Team:University College London/Module 2/Modelling
From 2012.igem.org
(→Reactions taking place in the model) |
(→Reactions taking place in the model) |
||
| Line 60: | Line 60: | ||
| R2 || POPex ↔ POPin || Forward: 0.6 <br /> Backward: 0.4 || Rate based on membrane permeability<sup>4</sup> and diffusion gradient | | R2 || POPex ↔ POPin || Forward: 0.6 <br /> Backward: 0.4 || Rate based on membrane permeability<sup>4</sup> and diffusion gradient | ||
|- | |- | ||
| - | | R3 || POPin + mRNA.Nahr | + | | R3 || POPin + mRNA.Nahr ↔ POPin.mRNA.Nahr || Forward: 1 <br /> Backward: 0.0001 || Based on the assumption that the chemical structure/size of POPs is similar to salycilate<sup>7</sup>. Salycilate binds to the NahR mRNA product, which complex then binds to the pSal promoter. |
|- | |- | ||
| - | | R4 || POPinmRNANahr | + | | R4 || POPinmRNANahr ↔ POPinmRNANahr.Psal || Forward: 78200 <br /> Backward: 0.191 <sup>10</sup> || NahR to pSal binding based on the assumption that POP-NahR binding has no effect on NahR-pSal binding |
|- | |- | ||
| R5 || POPexmRNANahr.Psal → mRNAcurli || 0.054 || Transcription rate of curli cluster in molecules/sec (for cluster size 1500bp<sup>11</sup>, transcription rate in E.coli 80bp/sec<sup>9</sup>) | | R5 || POPexmRNANahr.Psal → mRNAcurli || 0.054 || Transcription rate of curli cluster in molecules/sec (for cluster size 1500bp<sup>11</sup>, transcription rate in E.coli 80bp/sec<sup>9</sup>) | ||
| Line 72: | Line 72: | ||
| R8 || CsgB + CsgA -> fibril || ARGH || ARGH | | R8 || CsgB + CsgA -> fibril || ARGH || ARGH | ||
|- | |- | ||
| - | | R9 || 0 | + | | R9 || 0 ↔ mRNA.Nahr || Forward: 0.088 <br /> Backward: 0.6 || Transcription rate of NahR in molecules/sec (for NahR size 909 bp<sup>8</sup>, transcription rate in E.coli 80bp/sec<sup>9</sup>) under constitutive promoter control |
|- | |- | ||
| R10 || mRNA.Nahr → 0 || 0.03 || Degradation rate of NahR mRNA product<sup>14</sup> | | R10 || mRNA.Nahr → 0 || 0.03 || Degradation rate of NahR mRNA product<sup>14</sup> | ||
Revision as of 18:31, 25 September 2012
Contents |
Module 2: Aggregation
Description | Design | Construction | Characterisation | Shear Device | Modelling | Results | Conclusions
Modelling
Our cell model for aggregation describes the pathways through which the proteins CsgA and CsgB, which make up curli fibrils, are produced on detection of microplastics. Using this we aim to find out how many curlis each cell might express, which we could then use to estimate the binding strength of our bacteria.
As with our degradation model, this SimBiology model is divided into three compartments:
1. Outside1: in this compartment we see the association, adherence, and dissociation of persistent organic pollutants (POPs) from polyethylene (PE) (R1). We rely on the POPs to induce our degradation system, increasing by (105 to 106 the specificity to our system.
2. Cell: NahR is a constitutively produced mRNA product (R9) of which around 3% degrades (R10). When POP diffuses into the cell (R2), it forms a complex with intact NahR (R3) which then binds to the pSal promoter (R4) to induce the curli gene cluster (R5). This starts translation of five proteins which make up curli fibrils. Here we focus on those controlled by the CsgAB operon, CsgA (R7) and CsgB (R6), as these are the most important in curli synthesis as described on our [| research page]. The CsgA and CsgB that do not degrade (R11, R12) diffuse out of the cell.
3. Outside2: Outside of the cell the polymerization of CsgA by CsgB takes place (R8) to make a curli fibril.
Species
| Species | Initial value (molecules) | Notes & Assumptions |
|---|---|---|
| PE | 0.044 | Polyethylene found in North Pacific Gyre (value per cubic metre)1,2 |
| POPex | 0.0 | Persistent organic pollutants (ex = extracellular) that are not adhered to plastic surface |
| PEPOPex | 9.24E-5 | Persistent organic pollutants (ex = extracellular) that are adhered to the plastic surface3 |
| POPin | 0.5 | Persistent organic pollutants (in = intracellular) assumed from E. coli membrane permeability 4 |
| mRNANahR | 0.0 | NahR mRNA product |
| POPinNahR | 0.0 | Complex of the above two molecules |
| POPinNahRpSal | 0.0 | Complex of the above molecule and pSal (promoter that induces laccase transcription) |
| mRNACurli | 0.0 | Polycistronic mRNA as it codes for more than one protein, in reality curli cluster contains five or more proteins, in our model mRNA is present as CsgE5 |
| CsgA | 0.0 | One of the polypeptides that is coded for in curli cluster, it is a structural component secreted in subunits outside of the cell |
| CsgB | 0.0 | One of the polypeptides that is coded for in curli cluster, it is secreted outside of the cell allowing polymerization of CsgA |
| Fibril | 0.0 | Result of CsgA and CsgB interaction |
Reactions taking place in the model
| Number | Reaction | Reaction rate (molecules/sec) | Notes & Assumptions |
|---|---|---|---|
| R1 | PE + POPex ↔ PEPOPex | Forward: 1000 Backward: 1 | Pops have 1000 to 10000 times greater tendency to adhere to plastic than float free in the ocean6 |
| R2 | POPex ↔ POPin | Forward: 0.6 Backward: 0.4 | Rate based on membrane permeability4 and diffusion gradient |
| R3 | POPin + mRNA.Nahr ↔ POPin.mRNA.Nahr | Forward: 1 Backward: 0.0001 | Based on the assumption that the chemical structure/size of POPs is similar to salycilate7. Salycilate binds to the NahR mRNA product, which complex then binds to the pSal promoter. |
| R4 | POPinmRNANahr ↔ POPinmRNANahr.Psal | Forward: 78200 Backward: 0.191 10 | NahR to pSal binding based on the assumption that POP-NahR binding has no effect on NahR-pSal binding |
| R5 | POPexmRNANahr.Psal → mRNAcurli | 0.054 | Transcription rate of curli cluster in molecules/sec (for cluster size 1500bp11, transcription rate in E.coli 80bp/sec9) |
| R6 | mRNAcurli → CsgB | 0.13 | Translation rate of CsgB in molecules/sec (for CsgB size 151aa12, translation rate in E.coli 20aa/sec9) |
| R7 | mRNAcurli → CsgA | 0.13 | Translation rate of CsgA in molecules/sec (for CsgA size 151aa13, translation rate in E.coli 20aa/sec9) |
| R8 | CsgB + CsgA -> fibril | ARGH | ARGH |
| R9 | 0 ↔ mRNA.Nahr | Forward: 0.088 Backward: 0.6 | Transcription rate of NahR in molecules/sec (for NahR size 909 bp8, transcription rate in E.coli 80bp/sec9) under constitutive promoter control |
| R10 | mRNA.Nahr → 0 | 0.03 | Degradation rate of NahR mRNA product14 |
| R11 | CsgA → 0 | 0.03 | Degradation rate of CsgA 14 must be taken into account due to suboptimal conditions |
| R12 | CsgB → 0 | 0.03 | Degradation rate of CsgB 14 must be taken into account due to suboptimal conditions |
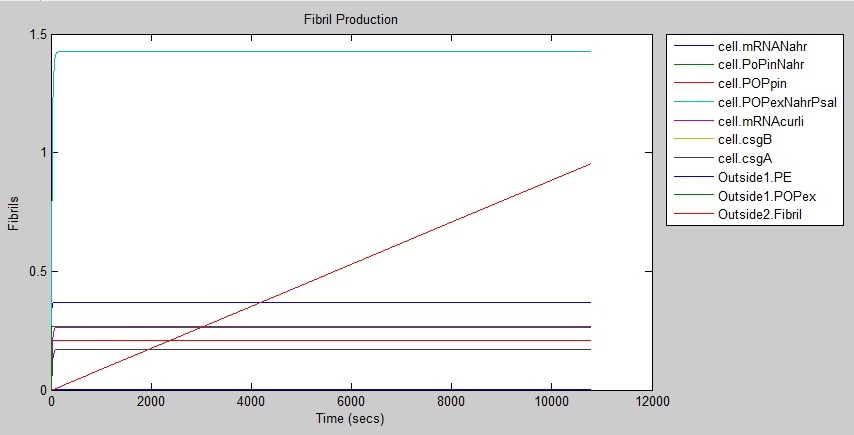
Results
The graph shows that fibrils are produced at a rate of around 1 fibril per second.
References
1. Goldstein M, Rosenberg M, Cheng L (2012) Increased oceanic microplastic debris enhances oviposition in an endemic pelagic insect. Biology Letters 10.1098
2. Andrady AL (2011) Microplastics in the marine environment. Marine Pollution Bulletin 62: 1596-1605
3. To follow
4. To follow
5. Nenninger AA, Robinson LS, Hammer ND, Epstein EA, Badtke MP, Hultgren SJ, Chapman MR (2011) CsgE is a curli secretion specificity factor that prevents amyloid fibre aggregation. Mol Microbiol 81: 486-499
6. Mato Y, Isobe T, Takada H, Kanehiro H, Ohtake C, Kaminuma T (2001) Plastic Resin Pellets as a Transport Medium for Toxic Chemicals in the Marine Environment. Environ. Sci. Technol. 35: 318-324
7. https://2011.igem.org/Team:Peking_S/project/wire/harvest
8. http://www.xbase.ac.uk/genome/azoarcus-sp-bh72/NC_008702/azo2419;nahR1/viewer
9. http://kirschner.med.harvard.edu/files/bionumbers/fundamentalBioNumbersHandout.pdf
10. Park H, Lim W, Shin H (2005) In vitro binding of purified NahR regulatory protein with promoter Psal. Biochimica et Biophysica Acta 1775: 247-255
11. Shala AA, Restrepo S, Gonzalez Barrios AF (2011) A network model for biofilm development in Escherichia coli K-12. Theoretical Biology and Medical Modelling 8: 34 doi:10.1186/1742-4682-8-34
12. http://www.ecogene.org//?q=gene/EG12621
13. http://www.ecogene.org/?q=gene/EG11489
14. Kushner S (2002) mRNA Decay in Escherichia coli Comes of Age. J Bacteriol. 184: 4658-4665
Where does this go? Walsh DM, Lomakin A, Benedek GB, Condron MM, Teplow DB (1997) Amyloid b-Protein Fibrillogenesis: Detection of a protofibrillar intermediate. The Journal of Biological Chemistry 272: 22364–22372
 "
"