|
|
| (77 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| | <!-- Include the next line at the beginning of every page --> | | <!-- Include the next line at the beginning of every page --> |
| | {{:Team:LMU-Munich/Templates/Page Header|File:Team-LMU_eppis.resized.jpg|3}} | | {{:Team:LMU-Munich/Templates/Page Header|File:Team-LMU_eppis.resized.jpg|3}} |
| | + | [[File:LMU Glow Spore2 cutII.jpg|620px|link=]] |
| | | | |
| | | | |
| - | [[File:LMU Glow Spore2 cut.jpg|620px]] | + | [[File:SporeCoat.png|100px|right|link=]] |
| | | | |
| - | [[File:SporeCoat.png|100px|right|link=Team:LMU-Munich/Spore_Coat_Proteins]]
| |
| | | | |
| | | | |
| | + | =='''Sporo'''beads - What Protein Do ''You'' Want to Display?== |
| | + | <br> |
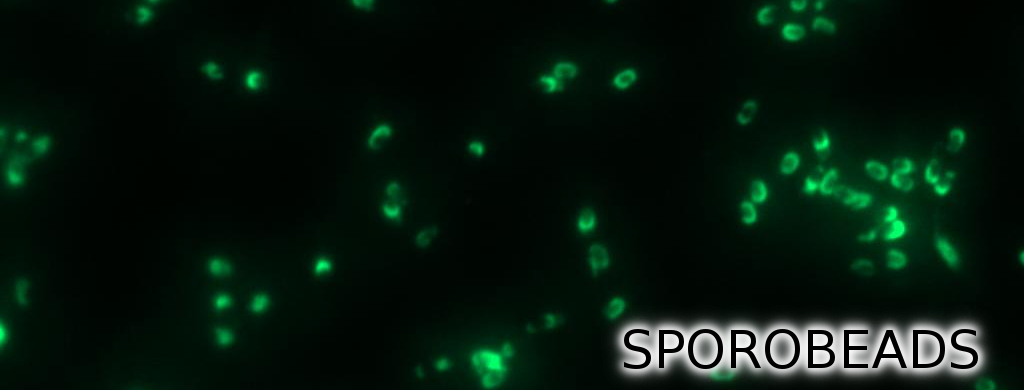
| | + | <p align="justify">'''Sporo'''beads are the first generation of ''Bacillus subtilis'' endospores displaying a protein of our choice on their outermost layer, the spore crust. In future '''Sporo'''beads could serve as a platform for protein display and thus be used for numerous versatile [https://2012.igem.org/Team:LMU-Munich/Application applications]. Our goal was to show that ''B. subtilis'' spores have the ability to do so. As a [https://2012.igem.org/Team:LMU-Munich/Spore_Coat_Proteins/result proof of principle] we successfully fused GFP to the spore crust and obtained fluorescence with microscopy.</p> |
| | | | |
| | | | |
| - | ==Spore Crust Proteins== | + | <div class="box"> |
| | + | ===Scientific Background=== |
| | + | {| "width=100%" style="text-align:center;" style="align:right"| |
| | + | |<p align="justify">Introduction to ''B. subtilis'' spores and their use in our project</p> |
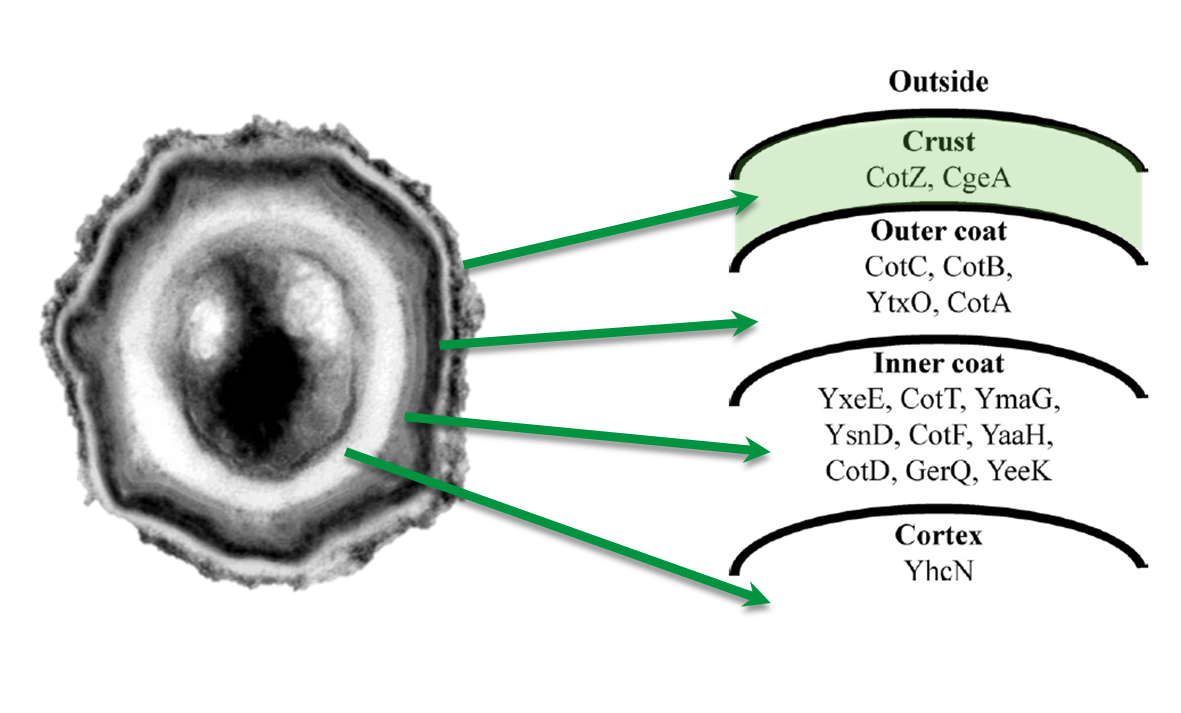
| | + | |[[File:Imamura, 2011 & McKenney, 2010.png|200px|right|link=Team:LMU-Munich/Spore_Coat_Proteins/Background]] |
| | + | |- |
| | + | ! colspan="2" |[[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Spore_Coat_Proteins/Background]] |
| | + | |} |
| | + | </div> |
| | | | |
| - | | + | <div class="box"> |
| - | <p align="justify">The aim of this module is to create spores that display fusion proteins on their crust. There are several different proteins forming the spore coat layers of ''Bacillus subtilis'' spores. On the outermost layer, the so called spore crust, the CotZ and CgeA proteins are located ([http://www.ncbi.nlm.nih.gov/pubmed?term=imamura%20et%20al.%202011%20spore%20crust Imamura et al., 2011]). This is why we used them to create functional fusion proteins to be expressed on our '''Sporo'''beads.</p> | + | ===Cloning Strategy=== |
| - | | + | {| "width=100%" style="text-align:center;" style="align:right"| |
| - | | + | |<p align="justify">Cloning strategy to create different variants of our ''' Sporo'''beads</p> |
| - | {| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:left;" | + | |[[File:Final construct.png|200px|link=Team:LMU-Munich/Spore_Coat_Proteins/cloning]] |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | |
| - | {|
| + | |
| - | |[[File:Imamura, 2011 & McKenney, 2010.png|Protein distribution in spore coat of ''Bacillus subtilis''|610px|center]] | + | |
| | |- | | |- |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | ! colspan="2" |[[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Spore_Coat_Proteins/cloning]] |
| - | {| style="color:black;" cellpadding="0" width="70%" cellspacing="0" border="0" align="center" style="text-align:center;"
| + | |
| - | |style="width: 70%;background-color: #EBFCE4;" | | + | |
| - | <font color="#000000"; size="2">Protein distribution in spore coat of ''Bacillus subtilis''</font>
| + | |
| - | |}
| + | |
| - | |}
| + | |
| | |} | | |} |
| | + | </div> |
| | | | |
| - | | + | <div class="box"> |
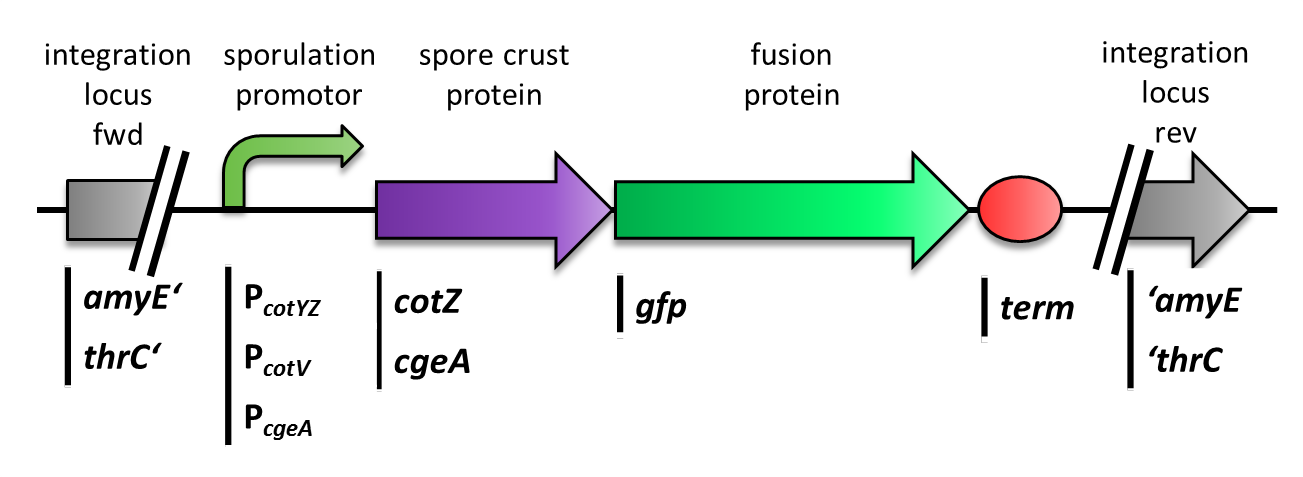
| - | <p align="justify">The gene ''cgeA'' is located in the ''cgeABCDE'' cluster and is regulated by its own promoter P<sub>''cgeA''</sub>. The cluster ''cotVWXYZ'' contains the gene ''cotZ'' which is cotranscribed with ''cotY'' regulated by the promoter P<sub>''cotYZ''</sub>. Another promoter of this cluster P<sub>''cotV''</sub> is responsible for the transcription of the other three genes. Those three promoters were [https://2012.igem.org/Team:LMU-Munich/Data/crustpromoters evaluated] with ''lux'' reporter genes to get an impression of their time of activation and their strength (see for more details [http://partsregistry.org/Part:BBa_K823025 pSB<sub>''Bs''</sub>3C-''lux''ABCDE]) so they could be used for expression of spore crust fusion proteins.</p> | + | ===GFP as a Proof of Principle=== |
| - | | + | {| "width=100%" style="text-align:center;" style="align:right"| |
| - | | + | |<p align="justify">Main results of the various constructs that were created to find the best one!</p> |
| - | {| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:left;" | + | |[[File:LMU Firstspore.jpg|200px|right|link=Team:LMU-Munich/Spore_Coat_Proteins/result]] |
| - | | style="width: 100%;background-color: #EBFCE4;" |
| + | |
| - | {|
| + | |
| - | |[[File:Operons.png|610px|center]] | + | |
| | |- | | |- |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | ! colspan="2" |[[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Spore_Coat_Proteins/result]] |
| - | {| style="color:black;" cellpadding="0" width="70%" cellspacing="0" border="0" align="center" style="text-align:center;"
| + | |
| - | |style="width: 70%;background-color: #EBFCE4;" | | + | |
| - | <font color="#000000"; size="2">Gene clusters of ''cotZ'' and ''cgeA''</font>
| + | |
| - | |}
| + | |
| - | |}
| + | |
| | |} | | |} |
| | + | </div> |
| | | | |
| - | | + | <div class="box"> |
| - | <p align="justify">The first step was to fuse [http://partsregistry.org/Part:BBa_K823039 ''gfp''] to ''cgeA'' and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823031 ''cotZ''] as a proof of principle. This way we would determine if it is possible to display proteins on the spore crust and if their expression has any effect on spore formation.</p> | + | ===Laccases as functional enzymes=== |
| - |
| + | {| "width=100%" style="text-align:center;" style="align:right"| |
| - | | + | |<p align="justify">Creation of functional Laccase-''' Sporo'''beads</p> |
| - | <p align="justify">Therefore we first fused ''cotZ'' to its two native promoters, P<sub>''cotV''</sub> and P<sub>''cotYZ''</sub>, and to P<sub>''cgeA''</sub>, which regulates the transcription of ''cgeA''. For cgeA we only used its native promoter P<sub>''cgeA''</sub> and the stonger one of the two promoters of the ''cotVWXYZ'' cluster, P<sub>''cotYZ''</sub> (for more details see [https://2012.igem.org/Team:LMU-Munich/Data/crustpromoters crust promotor evaluation]. While [http://partsregistry.org/Part:BBa_K823039 ''gfp''] was ligated to the terminator B0014 (see [http://partsregistry.org/wiki/index.php?title=Part:BBa_B0014 Registry]). When these constructs were finished and confirmed by sequencing, we fused them together applying the Freiburg standard to create in frame fusion proteins, flanked by one of the three promoters and the terminator.This way we created C-terminal fusion proteins.
| + | |[[File:Final construct.png|200px|link=Team:LMU-Munich/Spore_Coat_Proteins/laccases]] |
| - | <br>But as we did not know if C- or N-terminal fusion would influence the fusion protein expression, our second aim was to construct N-terminal fusion proteins as well. For this purpose we wanted to fuse the genes for the crust proteins ''cotZ'' and ''cgeA'' to the terminator and ''gfp'' to the three chosen promoters. Unsuccessfully, there occured a mutation in the XbaI site during construction of ''gfp'' in Freiburg Standard which is why we were not able to finish these constructs.
| + | |
| - | </p>
| + | |
| - | | + | |
| - | | + | |
| - | | + | |
| - | {| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:left;" | + | |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | |
| - | {|
| + | |
| - | |[[File:Final construct.png|610px|center]] | + | |
| - | |}
| + | |
| | |- | | |- |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | ! colspan="2" |[[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Spore_Coat_Proteins/laccases]] |
| - | {| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;"
| + | |
| - | |style="width: 70%;background-color: #EBFCE4;" | | + | |
| - | <font color="#000000"; size="2">Section of the genome of ''B. subtilis'' with the various integrated constructs.</font>
| + | |
| - | |}
| + | |
| | |} | | |} |
| | + | </div> |
| | | | |
| - | | + | <div class="box"> |
| - | <p align="justify">As we are working with B. subtilis spores, we needed to clone our final constructs into an empty Bacillus vector, so that they could get integrated into the genome of ''B. subtilis'' after transformation. Thus we picked the empty vector pSB<sub>BS</sub>1C from our '''''Bacillus''B'''io'''B'''rick'''B'''ox, for the ''cotZ'' constructs. This vector integrates into the ''amyE'' locus in the ''B. subtilis'' genome. Therefore we checked the integration of our construct via a starch test. The clones with the right integrated device have then been chosen for further analysis. In oder to express both crust protein constructs in one strain the ''cgeA'' fusion proteins had to be cloned into one of the other empty vectors. Unfortunately for unknown reasons, the cloning of the constructs with ''cgeA'' into this vector have been unsuccessful so far.</p> | + | ===Purification Methods=== |
| - | | + | {| "width=100%" style="text-align:center;" style="align:right"| |
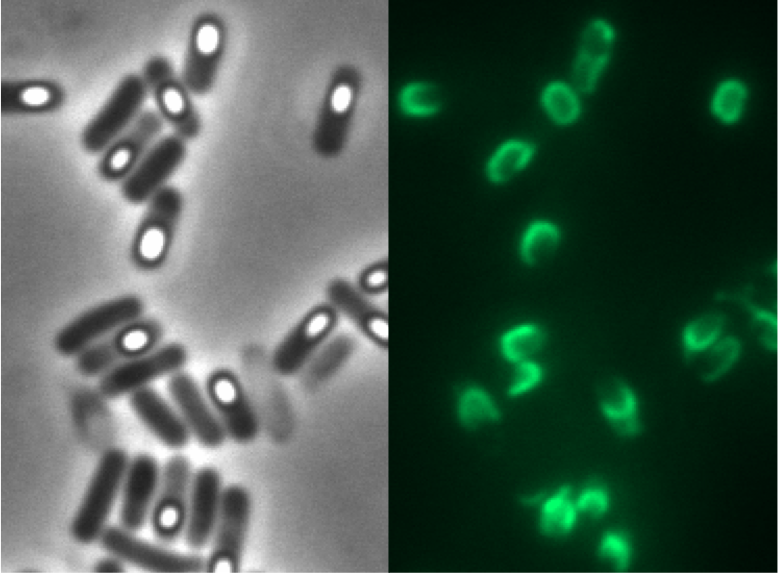
| - | <p align="justify">Finally we could start with the important experiment for our GFP-'''Sporo'''beads, fluorescence microscopy. Therefore we developed a sporulation protocol, that increases the rates of mature spores in our mutant samples (for details see link). The cells were fixed on agarose-pads and imaged in bright field and excited in blue wavelength. Because of the low but distinct fluorescence of wildtype sores, we measured and compared the fluorescence intensity of 100 spores per mutant. We obtained significant differences between wildtype spores and all our '''Sporo'''beads [link data]. We only worked with the P<sub>''cotYZ''</sub>-''cotZ''-''gfp''-''terminator'' spores for further experiments as these showed the brightest fluorescence.</p>
| + | |<p align="justify">Description of the different purification methods of the spores</p> |
| - | | + | |[[File:Treatments.png|200px|right|link=Team:LMU-Munich/Spore_Coat_Proteins/purification]] |
| - | [[File:Example.jpg PC+FL wt Pyz]]
| + | |
| - | | + | |
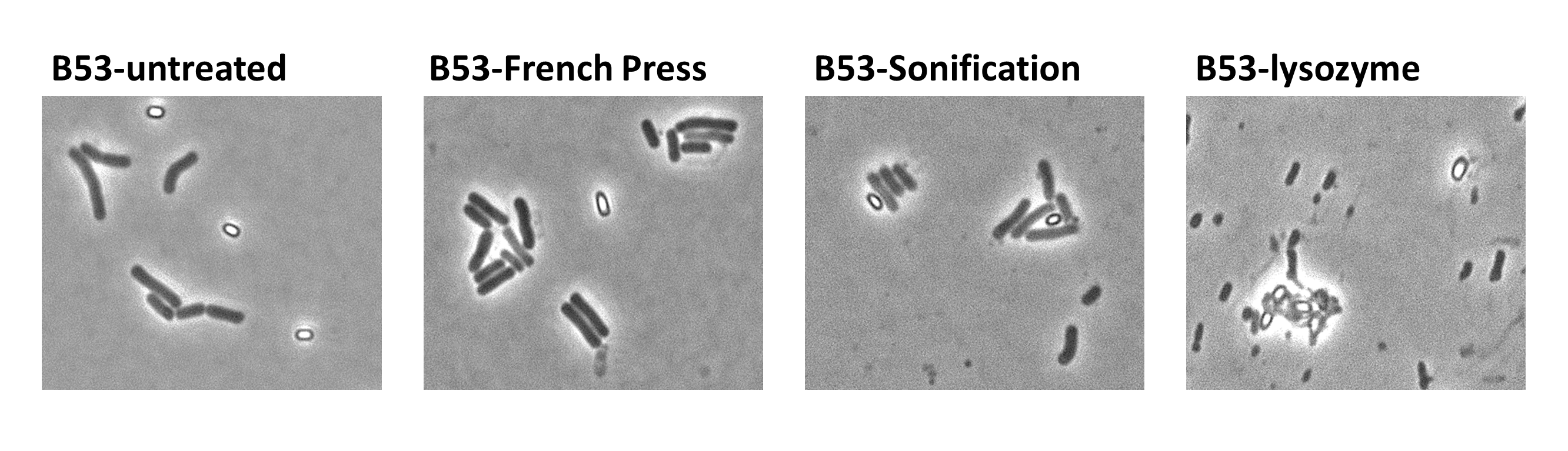
| - | <br>Since there were still some vegetative cells left after 24 hours of growth in DS-Medium, we wanted to purify the '''Sporo'''beads from them, which thereby should be deadened. We chose three different methods for this approach, the treatment with French Press, ultrasound (sonification) or lysozyme. By means of the microscopy results we were able to conclude that lysozyme treatment was the only successful method [link to data]. Additionally, it did not harm the crust fusion proteins as green fluorescence was detectable afterwards [link zu data]. This is why we use this treatment for purifying spores since.</p>
| + | |
| - | | + | |
| - | <p align="justify">Eventually, clean deletions of the native genes should reveal if there is any difference in fusion protein expression in our '''Sporo'''beads. Thus we deleted the native ''cotZ'' and ''cgeA'' using the cloning method described by [http://www.ncbi.nlm.nih.gov/pubmedterm=New%20Vector%20for%20Efficient%20Allelic%20Replacement%20in%20Naturally%20Nontransformable%2C%20Low-GC-Content%2C%20Gram-Positive%20Bacteria Arnaud ''et al''., 2004].</p>
| + | |
| - | | + | |
| - | | + | |
| - | {| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:left;" | + | |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | |
| - | {|
| + | |
| - | |[[File:Final construct 2.png|610px|center]] | + | |
| - | |}
| + | |
| | |- | | |- |
| - | | style="width: 70%;background-color: #EBFCE4;" |
| + | ! colspan="2" |[[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Spore_Coat_Proteins/purification]] |
| - | {| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;"
| + | |
| - | |style="width: 70%;background-color: #EBFCE4;" | | + | |
| - | <font color="#000000"; size="2">Section of the genome of ''B. subtilis'' with the various integrated constructs and the deletion of ''cotZ''.</font>
| + | |
| | |} | | |} |
| - | |}
| + | </div> |
| - | | + | <br> |
| - | | + | <br> |
| - | <p align="justify">The '''Sporo'''beads-Δ''cotZ'' were investigated by fluorescence microscopy and analysed like the other '''Sporo'''beads. The intensity bar charts should thereby show the fluorescence difference between wildtype (W168), B53- and B70-'''Sporo'''beads. To demonstrate the distribution of the fusion proteins we created 3D graphs, which show the fluorescence intensity spread across the spore surface. For analysis we measured the fluorescence intensity of a area of 750px per spore by using ImageJ and evaluated it with the statistical software '''R'''. The following graph shows the results of microscopy and ImageJ analysis.</p>
| + | <br> |
| - | | + | <br> |
| - | [[File:Example.jpg endbild]]
| + | |
| - | | + | |
| - | | + | |
| - | ==Applications==
| + | |
| - | | + | |
| - | <p align="justify">There are many possible applications for our '''Sporo'''beads and to easily create any kind of spore we designed a [https://2012.igem.org/Team:LMU-Munich/Bacillus_BioBricks/Sporovector '''Sporo'''vector]. With the help of this vector one can produce any individually desired spore. We picked three exemplary applications for our '''Sporo'''beads to be explained in detail: </p> | + | |
| - | | + | |
| - | | + | |
| - | <p align="justify"> | + | |
| - | <br>'''Kumamolisin:''' | + | |
| | <br> | | <br> |
| - | Kumamolisin is an [http://partsregistry.org/wiki/index.php?title=Part:BBa_K590087 enzyme] that cleaves peptides and was produced by the iGEM-Team from the University of [https://2011.igem.org/Team:Washington Washington]] last year. The substrate includes a specific sequence of amino acids which causes celiac disease in sensitive people when they consume food containing gluten. Our beads could carry this enzyme and offer a protected passage through the stomach, so that the enzyme can work where it is needed in the intestines. The <b>Germination</b>STOP we put in place in our spores would ensure a correct dosage. This project is a pharmaceutical application and therefore would have to fulfill the laws for pharmaceuticals. This includes several verification steps of non-toxicity and efficacy.
| |
| - | </p>
| |
| - |
| |
| - | <p align="justify">
| |
| - | B. subtilis spores are part of several dietary and health supplements and have a GRAS (Generally regarded as safe) status. Theoretically it might be possible to use the Sporobeads for dietary applications, but this includes several years of work and studying regulations. The [http://www.gesetze-im-internet.de/gcp-v/BJNR208100004.html GCP Verordnung] (“Good Clinical Practice ordinance”) has to be followed and the [http://www.pei.de/cln_101/DE/home/de-node.html?__nnn=true Paul-Ehrlich-Institute] would have to permit all the studies.
| |
| - | </p>
| |
| - |
| |
| - | '''CPX:'''
| |
| - | <p align="justify">
| |
| - | The [http://partsregistry.org/wiki/index.php/Part:BBa_I728500 CPX-peptide] generated by MIT (2007) can bind to Polystyrene which is a common plastic. The waste from Polystyrene is currently a big challenge in the field of environmental protection and remediation. In the ocean, large piece of Polystyrene litter are ground by sea currents into very small pieces, so called plastik plankton, that are consumed by fish, filter feeders, and other organisms living in the oceans. Such plastic uptake leads to poisoning, sterility and death. To remove Polystyrene from the oceans, huge filter boxes could be put into place to mechanically filter out large pieces; our CPX-Sporobeads could remove the microscopic plastic particles.
| |
| - | </p>
| |
| - |
| |
| - | <p align="justify">
| |
| - | Accidentally, some spores could loosen from the surface they would be affixed to and float into the sea. Therefore, we would have to ask the ZKBS([http://www.bvl.bund.de/DE/06_Gentechnik/03_Antragsteller/06_Institutionen_fuer_biologische_Sicherheit/01_ZKBS/gentechnik_zkbs_node.html Zentrale Kommission für biologische Sicherheit]; “Central Committee for Biological Safety”) for permission to release of the spores. They would require details about our spores and check whether our spores would be safe for environmental release.
| |
| - | Another application could be the removal of plastic particles in paper recycling facilities. The applicable laws and regulations should be similar.
| |
| - | </p>
| |
| - |
| |
| - | '''Virus binding:'''
| |
| - | <p align="justify">
| |
| - | Viruses bind specifically to certain proteins and domains to infect cells. To capitalize on this mechanism, the <b>Sporo</b>beads could be coated with such domains. The viruses would then bind to the spores and could be removed from liquid samples. This could be used to filter human immunodeficiency virus (HIV) from blood for transfusions, or even to help people with deadly virus infections by dialyzing their blood and thereby lowering their virus titer.
| |
| - | </p>
| |
| - |
| |
| - | <p align="justify">
| |
| - | Of course, permission from the ZKBS would be necessary, but because virus binding would be a medical application, rules from the medical field must be taken into account. The Robert-Koch-Institute in charge of hospital hygiene and infection prevention would check for the hygienic methods of such an application and whether it would work according to the [http://www.gesetze-im-internet.de/ifsg/ Infektionsschutzgesetz] (“prevention of infections law”).
| |
| - | </p>
| |
| - |
| |
| - | Consideration of medical laws would also be necessary, if our <b>Sporo</b>beads were to be used to filter viruses from drinking water.
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| - |
| |
| | <div class="box"> | | <div class="box"> |
| | ====Project Navigation==== | | ====Project Navigation==== |
| Line 154: |
Line 81: |
| | |} | | |} |
| | </div> | | </div> |
| | + | |
| | + | |
| | | | |
| | <!-- Include the next line at the end of every page --> | | <!-- Include the next line at the end of every page --> |
| | {{:Team:LMU-Munich/Templates/Page Footer}} | | {{:Team:LMU-Munich/Templates/Page Footer}} |







 "
"