Team:Colombia/Project/Experiments/Ralstonia
From 2012.igem.org
m (→Ralstonia BioBricks) |
m (→PxpsR Reporter) |
||
| Line 9: | Line 9: | ||
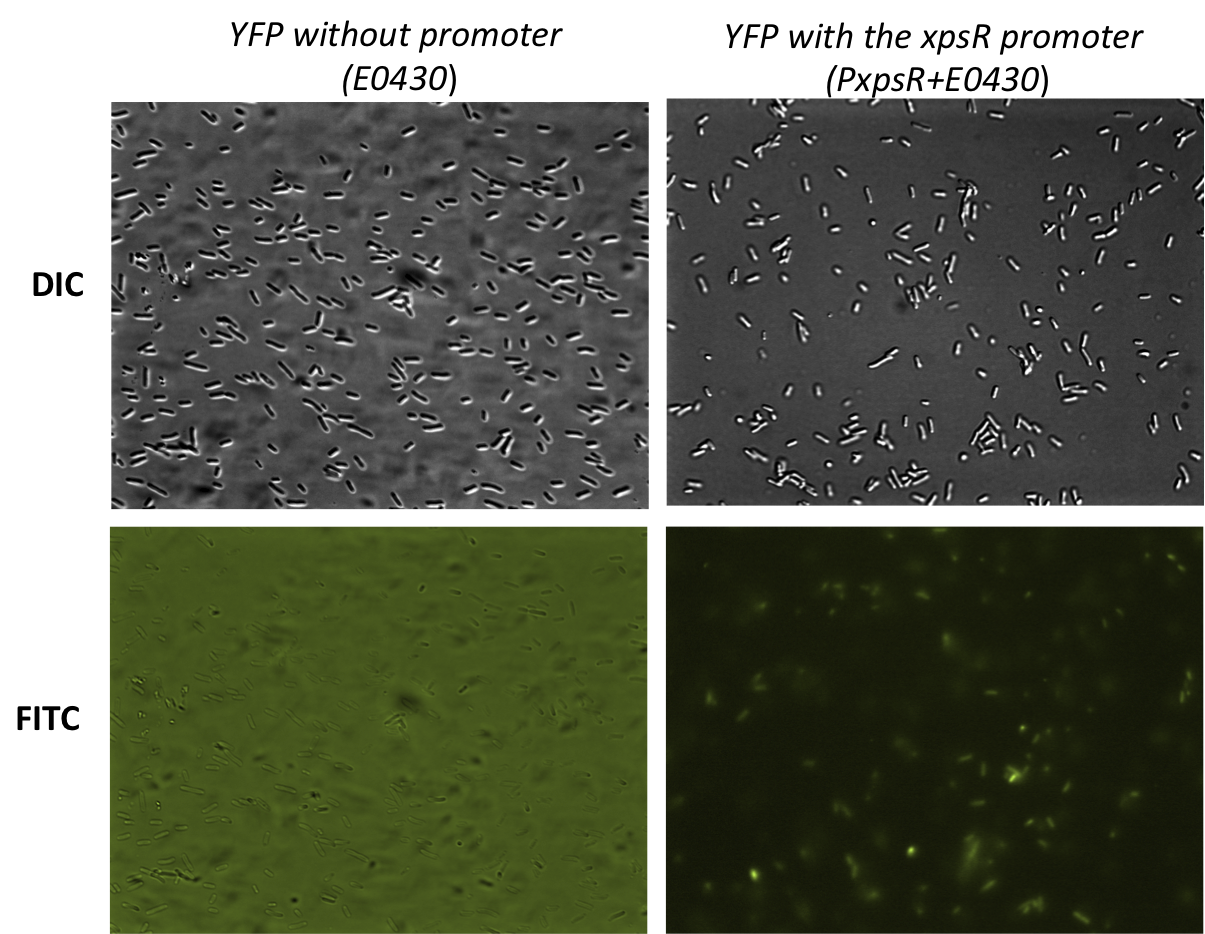
In order to quantify the responsiveness of PxpsR to the presence of 3 OH-PAME we generated a PxpsR-eYFP reporter. The reporter was tested in the pSB1A2 however the part was ensambled into de pSB1C3 backbone and sent to the iGEM Registry Parts. Here we show the functionality and basal activity (in absence of 3 OH-PAME) of the PxpsR-eYFP reporter (Image 2). | In order to quantify the responsiveness of PxpsR to the presence of 3 OH-PAME we generated a PxpsR-eYFP reporter. The reporter was tested in the pSB1A2 however the part was ensambled into de pSB1C3 backbone and sent to the iGEM Registry Parts. Here we show the functionality and basal activity (in absence of 3 OH-PAME) of the PxpsR-eYFP reporter (Image 2). | ||
| - | [[File:PxpsR-YFP.png|450px|thumb|center]] | + | [[File:PxpsR-YFP.png|450px|thumb|center|''Image 2''. '''Functionality of the reporter PxpsR-eYFP'''. Bacterials strains were photograph while using Differential Interference Contrast (DIC) microscopy and Fluorescence microscopy using FITC filter. Bacteria transformed with PxpsR-eYFP construct show basal levels of expresión of the reporter while none of the bacteria transformed with the promotorless eYFP shows fluorescens. High background levels in the promotorless eYFP is due to the autoflorescence of the LB medium and the high exposition.]] |
==Response of PxpsR to synthetic 3 OH-PAME== | ==Response of PxpsR to synthetic 3 OH-PAME== | ||
Revision as of 15:07, 26 September 2012
Template:Https://2012.igem.org/User:Tabima
Contents |
Ralstonia Experiments
Ralstonia BioBricks
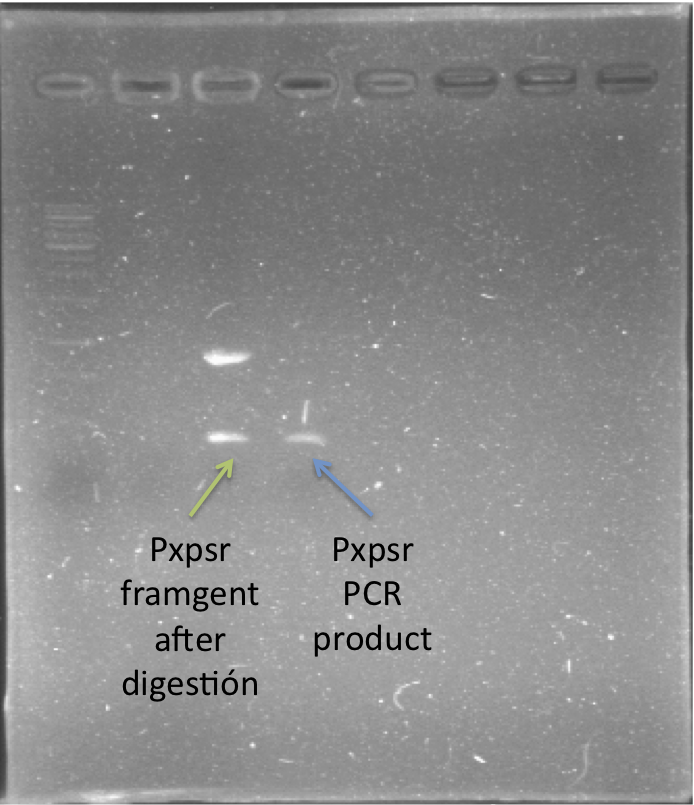
We cloned and generated a biobrick for the promoter of xpsR (PxpsR) which responds to the Quorum Sensing signal 3 OH-PAME especific of Ralstonia solanacearum, this part was also confirmed by sequenciation. In addition we cloned the three genes (phcA, phcS and phcR) involved in the sensing of the 3 OH-PAME, further confirmation of these parts are still needed. Image 1 shows enzymatic confirmation for PxpsR in the backbone pBS1C3. For a more extensive and more detailed cloning procedures here Ralstonia Journal
PxpsR Reporter
In order to quantify the responsiveness of PxpsR to the presence of 3 OH-PAME we generated a PxpsR-eYFP reporter. The reporter was tested in the pSB1A2 however the part was ensambled into de pSB1C3 backbone and sent to the iGEM Registry Parts. Here we show the functionality and basal activity (in absence of 3 OH-PAME) of the PxpsR-eYFP reporter (Image 2).
Response of PxpsR to synthetic 3 OH-PAME
Objective To determine the induced response of PxpsR in a mute Ralstonia solanacearum with the presence of synthetic 3 OH-PAME at different concentrations, using eYFP as a fluorescent reporter.
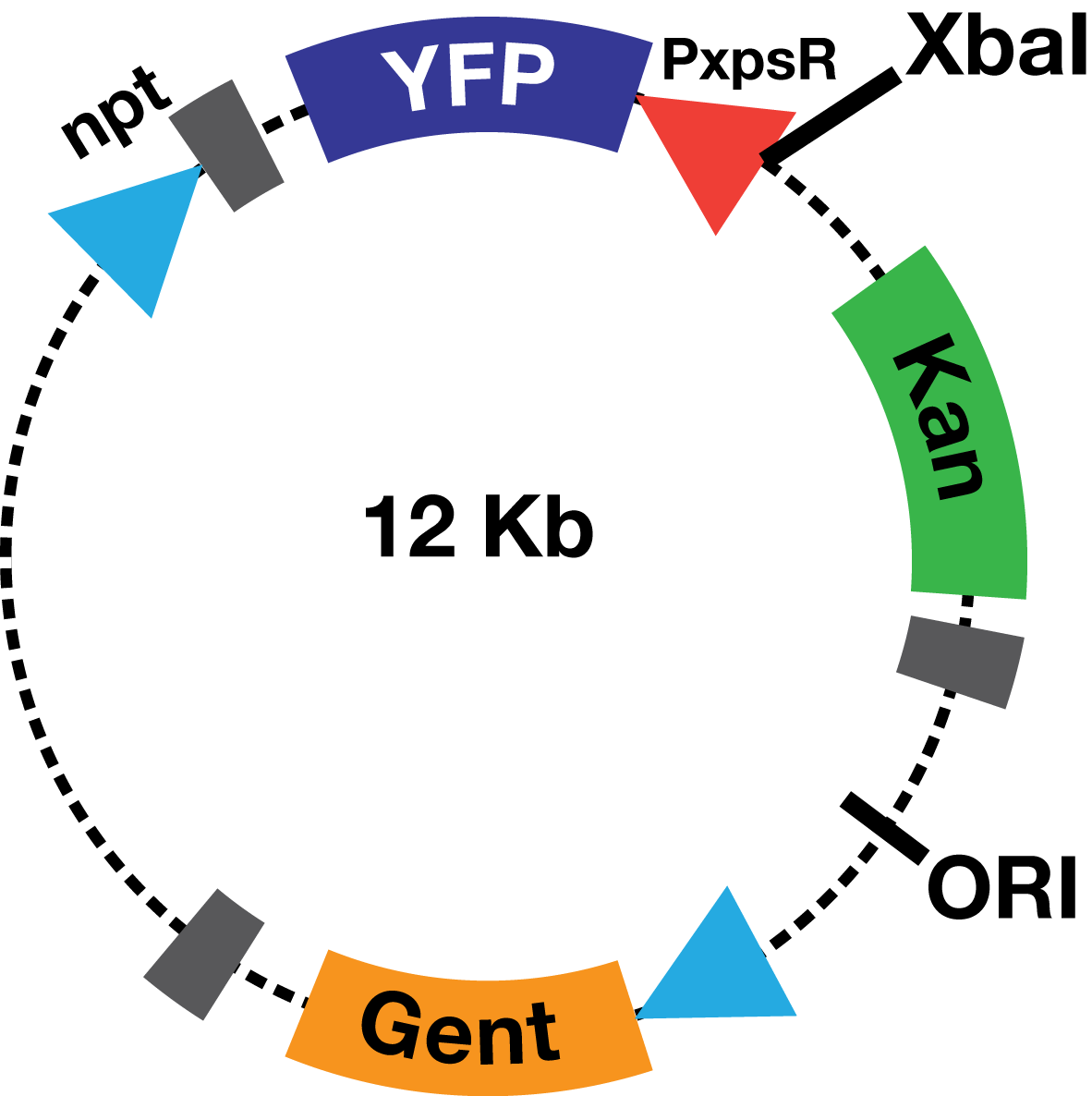
Description The PxpsR-eYFP biobrick was digested with XbaI and SpeI, and then was ligated to an XbaI digested ML123 plasmid. This one has a replication origin derived from pVS1, which allows replication in Xanthomonas, Pseudomonas and Ralstonia. To determine whether the biobrick had ligated in the opposite direction to the strong promoter npt (Figure 1), transformed colonies were confirmed via enzymatic digestions. This way the biobrick would not be induced by the npt promoter present in the plasmid, and therefore, the fluorescent response due to the presence of 3OH-PAME would depend solely on PxpsR.
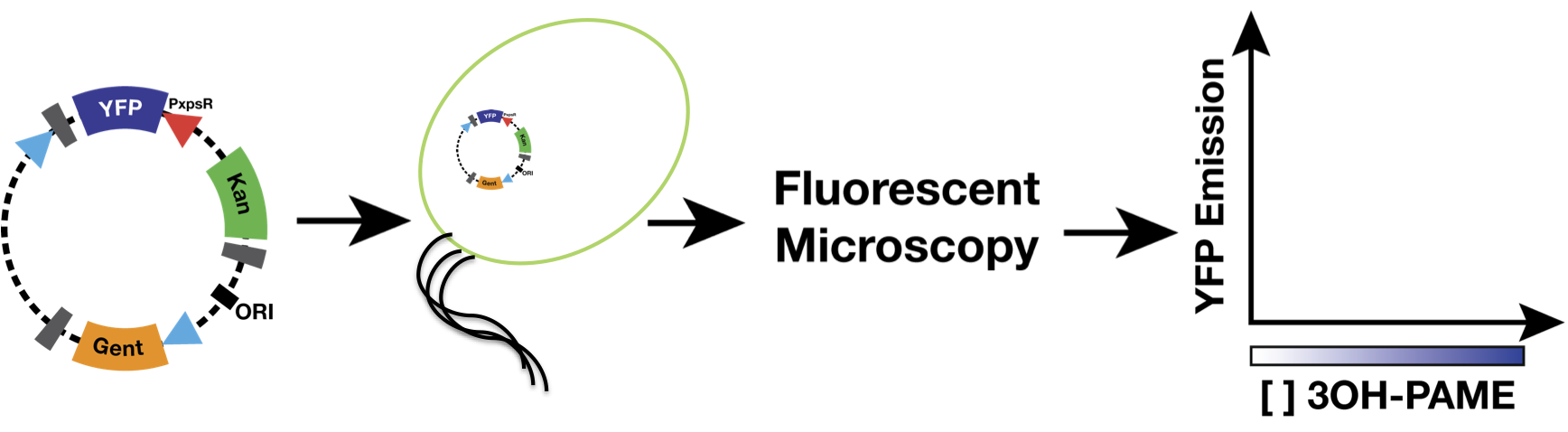
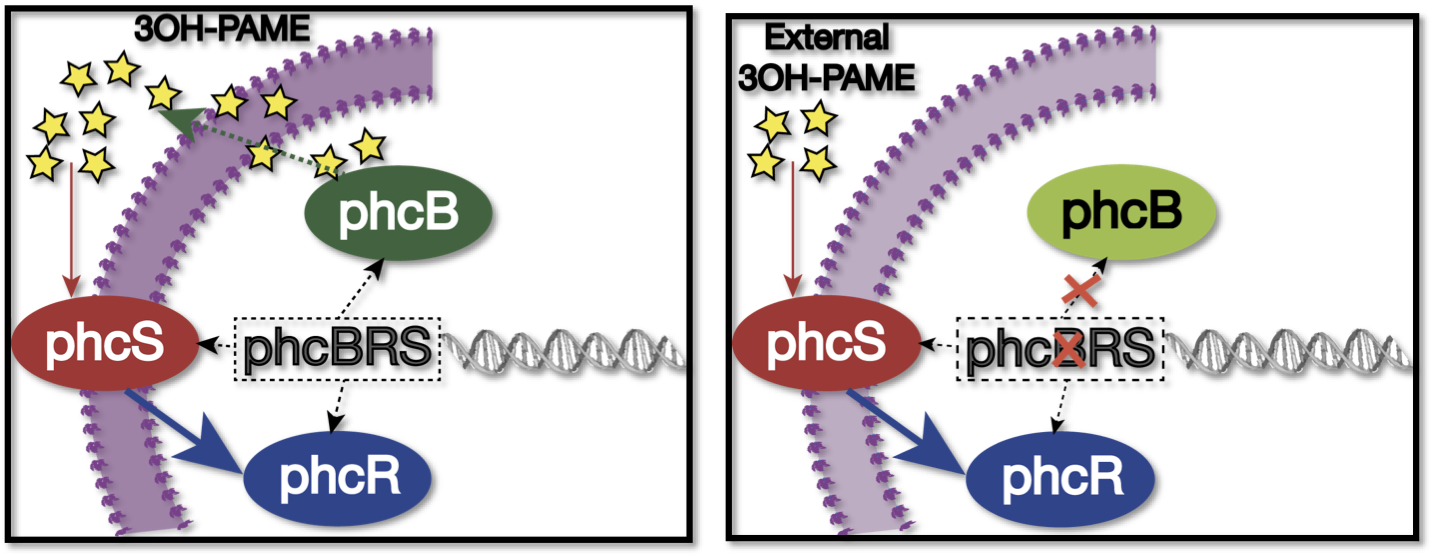
We used as host a mute mutant of Ralstonia Solanacearum, which has a mutation in the gene phcB (3 OH-PAME synthetase). Although it has the ability to sense 3 OH-PAME, it cannot produce it (Figure 2), which allows to create an environment with a known concentration of 3 OH-PAME at all times, indepent of the number of bacterias present. We are currently transforming the mute strain with the PxpsR-eYFP reporter in pML123, once we obtain the transformants we will determine the fluorescent response at different concentrations of synthetic 3-OH-PAME (Figure 3). Based on the fluorescent response, we will be able to determine the inducibility of PxpsR.
 "
"