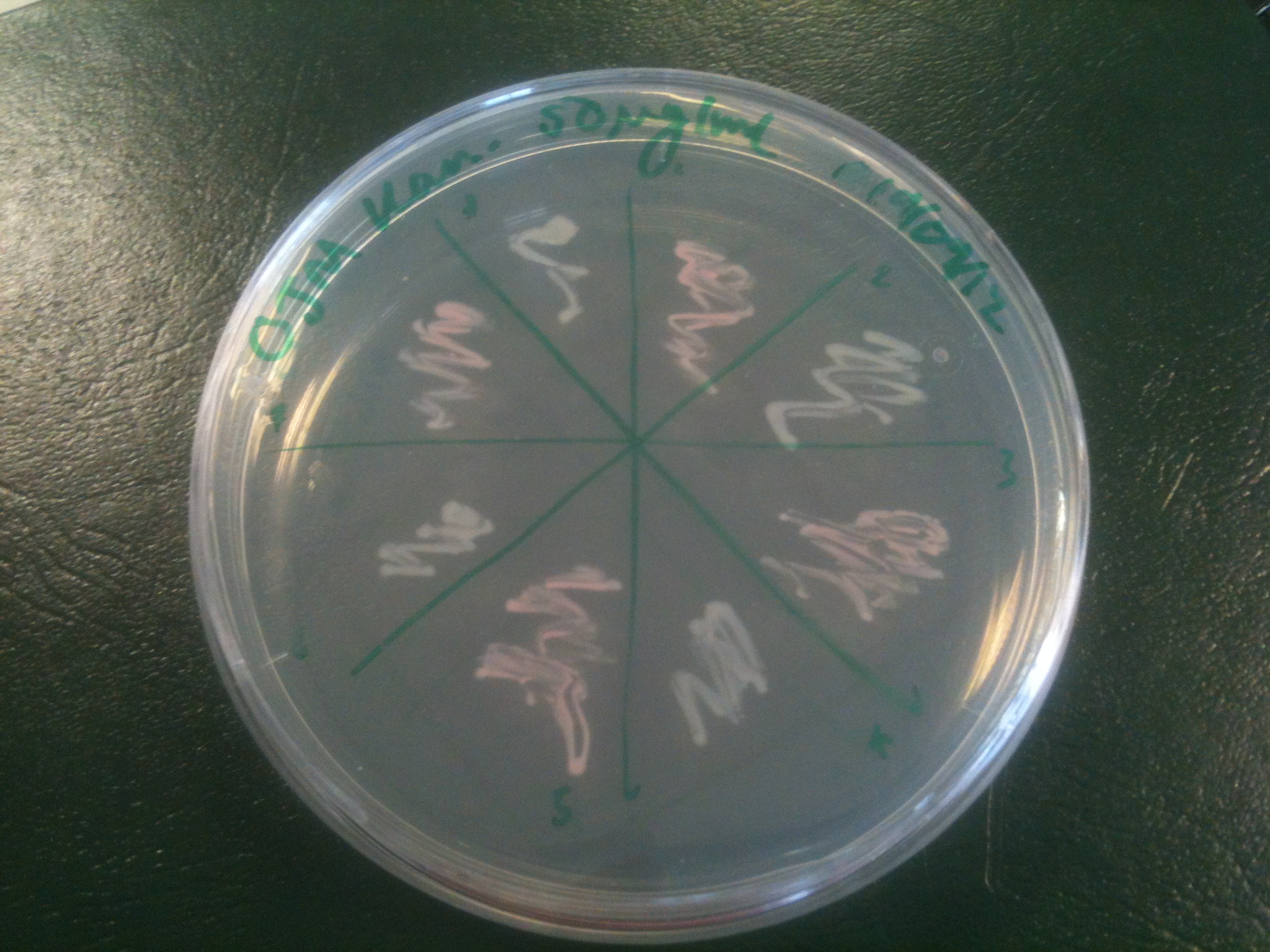
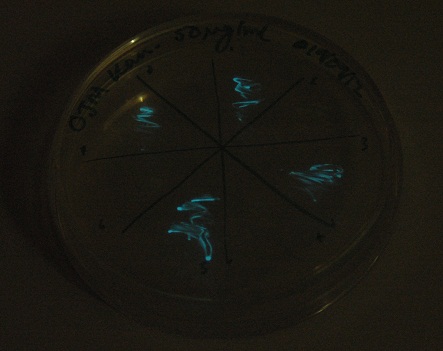
Starting from the ground up, we first showed that the part was constitutively luminescing. We also observed that the colonies were orange. This was not initially expected, however this is probably because we designed the construct with consensus RBSes, so any leaky transcription through the inducible promoter would result in enough protein to be visible. We showed that orange colour and luminescence cosegregated, implying that the entire construct is being lost, not just part of it.
Given that IPTG induction seemed unnecessary for production of mOrange, we reasoned that any spectral shift in the emission of the luciferase should be visible without induction. Our next move was to see if we could detect a difference with the filters we used for the instrumentation.
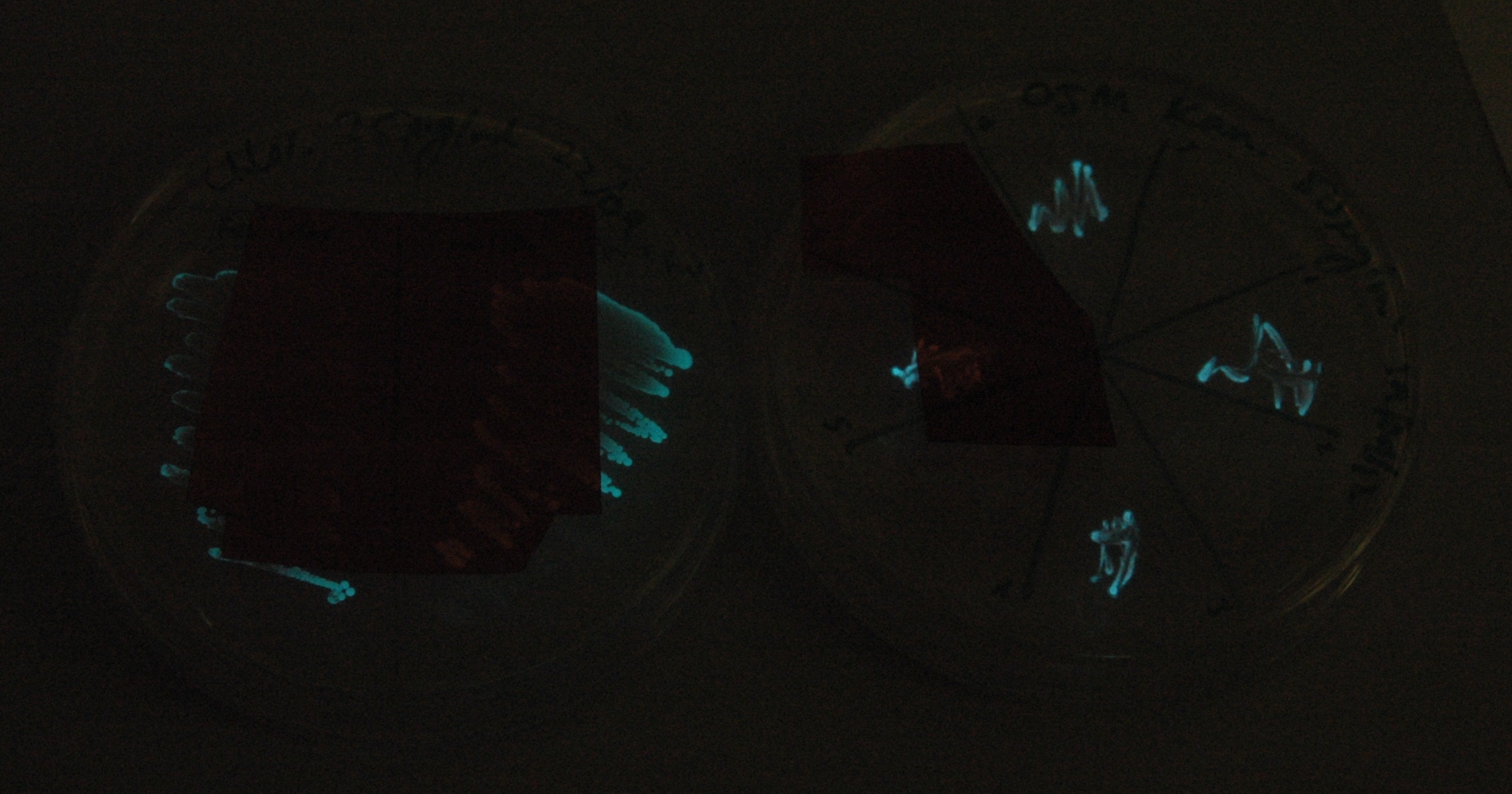
This was done at quite a late stage and in a slightly impromptu fashion. The photographs above directly compare the normal lux biobrick (BBa_K325909) on the left, and our construct on the right, with the same filter. There does seem to be a difference in the quantity of light coming through the filter. However it could be due to colony density (although they seem to be of similar intensities without the filter). Interesting though these results are, they are not quantitative or well-controlled enough to constitute confirmation that the luxA-mOr fusion is behaving as expected, rather they are an indication that it may be.
To categorically confirm that the spectrum is as we expected we would want to fully characterise the emission spectrum with a scanning luminometer. We do not currently have access to a scanning luminometer in time to submit the data here before the wiki freeze, however we hope to obtain this data in the next week, and to be able to present it at the jamboree.
UPDATE: We have obtained emission spectra for the construct when uninduced and induced. The spectra are a little noisy due to the difficulties in growing and expressing the toxic construct in culture, meaning that the emission is not very bright. Both in the uninduced and induced state, there does not seem to be a spectral shift. This is not what we had hoped for, but we have a number of hypotheses as to why. The rest of the lux genes are arranged in a cotranscribed operon, so it's possible the wildtype luxA subunit has a huge advantage over the modified one in terms of binding luxB as it is made. Alternatively, the spectral shift produced by the mOrange - luxA fusion is simply not as the original paper described.
 "
"