Team:Paris-Saclay/Project/Notebook/Week 13
From 2012.igem.org
(Difference between revisions)
YohannPetiot (Talk | contribs) (Created page with "<div id="large-single-paris-saclay"> {{Team:Paris-Saclay/Header}} <div id="single-paris-saclay"> {{Team:Paris-Saclay/Menu}} <div id="single-left-column"> <html> <head> <l...") |
(→27th August) |
||
| (13 intermediate revisions not shown) | |||
| Line 23: | Line 23: | ||
</head> | </head> | ||
<body> | <body> | ||
| - | <div id="single- | + | <div id="tiles"> |
| - | <div | + | <div id="single-tile3" class="red live-tile" data-mode="flip" data-delay="4000"> |
| - | + | <div> | |
| - | + | <a href="https://2012.igem.org/Team:Paris-Saclay/Project/Abstract"> | |
| - | + | <div class="child-tile"><p class="child-tile">GEMOTE Project</p></div> | |
| + | <img src="https://static.igem.org/mediawiki/2012/7/7c/Project-1.png" alt="" /> | ||
| + | </a> | ||
| + | </div> | ||
<div> | <div> | ||
| - | <img src=" | + | <a href="https://2012.igem.org/Team:Paris-Saclay/Project/Abstract"> |
| - | + | <img src="https://static.igem.org/mediawiki/2012/d/d1/Project2.jpg" alt="" /> | |
| + | <div class="child-tile"><p class="child-tile">GEMOTE Project</p></div> | ||
| + | </a> | ||
</div> | </div> | ||
| - | <script type="text/javascript"> | + | </div> |
| + | <script type="text/javascript"> | ||
// apply regular slide universally unless .exclude class is applied | // apply regular slide universally unless .exclude class is applied | ||
// NOTE: The default options for each liveTile are being pulled from the 'data-' attributes | // NOTE: The default options for each liveTile are being pulled from the 'data-' attributes | ||
$(".live-tile, .flip-list").not(".exclude").liveTile(); | $(".live-tile, .flip-list").not(".exclude").liveTile(); | ||
| - | </script> | + | </script> |
| - | </div> | + | </div> |
</body> | </body> | ||
</html> | </html> | ||
| Line 43: | Line 49: | ||
</div> | </div> | ||
<div id="content-paris-saclay"> | <div id="content-paris-saclay"> | ||
| - | + | ='''Week 13'''= | |
| - | [[Category:Team:Paris-Saclay/Project Gemote/Notebook| | + | |
| + | [[Category:Team:Paris-Saclay/Project Gemote/Notebook|m]] | ||
__NOTOC__ | __NOTOC__ | ||
====27th August==== | ====27th August==== | ||
| Line 54: | Line 61: | ||
|- | |- | ||
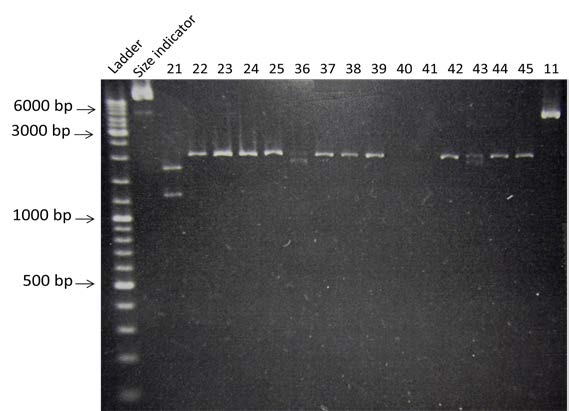
| style="width: 50%;"|Digestion by AseI of the 15 candidate colonies and #11. Visualization by electrophoresis on a 0.8% Agarose gel | | style="width: 50%;"|Digestion by AseI of the 15 candidate colonies and #11. Visualization by electrophoresis on a 0.8% Agarose gel | ||
| - | | style="width: 35%;"| [[File:Week13-2.jpg|right| | + | | style="width: 35%;"| [[File:Week13-2.jpg|right|380px]] |
|} | |} | ||
*Miniprep of colonies #11 and #21 | *Miniprep of colonies #11 and #21 | ||
| - | *Plasmids of colonies #11 and #21 have been | + | *Plasmids of colonies #11 and #21 have been sent for sequencing |
| Line 65: | Line 72: | ||
|- | |- | ||
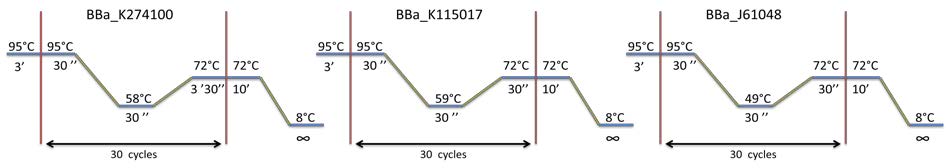
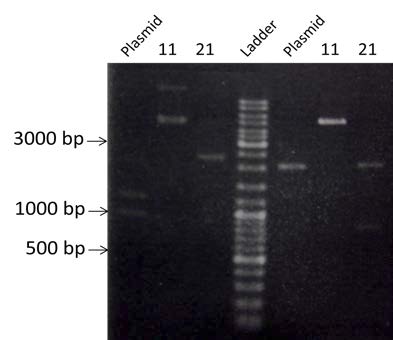
| style="width: 50%;"|Amplification of BBa_K274100, BBa_K115017 and BBa_J61048 by PCR using the colony #11. Visualization by electrophoresis on a 1% Agarose gel for BB3 and BB4 and a 2% Agarose gel for BB2. We are expecting a band at 123 bp for BBa_K115017, 133 bp for BBa_J61048 and 3408bp for BBa_K274100. | | style="width: 50%;"|Amplification of BBa_K274100, BBa_K115017 and BBa_J61048 by PCR using the colony #11. Visualization by electrophoresis on a 1% Agarose gel for BB3 and BB4 and a 2% Agarose gel for BB2. We are expecting a band at 123 bp for BBa_K115017, 133 bp for BBa_J61048 and 3408bp for BBa_K274100. | ||
| - | | style="width: 35%;"| [[File:Week13-3.jpg|right| | + | | style="width: 35%;"| [[File:Week13-3.jpg|right|350px]] |
|} | |} | ||
PCR program used for each biobrick amplification: | PCR program used for each biobrick amplification: | ||
| - | |||
| - | + | [[File:Week13-4.jpg|650px]] | |
| + | *Gibson assembly for the A construction (BBa_K098995+BBa_K274100+ BBa_J61048+Plasmid pSB1A2) and for the C construction (BBa_K115017 + BBa_C0051+166bp end of BBa_K098995+BBa_K274100+ BBa_J61048+Plasmid pSB1A2). Transformation of DH5αZ1 competent cell with construction A or construction C. Petri Dishes have been placed at 37°C for 1 day (construction A) or 2 days (construction C) | ||
====28th august==== | ====28th august==== | ||
| Line 129: | Line 136: | ||
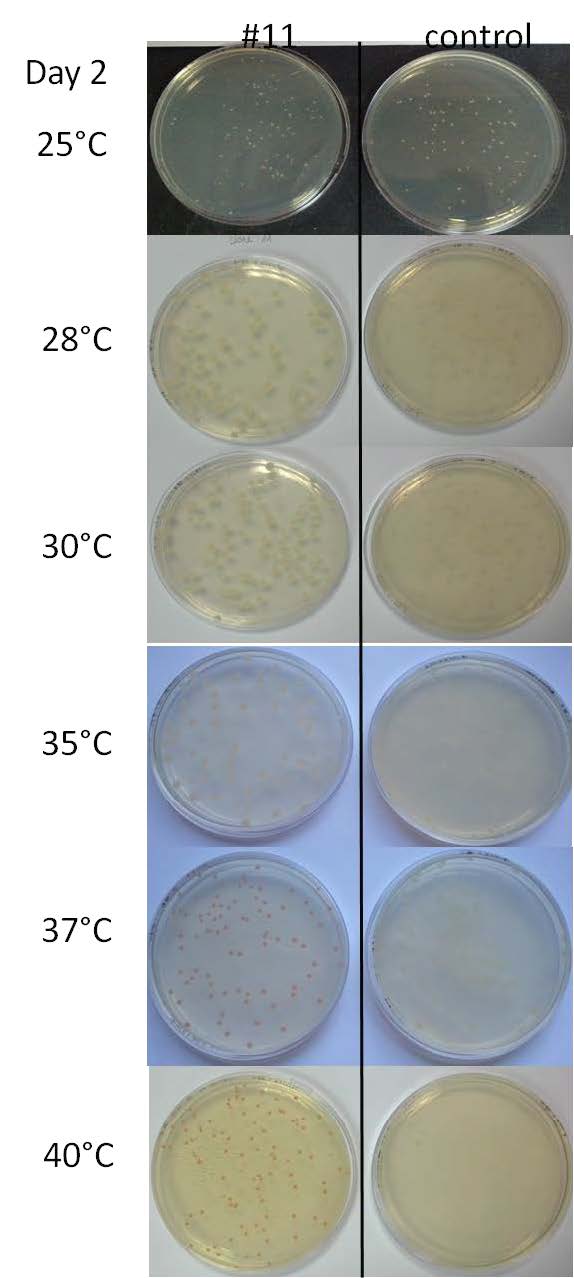
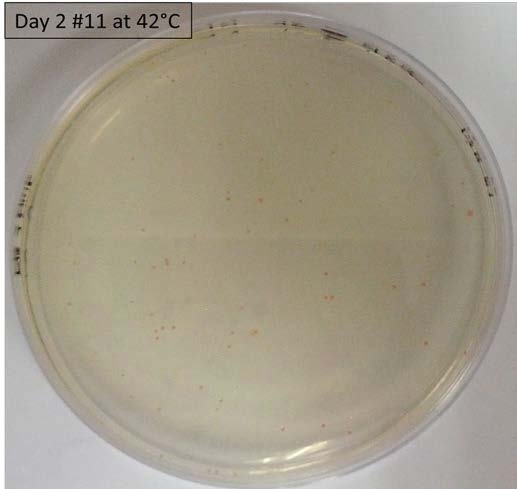
*Performing tests on a range of temperatures with the liquid culture of the colony #11 and the control 11811. On Petri Dishes LB+ agar+Ampicilline. 7 different temperatures have been tested: 25°C, 28°C, 30°C, 35°C, 37°C, 40°C and 42°C. | *Performing tests on a range of temperatures with the liquid culture of the colony #11 and the control 11811. On Petri Dishes LB+ agar+Ampicilline. 7 different temperatures have been tested: 25°C, 28°C, 30°C, 35°C, 37°C, 40°C and 42°C. | ||
| - | [[File:Week13-12.jpg]] | + | [[File:Week13-12.jpg|500px]] |
| - | [[File:Week13-13.jpg]] | + | [[File:Week13-13.jpg|500px]] |
| Line 138: | Line 145: | ||
| - | [[File:Week13-14.jpg| | + | [[File:Week13-14.jpg|500px]] |
| - | + | ||
====30th and 31th august==== | ====30th and 31th august==== | ||
| Line 146: | Line 152: | ||
| - | [[File:Week13-15.jpg| | + | [[File:Week13-15.jpg|600px]] |
PCR program used: | PCR program used: | ||
| - | [[File:Week13-16.jpg| | + | [[File:Week13-16.jpg|600px]] |
*Glycerol stock of the colonies #11 and #21. | *Glycerol stock of the colonies #11 and #21. | ||
| Line 157: | Line 163: | ||
{{Team:Paris-Saclay/Follow}} | {{Team:Paris-Saclay/Follow}} | ||
| + | </div> | ||
</div> | </div> | ||
</div> | </div> | ||
{{Team:Paris-Saclay/Footer}} | {{Team:Paris-Saclay/Footer}} | ||
Latest revision as of 00:45, 27 September 2012
Week 13
27th August
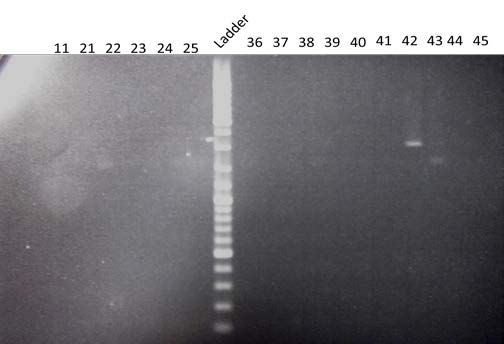
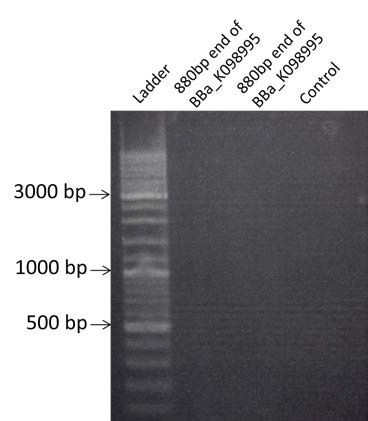
| Visualization by electrophoresis of the amplification of 880bp end of BBa_K098995 by PCR from the 15 candidate colonies and #11. Use of a 0.8% Agarose gel. We are expecting a band at 880 bp | |
| Digestion by AseI of the 15 candidate colonies and #11. Visualization by electrophoresis on a 0.8% Agarose gel |
- Miniprep of colonies #11 and #21
- Plasmids of colonies #11 and #21 have been sent for sequencing
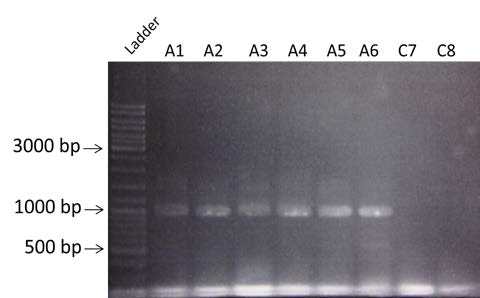
| Amplification of BBa_K274100, BBa_K115017 and BBa_J61048 by PCR using the colony #11. Visualization by electrophoresis on a 1% Agarose gel for BB3 and BB4 and a 2% Agarose gel for BB2. We are expecting a band at 123 bp for BBa_K115017, 133 bp for BBa_J61048 and 3408bp for BBa_K274100. |
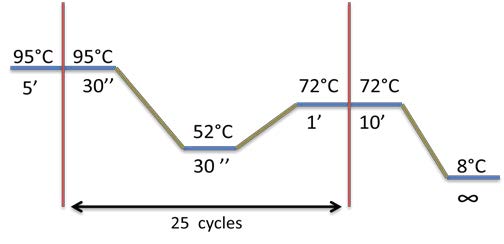
PCR program used for each biobrick amplification:
- Gibson assembly for the A construction (BBa_K098995+BBa_K274100+ BBa_J61048+Plasmid pSB1A2) and for the C construction (BBa_K115017 + BBa_C0051+166bp end of BBa_K098995+BBa_K274100+ BBa_J61048+Plasmid pSB1A2). Transformation of DH5αZ1 competent cell with construction A or construction C. Petri Dishes have been placed at 37°C for 1 day (construction A) or 2 days (construction C)
28th august
Digestion by AseI and NotI of the colonies #11, #21 and the linear plasmid.
- AseI:
- 1 restriction site in the Plasmid pSB1A2
- 1 restriction site in the 880bp end of BBa_K098995
- NotI:
- 1 restriction site either side of the B construction
| Visualization by electrophoresis on a 0.8% Agarose gel | |
| Amplification by PCR of the 880bp end of BBa_K098995. Visualization by electrophoresis on a 0.8% Agarose gel. We are expecting a band at 880bp. |
PCR program used:
- Amplification by PCR of the 880bp end of BBa_K098995 + BBa_K115017 contained in the colonies #11 and #21. PCR program used:
- Miniprep of the colony #11.
- Gibson assembly of the construction B (BBa_K115017+880bp end of BBa_K098995+ BBa_K274100+ BBa_J61048+ pSB1A2). Transformation of competent cells DH5α with the construction.
- Liquid culture of the colony #11 and a control colony 11811 to prepare a test of temperature range.
| Digestion of the colony #11 by AseI+HindIII. Visualization by electrophoresis on a 0.8% Agarose gel. |
29th August
| PCR on colonies to verify the construction A and C obtained by Gibson assembly. Amplification of BBa_K098995 for A and BBa_C0051 for C. Visualization by electrophoresis on a 0.8% Agarose gel. We are expecting a band at 935bp for A and 750pb for C. |
- Miniprep of the construction A and the construction C.
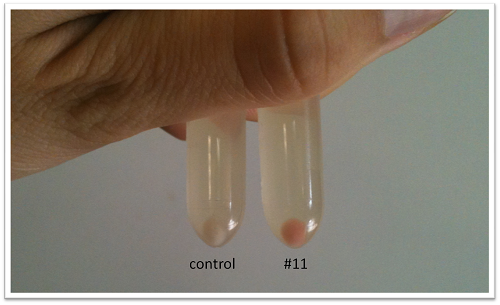
- Performing tests on a range of temperatures with the liquid culture of the colony #11 and the control 11811. On Petri Dishes LB+ agar+Ampicilline. 7 different temperatures have been tested: 25°C, 28°C, 30°C, 35°C, 37°C, 40°C and 42°C.
- The remaining of liquid cultures has been centrifuged to see the residue.
30th and 31th august
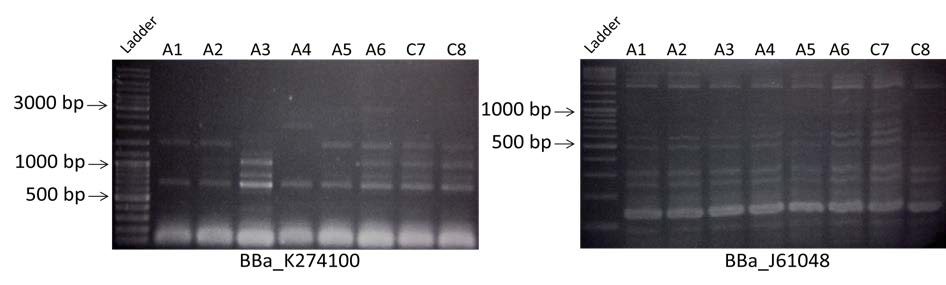
- Amplification of BBa_K274100 and BBa_J61048 of the colonies containing the construction A or the construction C. Visualization by electrophoresis on a 0.8% Agarose gel for BBa_K274100 and a 2% Agarose gel for BBa_J61048.
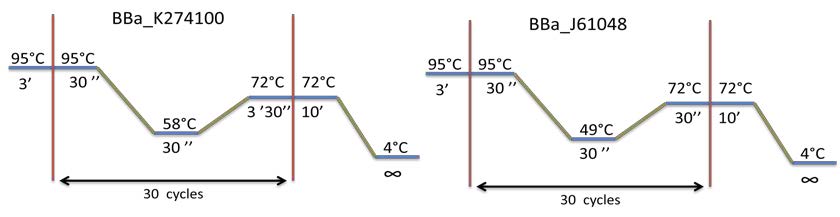
PCR program used:
- Glycerol stock of the colonies #11 and #21.
 "
"








Follow us !