Team:LMU-Munich/Results
From 2012.igem.org
| (5 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
| - | === | + | ===Seven novel vectors were constructed and four already proven to work in ''B. subtilis''.=== |
| - | + | <br> | |
{| class="colored" | {| class="colored" | ||
!Name | !Name | ||
| Line 85: | Line 85: | ||
===15 Promoters evaluated in ''B. subtilis'' and added to the registry!=== | ===15 Promoters evaluated in ''B. subtilis'' and added to the registry!=== | ||
| - | + | <br> | |
<p align="justify"> This section gives an overview on the strength of all evaluated promoters, which span a large range of activities. For more details and informations of the experiments see the [https://2012.igem.org/Team:LMU-Munich/Data Data] page of the promoters. Note that P<sub>''veg''</sub> was not evaluated with luminescence measurements and this bar is just projected from the results of the β-galactosidase assay.</p> | <p align="justify"> This section gives an overview on the strength of all evaluated promoters, which span a large range of activities. For more details and informations of the experiments see the [https://2012.igem.org/Team:LMU-Munich/Data Data] page of the promoters. Note that P<sub>''veg''</sub> was not evaluated with luminescence measurements and this bar is just projected from the results of the β-galactosidase assay.</p> | ||
<br> | <br> | ||
| Line 102: | Line 102: | ||
| - | ===Sporobeads - what protein do you want | + | ===Sporobeads - what protein do you want to display?=== |
| - | + | <br> | |
{| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:left;" | {| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:left;" | ||
| style="width: 70%;background-color: #EBFCE4;" | | | style="width: 70%;background-color: #EBFCE4;" | | ||
| Line 119: | Line 119: | ||
===Erase<sup>*</sup> germination ability! Knockout strains=== | ===Erase<sup>*</sup> germination ability! Knockout strains=== | ||
| - | + | <br> | |
{| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | {| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | ||
| style="width: 70%;background-color: #EBFCE4;" | | | style="width: 70%;background-color: #EBFCE4;" | | ||
| Line 128: | Line 128: | ||
{| style="color:black;" cellpadding="3" width="95%" cellspacing="0" border="0" align="left" style="text-align:left;" | {| style="color:black;" cellpadding="3" width="95%" cellspacing="0" border="0" align="left" style="text-align:left;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
| - | <font color="#000000"; size="2">'''The germination rates of ''B. subtilis'' germination mutants.''' All germination rates are relative to wild type 168 (W168), our positive control. The mutant ''spo0A''::tet is unable to form spores, and should therefore show no germination (our negative control). Mutants used were: <br /> | + | <font color="#000000"; size="2">'''The germination rates of ''B. subtilis'' germination mutants.''' All germination rates are relative to wild type 168 (W168), our positive control. W168 showed near 100% germination rates. The mutant ''spo0A''::tet is unable to form spores, and should therefore show no germination (our negative control). Our results were consistent with this expectation. Mutants used were: <br /> |
| - | '''W168''': | + | '''W168''': wild-type 168 <br /> |
| - | '''''spo0A''::tet''': | + | '''''spo0A''::tet''': a sporulation-deficient strain <br /> |
'''B29''': ''cwlD''::kan, ''sleB''::mls <br /> | '''B29''': ''cwlD''::kan, ''sleB''::mls <br /> | ||
'''B30''': ''gerD''::cm, ''sleB''::mls <br /> | '''B30''': ''gerD''::cm, ''sleB''::mls <br /> | ||
| Line 139: | Line 139: | ||
'''B43''': ''gerD''::cm, ''sleB''::mls, ''cwlJ''::spec <br /> | '''B43''': ''gerD''::cm, ''sleB''::mls, ''cwlJ''::spec <br /> | ||
'''B46''': ''cwlD''::kan, ''cwlJ''::spec, ''gerD''::cm, ''sleB''::mls <br /> | '''B46''': ''cwlD''::kan, ''cwlJ''::spec, ''gerD''::cm, ''sleB''::mls <br /> | ||
| - | '''B47''': ''gerD''::cm, ''sleB''::mls, ''cwlJ''::spec, ''cwlD''::kan</font> | + | '''B47''': ''gerD''::cm, ''sleB''::mls, ''cwlJ''::spec, ''cwlD''::kan<br></font> |
|} | |} | ||
|} | |} | ||
|} | |} | ||
| - | <p align="justify">*No germinating spores detected in 4.6x10<sup>9</sup> spores.</p> | + | <p align="justify">*No germinating spores detected in 4.6x10<sup>9</sup> spores. For a full explanation of these great results, see [https://2012.igem.org/Team:LMU-Munich/Data/Knockout All results].</p> |
===Inverter works!=== | ===Inverter works!=== | ||
| - | + | <br> | |
{| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | {| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | ||
| style="width: 70%;background-color: #EBFCE4;" | | | style="width: 70%;background-color: #EBFCE4;" | | ||
Latest revision as of 22:13, 26 September 2012
The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]


Here you will find the main results of our work.
Seven novel vectors were constructed and four already proven to work in B. subtilis.
| Name
BioBrick | E. coli res. | B. subt. res. | Insertion locus | Description | Cloning status | Works? |
|---|---|---|---|---|---|---|
| pSBBs1C [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823023 (BBa_K823023)] | Amp | Cm | amyE | empty | ||
| pSBBs4S [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823022 (BBa_K823022)] | Amp | Spec | thrC | empty | ||
| pSBBs2E [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823027 (BBa_K823027)] | Amp | MLS | lacA | empty | in work | not tested |
| pSBBs1C-lacZ [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823021 (BBa_K823021) ] | Amp | Cm | amyE | lacZ reporter | ||
| pSBBs3C-luxABCDE [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823025 (BBa_K823025)] | Amp | Cm | sacA | luxABCDE reporter | ||
| pSBBs4S-PXyl [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823024 (BBa_K823024)] | Amp | Spec | thrC | Xylose-promoter | refuses transformation | |
| pSBBs0K-Pspac [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823026 (BBa_K823026)] | Amp | Kan | replicative | IPTG-promoter | unexpected | |
| Sporovector [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823026 (BBa_K823054)] | Amp | Spec | thrC | to make Sporobeads | not tested |
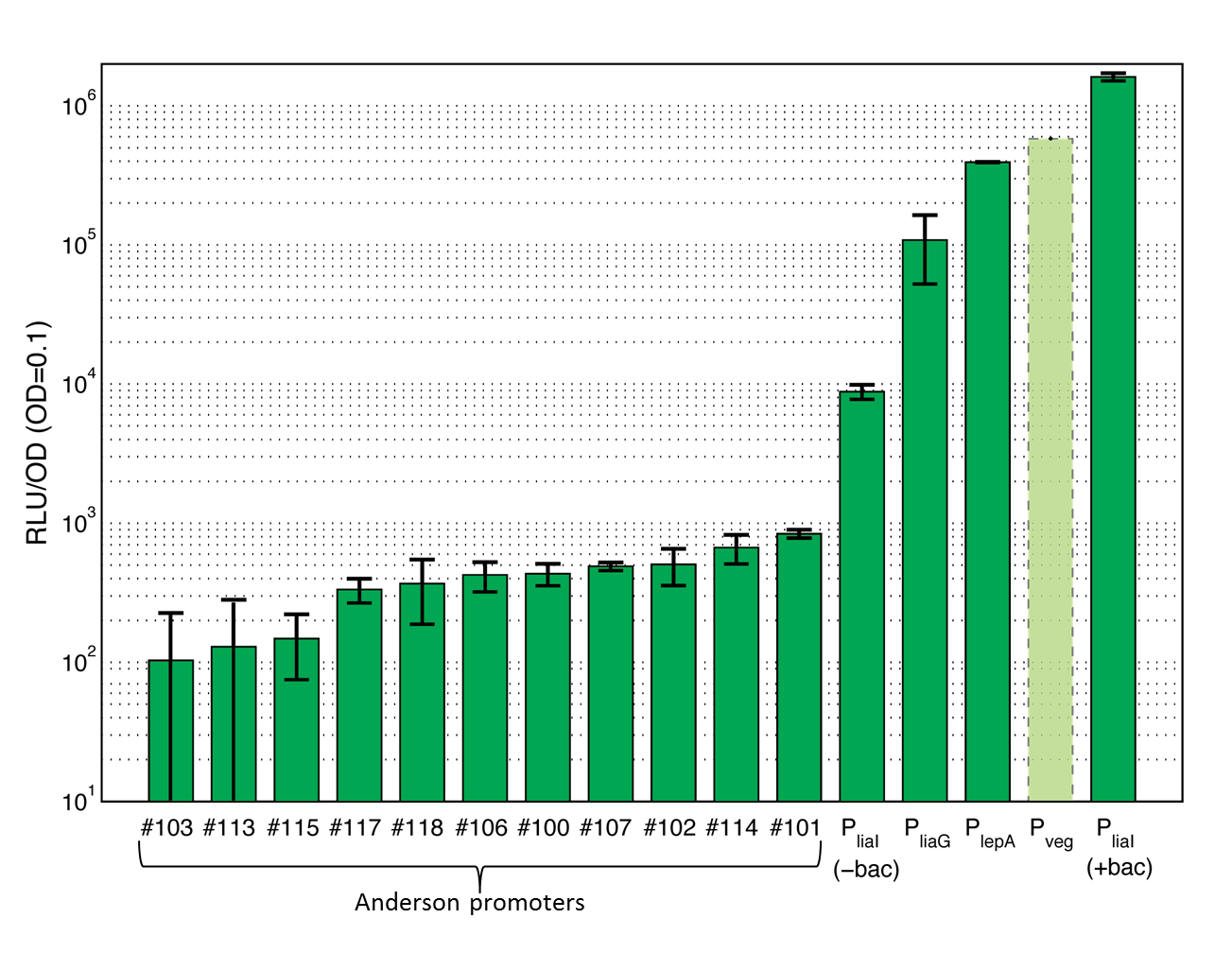
15 Promoters evaluated in B. subtilis and added to the registry!
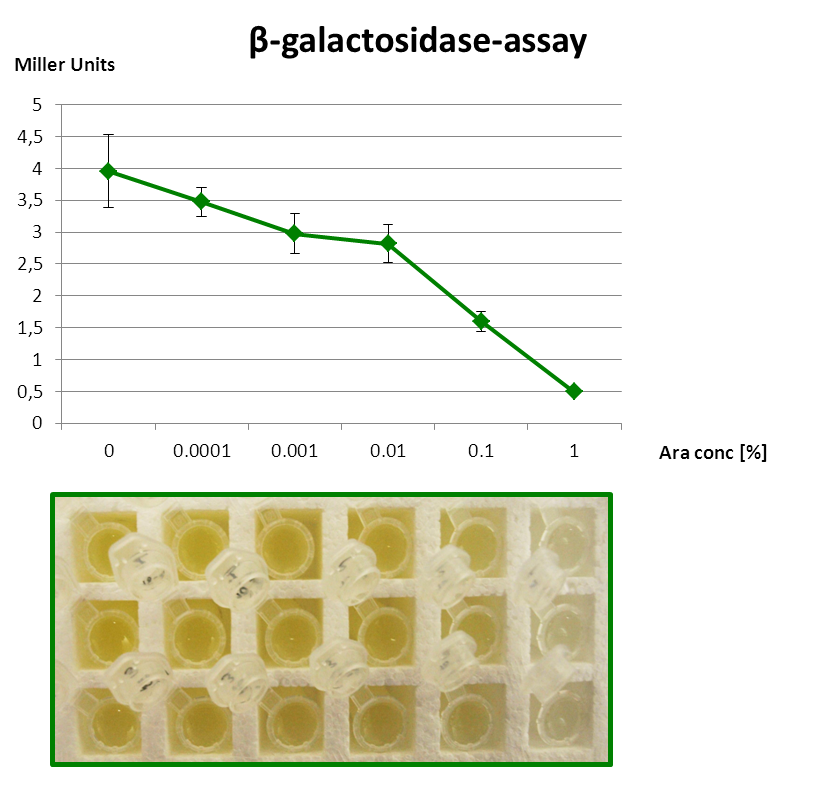
This section gives an overview on the strength of all evaluated promoters, which span a large range of activities. For more details and informations of the experiments see the Data page of the promoters. Note that Pveg was not evaluated with luminescence measurements and this bar is just projected from the results of the β-galactosidase assay.
|
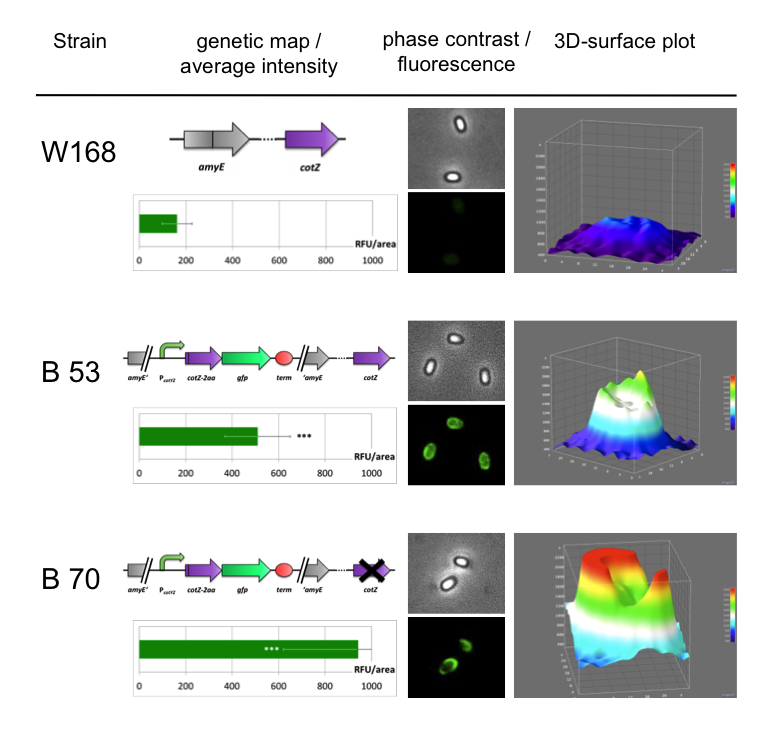
Sporobeads - what protein do you want to display?
|
| |
|
Erase* germination ability! Knockout strains
|
*No germinating spores detected in 4.6x109 spores. For a full explanation of these great results, see All results.
Inverter works!
|
For detailed results visit our data page and for all background info our project page.
 "
"