Team:LMU-Munich/Application/Further Applications tour
From 2012.igem.org
| Line 8: | Line 8: | ||
<p align="justify">Ultimately, the ability to express virtually any protein of interest shows great potential for laboratory work. Our '''Sporo'''beads could additionally express TAL effectors, for the binding of sequence-specific DNA stretches. This could allow simple GMO detection of food crops. Besides simply expressing proteins capable of binding elements of interest, our '''Sporo'''beads could express proteins which have enzymatic activity. The [https://2011.igem.org/Team:Washington 2011 University of Washington iGEM team] developed an enzyme, Kumamolisin, which cleaves peptides. The specific substrate of this enzyme is the specific amino acid sequence which causes reactions in individuals with Celiac disease. Such a cleaving enzyme could be used to eliminate irritants from gluten-containing food products.</p> | <p align="justify">Ultimately, the ability to express virtually any protein of interest shows great potential for laboratory work. Our '''Sporo'''beads could additionally express TAL effectors, for the binding of sequence-specific DNA stretches. This could allow simple GMO detection of food crops. Besides simply expressing proteins capable of binding elements of interest, our '''Sporo'''beads could express proteins which have enzymatic activity. The [https://2011.igem.org/Team:Washington 2011 University of Washington iGEM team] developed an enzyme, Kumamolisin, which cleaves peptides. The specific substrate of this enzyme is the specific amino acid sequence which causes reactions in individuals with Celiac disease. Such a cleaving enzyme could be used to eliminate irritants from gluten-containing food products.</p> | ||
[[File:LMU Sporo diversity.png|left|200px]] | [[File:LMU Sporo diversity.png|left|200px]] | ||
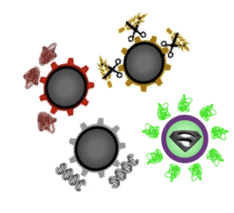
| - | <p align="justify">There are many possible applications for our | + | <p align="justify">There are many possible applications for our '''Sporo'''beads, as it is possible to display all kinds of different proteins on their surface. To easily create them, we designed a '''Sporo'''vector in which you just have to insert the gene encoding your fusion protein of choice.</p> |
Revision as of 16:52, 26 October 2012
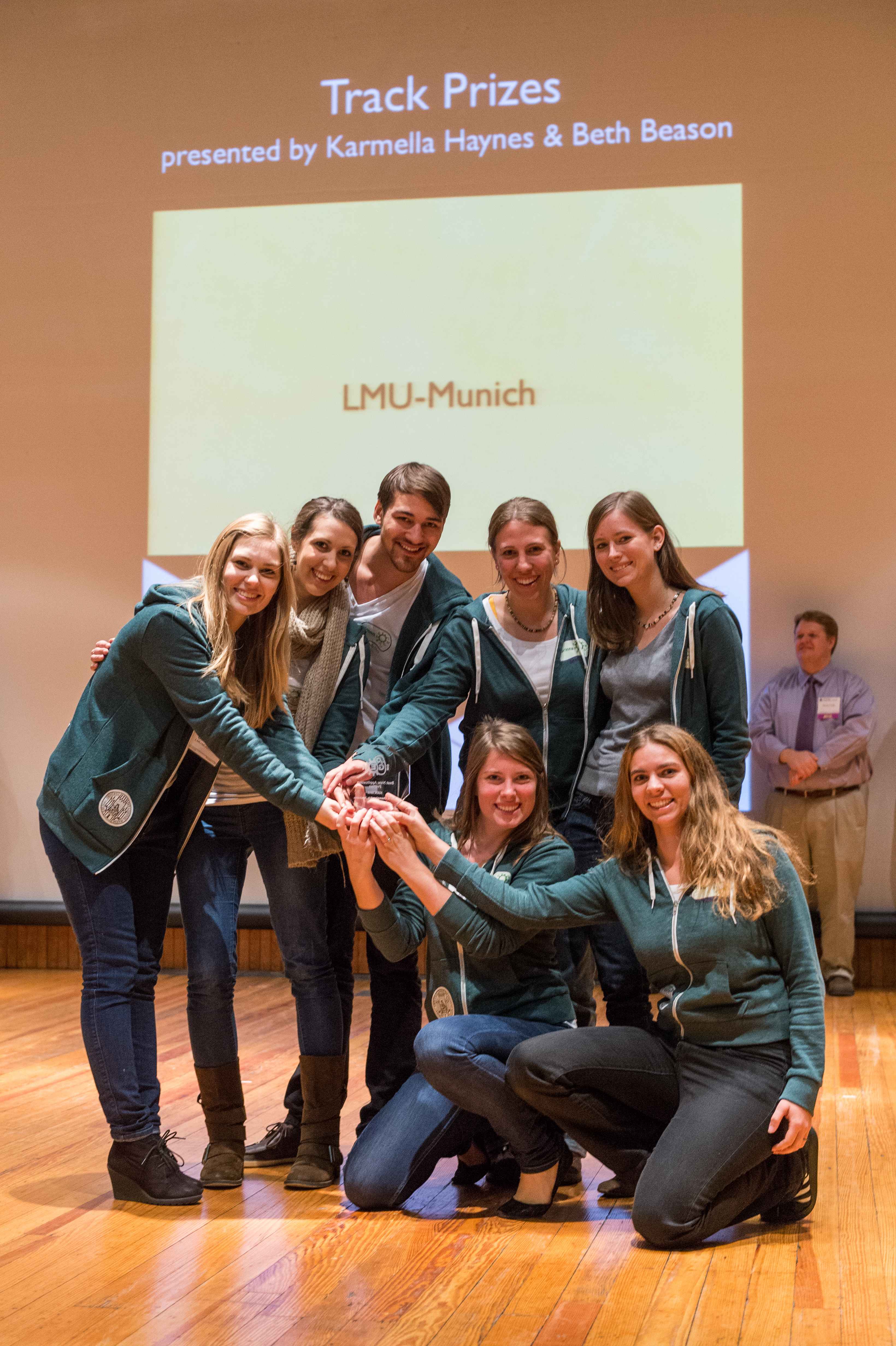
The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]


Further Applications
Ultimately, the ability to express virtually any protein of interest shows great potential for laboratory work. Our Sporobeads could additionally express TAL effectors, for the binding of sequence-specific DNA stretches. This could allow simple GMO detection of food crops. Besides simply expressing proteins capable of binding elements of interest, our Sporobeads could express proteins which have enzymatic activity. The 2011 University of Washington iGEM team developed an enzyme, Kumamolisin, which cleaves peptides. The specific substrate of this enzyme is the specific amino acid sequence which causes reactions in individuals with Celiac disease. Such a cleaving enzyme could be used to eliminate irritants from gluten-containing food products.
There are many possible applications for our Sporobeads, as it is possible to display all kinds of different proteins on their surface. To easily create them, we designed a Sporovector in which you just have to insert the gene encoding your fusion protein of choice.
 "
"