Team:Washington/Plastics
From 2012.igem.org
| (212 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{Template:Team:Washington/Templates/Top}} | {{Template:Team:Washington/Templates/Top}} | ||
__NOTOC__ | __NOTOC__ | ||
| - | |||
==<i>“Plastics: made to last forever, designed to throw away”--5gyres.org</i>== | ==<i>“Plastics: made to last forever, designed to throw away”--5gyres.org</i>== | ||
| - | + | ||
| + | <h1 id='Background'>Background</h1> | ||
| + | [[Image:Washington_Sadfish.png|border|440px|right|thumb|Much of the plastic waste in the environment accumulates in the ocean, where the toxic plasticizers and additives contaminate the water.]] | ||
| + | Playing an integral role in our modern world, plastics account for over a third of products made today. They have a relatively low cost of production, serve to promote the development of industry, lower the cost of consumer goods, and are easy to produce on a large scale. Plastics, regardless of disposability, compose 10% of the waste generated in the world and an even higher percentage is carelessly thrown into our environment<html><a href="#Sources"><font size="1"><sup>[1]</sup></font></a></html>. Plastic pollution discharges debris into the ocean that are ingested by wildlife, often resulting in injury or death<html><a href="#Sources"><font size="1"><sup>[2]</sup></font></a></html>. | ||
===The PUR Problem === | ===The PUR Problem === | ||
| - | + | A commonly used plastic, Polyurethane (PUR), is used to create a wide range of products including dense solids, rigid insulating foam, and thermoset elastics. Because of its extreme versatility, the world consumption of PUR is continually increasing. According to current market analysis, the global production of PUR is estimated at 27 billion pounds per year and is expected to increase to 36 billion pounds by the year 2016<html><a href="#Sources"><font size="1"><sup>[3]</sup></font></a></html>. The leading method of recycling polyurethane is to mechanically grind the waste and rebind it into carpet cushioning, accounting for about 4% of waste polyurethane<html><a href="#Sources"><font size="1"><sup>[4]</sup></font></a></html>. However, due to the high costs of transportation and chemical degradation, most PURs are incinerated to recapture some of the energy used to make them<html><a href="#Sources"><font size="1"><sup>[5]</sup></font></a></html>. | |
| + | |||
| + | [[Image:Washington_PURBURN.png|border|280px|left|thumb|Incineration is currently the most popular means of waste PUR disposal. This method is not only harmful to our environment, the products have been shown to be carcinogenic.]] | ||
===Current Means of Disposal are Undesirable=== | ===Current Means of Disposal are Undesirable=== | ||
| - | + | Non-biodegradable plastics are often disposed of through waste incineration as it is the most efficient method of degradation. However, a major issue with this is that the products of plastic burning, which include polychlorinated di-benzo-p-dioxins/furans and carbon dioxide, are known to be carcinogenic<html><a href="#Sources"><font size="1"><sup>[6]</sup></font></a></html>. Upon incineration, these gaseous products are released into the atmosphere and have the potential to cause future problems to human health as well as contribute to global warming. As an additional concern, plastics constitute large volumes of space as they are habitually disposed of in landfills. It has been shown that plasticizers and plastic additives leak from the these plastics and contaminate aquatic environments<html><a href="#Sources"><font size="1"><sup>[7]</sup></font></a></html>. Thus, because of the environmental damage that current disposal methods incur, there is a large need for safer methods of plastic degradation. | |
===Engineering Microbes to Degrade PUR=== | ===Engineering Microbes to Degrade PUR=== | ||
| - | + | To solve the problem of PUR recycling, we propose a bacteria that is both able to degrade PUR and subsist off of the products of degradation as its sole carbon source. To this end, we propose a two plasmid system. The first plasmid would have two genes. The first gene, polyeurethane esterase, will encode an enzyme that is able to break down the PUR polymer structure into two molecules, one of which, ethylene glycol, can diffuse across the membrane of the bacterium<html><a href="#Sources"><font size="1"><sup>[8]</sup></font></a></html>. The second gene, osmotic inducible protein Y (osmY), would encode a protein that fuses to PUR esterase and exports the enzyme through the cell membrane and into the supernatant<html><a href="#Sources"><font size="1"><sup>[9]</sup></font></a></html>. The second plasmid would have an operon composed of, glycolaldehyde reductase (fucO) and glycoaldehyde dehydrogenase (aldA), that allows the bacterium to use ethylene glycol as its central metabolite<html><a href="#Sources"><font size="1"><sup>[10]</sup></font></a></html>. This system would allow the bacteria to turn the plastics plaguing our landfills into bacterial biomass which would in turn degrade more PUR. | |
| - | To solve the problem of PUR recycling, we propose a bacteria that is able to | + | [[Image:Washington_Plastic_Overview.png|border|950px|center|thumb|The complete degradation pathway takes polyurethane, an undesirable waste product and converts it into a central metabolite for bacterium.]] |
| + | |||
| + | |||
| + | ---- | ||
| + | <h1 id='App'>Our App: The Turbidostat<html><a href="#App"><font size="3">[Top]</font></a></html></h1> | ||
| + | <center>[http://students.washington.edu/cnt/pmwiki/pmwiki.php?n=Main.HomePage <b><font size='4'>Click here for the Turbidostat Wiki</font></b>]</center> | ||
| + | [[Image:Washington_Turbidostat.png|border|400px|left|thumb|Schematic diagram of our turbidostat. It uses the optical density measurements to modulate how much media is pumped into the culture vessel every minute to maintain a constant cell density. Through this method of cell density control, we can measure how fast the cells are growing and thus gain an understanding for relative fitness increases thorough out the course of an experiment.]] | ||
| + | [[Image:Recycleapp.png|150px|right]] | ||
| + | To characterize and optimize growth of specific parts of our overall polyurethane degradation pathway we developed a <i><b>homemade software application (App)</b></i>. However, devices that can accomplish this are not commercially available. Additionally, details for such tools are often sparsely detailed and frequently require exotic reagents and specialized training. To circumvent these issues, we instead decided to use our expertise in electrical, biological, and computer engineering to create an application that could control cheaply and readily available hardware to achieve the desired directed evolution and characterization functionality. | ||
| + | |||
| + | |||
| + | Our App uniquely controls an assortment of hardware devices easily found online, creating a fully functional turbidostat. A turbidostat is a closed-loop continuous culture device that maintains a desired constant cell density and chemical environment. The constant environment allows for continuous log phase growth and constant selective pressures (in our case polyurethane or ethylene glycol), which enables proper characterization (seen through media input per dilution cycle) and reduces evolutionary trajectory drift. By maintaining constant selective pressures, the rate of evolution is also dramatically improved<html><a href="#Sources"><font size="1"><sup>[11]</sup></font></a></html>. By creating this turbidostat App (written in Python), given sufficient time, we are able to properly assess and optimize our individual parts as well as the entire polyurethane degrading circuit in a controlled and reliable manner. Please click the link for complete instructions on how to build your own turbidostat. | ||
| + | |||
---- | ---- | ||
<h1 id='Methods'>Methods <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | <h1 id='Methods'>Methods <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | ||
| - | |||
| - | |||
| - | |||
| + | [[Image:Washington_PUR.png|border|250px|left|thumb|To assemble the polyurethane esterase, we used 4 IDT gBlocks, each containing a unique set of homology sequences that only allow for one orientation of binding, and Gibsoned them together onto a pGA backbone.]] | ||
| + | |||
| + | ===PUR Esterase Assay=== | ||
| + | Polyurethane esterase degrades polyurethane by cleaving the ester bonds to produce one of a variety of monomers with isocyanate groups on either end and an ethylene glycol. For our circuit, we have chosen to use the esterase estCS2 ([http://partsregistry.org/Part:BBa_K892012 BBa_K892012]). Originally isolated and characterized by Kang et al, the sequence of estCS2 is available at [http://www.ncbi.nlm.nih.gov/nuccore/283462288 GenBank (GU256649)]<html><a href="#Sources"><font size="1"><sup>[8]</sup></font></a></html>. Since their DNA constructs were unavailable and the authors did not respond to requests, we synthetically constructed the entire 1.7kb estCS2 gene from four 500-base gBlocks provided by IDT using Gibson cloning. | ||
| - | |||
| - | |||
| + | [[Image:Washington_PUR_Assay.png|border|350px|right|thumb|To assemble for activity of polyurethane esterase, cells transformed with the PUR esterase plasmid were grown to | ||
| + | saturation, sonicated, then a piece of pre-weighed polyurethane foam was inserted into the culture tube.]] | ||
| + | To assay esterase activity we made one 50mL TB + kanamycin overnight culture for the cells containing the esterase plasmid and a 50mL TB overnight culture with untransformed cells. After incubating the cultures for a day, we spun them down to form a pellet. The supernatant was then poured out and then the remaining pellets were resuspended the in enough water for each culture to be equalized at an arbitrary OD of 1.4. Three milliliters of each resuspended culture was aliquoted out into three microcentrifuge tubes (1 mL of one culture in each tube, 6 total microcentrifuge tubes). These tubes, along with 6 microcentrifuge tubes that contained only 1mL of LB were then sonicated at an amplitude of 20 with 1 second pulses on and off for 30 seconds. After sonication we poured each lysate and LB aliquots into 25mL culture tubes containing 1mL of LB and a preweighed sample of foam. This gave us a total of 12, 25mL culture tubes containing a pre weighed foam piece with LB+lysate or LB+sonicated LB. These culture tubes were left on the bench top at room temperature overnight. In the morning we washed the foam with DI water and dried them in a 65º C incubator. After drying, we weighed the samples to see if there was any change in the mass of the foam samples. | ||
===Exporter Protein OsmY Assay=== | ===Exporter Protein OsmY Assay=== | ||
| - | Since polyurethane is a large polymer, larger than the cell itself, it cannot be imported for degradation and therefore our esterase must be exported. It has been shown in Bolkinsky, Gregory et al | + | Since polyurethane is a large polymer, larger than the cell itself, it cannot be imported for degradation and therefore our esterase must be exported. It has been shown in Bolkinsky, Gregory et al that a protein, osmotically inducible protein Y (osmY, [http://partsregistry.org/Part:BBa_K892008 BBa_K892008]), is naturally exported by <i>E. coli</i> [9]. Furthermore this paper shows that osmY can be used as an export tag in <i>E. coli</i> when it is translationally fused to a protein. |
| - | + | [[Image:Washington_osmY.png|border|348px|left|thumb|Our exporter protein (osmY) is linked to sfGFP through a serine glycine linker.]] | |
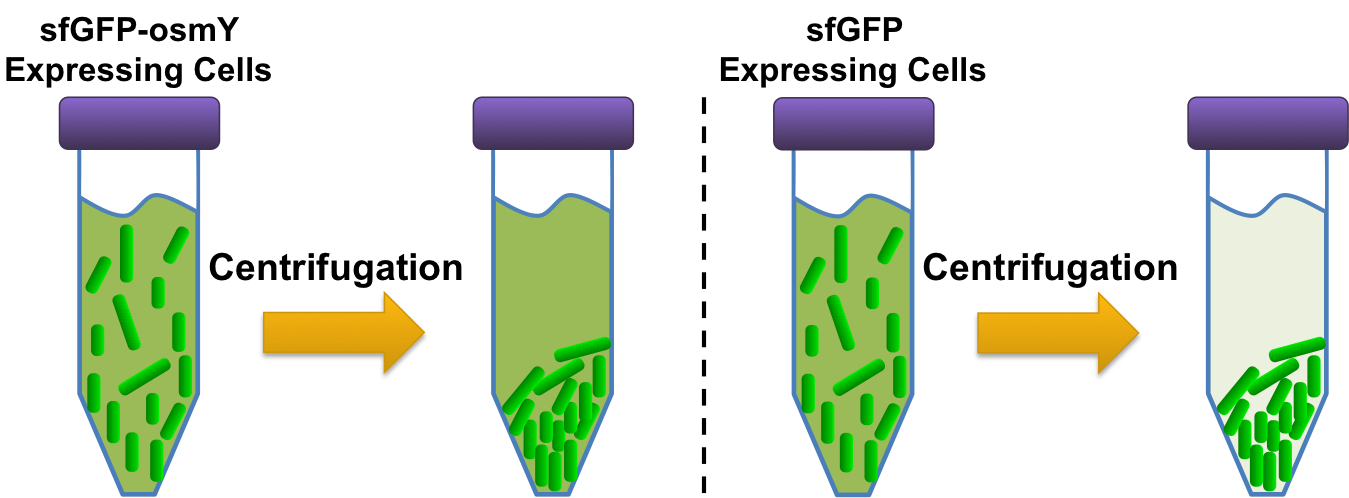
| + | [[Image:Washington_osmY_Assay.png|border|546px|right|thumb|We used osmY fused to sfGFP and as a control sfGFP fused to mamI in the same translational conformation to test for osmY functionality. These cultures were spun down and the fluorescence (480nm excitation, 509nm emission) of the cells+supernatant and supernatant were measured.]] | ||
| - | |||
| + | To assay the exportation system we created a fusion protein. The fusion protein consisted of super folder GFP (sfGFP) fused to osmY ([http://partsregistry.org/Part:BBa_K892008 BBa_K892011]), which allowed for visual assessment of whether or not osmY was correctly exported into the supernatant. For the control, sfGFP fused to mamI ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K590016 BBa_K590016]), a protein that binds to the cell membrane, was used. The sfGFP-osmY and sfGFP-mamI plasmids were transformed into BL21 and then were grown up overnight (after picking a colony of each from their transformation plates) in M9 1% glucose media. To quantify whether or not osmY properly exported proteins outside of the cell membrane, 4, 500µL aliquots of each overnight culture was made. From each of these aliquots, 200µL of the culture was pipetted into a 96 well plate and then the remaining culture was spun down in a centrifuge at 3000g for 3 minutes. Carefully, from each of the spun down aliquots, 200µL of the supernatant was pipetted into the 96 well plate. The remaining supernatant for each aliquot was disposed of and the cells were resuspended with 300 µL of fresh M9 1% glucose media. From the each of the resuspended aliquots, 200µL was pipetted into the 96 well plate. The 96 well plate was then read with a plate reader; each well was excited with a 480+/-20 nm light and then a detector quantified how much 509 +/-20 nm light was give off. | ||
| - | ===Turbidostat Assay=== | + | Using the information from the plate reader we could determine if osmY properly exports the fused protein. If the supernatant from the cells expressing the osmY construct had high fluorescence with respect to cells+supernatant while the supernatant of the control sfGFP-mamI had low fluorescence compared to its overall cells+supernatant fluorescence than it would be concluded that the protein was properly exported when fused to osmY. But if there was little to no supernatant fluorescence of the sfGFP-osmY construct or sfGFP-mamI and sfGFP-osmY illustrated the same ratio of supernatant fluorescence to cells+supernatant fluorescence then it would be concluded that osmY did not export proteins outside of the cell or our designed fusion between sfGFP and osmY disrupted sfGFP/osmY function. |
| - | + | ||
| + | |||
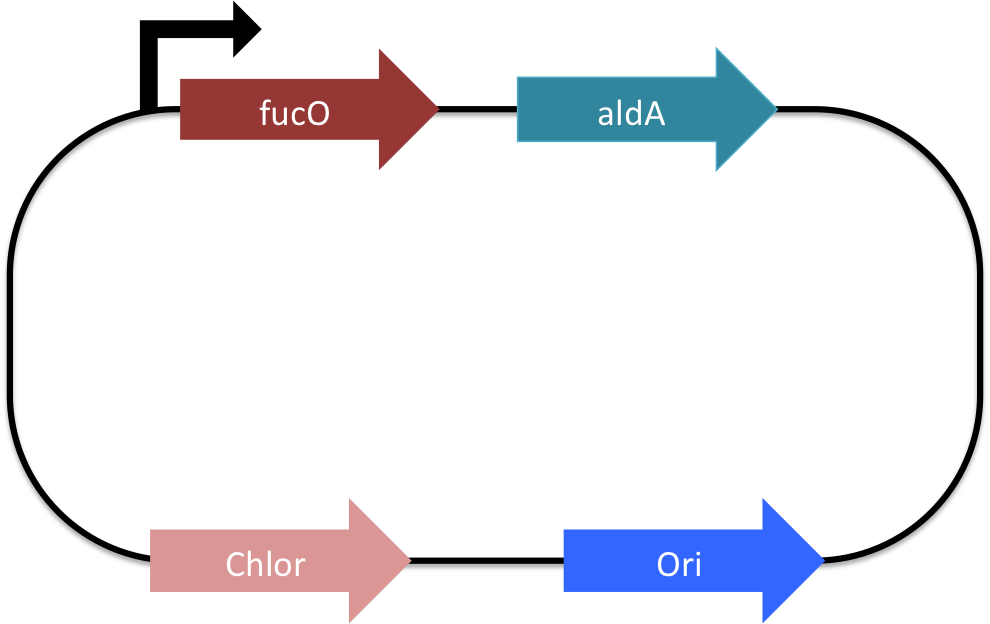
| + | [[Image:Washington_fucO_aldA.png|border|350px|left|thumb|By putting fucO and aldA onto a single plasmids with a single promote, we were able to create an operon that when transcribed and translated allows microbes to utilize ethylene glycol as a sole carbon source [10].]] | ||
| + | |||
| + | ===The Turbidostat Assay=== | ||
| + | The media used for the turbidostat assay was comprised exclusively of M9 salts and ethylene glycol. Because of this choice of media, the only carbon source available to <i>E. coli</i> was ethylene glycol and because this cannot be digested by normal lab strains of <i>E. coli</i>, only <i>E. coli</i> that can digest ethylene glycol will survive<html><a href="#Sources"><font size="1"><sup>[10]</sup></font></a></html>. The turbidostat App tracks cell growth by first blanking (setting optical density (OD) to 0) and then measuring and recording OD values after inoculation. Because OD and cell density are both linearly related, OD is a proxy for cell density measurements. After blanking on the M9 30mM ethylene glycol media, MG1655 cells that were transformed with fucO ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892009 BBa_K892009]) and aldA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892010 BBa_K892010]), which were grown in TB to log phase and then washed with PBS, were added to the turbidostat's culture vessel to an OD of 0.2. This allowed cells still in log phase to divide without exceeding the OD threshold of 0.3 (our desired constant cell density). After inoculation, the turbidostat was left to run overnight. Cell density and amount of media added per dilution cycle was captured in the morning. With this data, an understanding for how well the transformed <i>E. coli</i> subsists using ethylene glycol as its sole carbon source was gained. | ||
| + | |||
| + | |||
| + | The next assay ran was to transform MG1655 with a fucO-aldA operon plasmid ([http://partsregistry.org/Part:BBa_K892013 BBa_K892013]). This operon consisted of fucO and aldA being on a single plasmid that contains a single promoter. With this present, transformed MG1655 cells would be theoretically more viable in ethylene glycol since the cells would not have to use energy to create two antibiotic resistance genes (although no antibiotics were used in the the M9 ethylene glycol media, the cells still transcribe and translate the genes). By running the turbidostat in the same way as the dual transformation experiment, an idea of how well the transformed operon faired versus how well the two plasmid transformation could be gained. | ||
| - | |||
---- | ---- | ||
<h1 id='Results'>Results Summary <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | <h1 id='Results'>Results Summary <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | ||
| + | |||
| + | ===PUR Esterase Assay Results=== | ||
| + | We ran the PUR esterase assay and we found that the foam we used did not degrade. We quantified this by measuring the weight of the foam pieces to the nearest milligram before and after we ran the assay. We actually ended up leaving the lysate with the foam for three days to improve degradation but this had no effect on the end result. The reason we think the foam was not degraded was because we did not know the full composition of the foam, it could of had other polymers mixed in with the polyurethane that prevented our enzyme from properly degrading it. Further testing will need to be done to properly assess the functionality of our PUR esterase. | ||
| + | |||
| + | ===Exporter Protein OsmY Assay Results=== | ||
| + | [[Image:Washington_osmY_Assay_Results.png|border|940px|center|thumb|Each histogram depicts the average normalized fluorescence values for cells plus supernatant, cells only, and supernatant only. The image in the center is a visual representation of the presented data.]] | ||
| + | When running the osmY assay described in the methods section, we saw that sfGFP fused to osmY ([http://partsregistry.org/Part:BBa_K892008 BBa_K892011]) resulted in a much higher fluorescence ratio of cells+supernatant to supernatant than the fluorescence ratio of cells+supernatant to supernatant of sfGFP fused to mamI. Even though the cells only fluorescence of sfGFP-osmY plasmid exhibited nearly the same level of fluorescence as supernatant only, it does not go against the argument that osmY is properly exporting sfGFP into the supernatant. sfGFP-osmY is produced within the cell and it cannot be expected that all osmY will be secreted from the cell at a given moment. Thus from the above data, it can be gathered that <b>a protein fused to osmY will be exported into the supernatant.</b> | ||
| + | |||
| + | ===Turbidostat Assay Results=== | ||
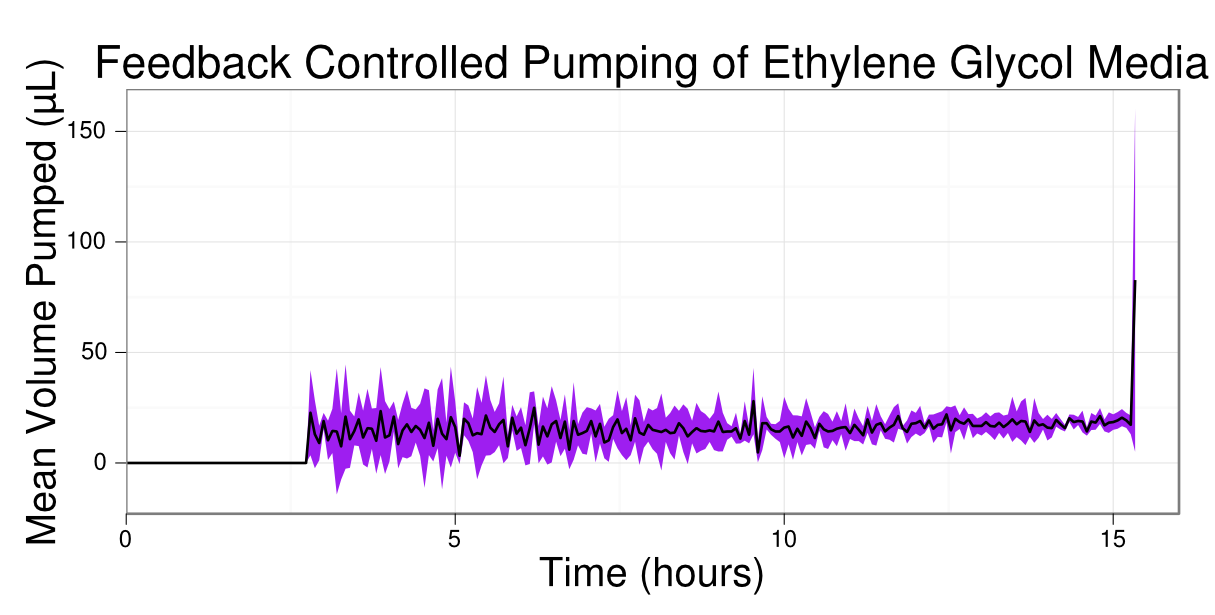
| + | [[Image:Washington_Turbidostat_Pump.png|border|450px|left|thumb|M9 30 mM ethylene glycol media injected into the culture vessel every minute to maintain the arbitrary optical density of 0.3.]] | ||
| + | |||
| + | [[Image:Washington_Turbidostat_OD.png|border|450px|right|thumb|Optical density measurements of <i>E. coli</i> transformed with fucO pGA3K3 + aldA pGA1C3 within the turbidostat growing on M9 30 mM ethylene glycol media.]] | ||
| + | |||
| + | |||
| + | Following the turbidostat assay protocol, we used transformed MG1655 with fucO ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892009 BBa_K892009]) on a medium copy Kanamycin resistance backbone and aldA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892010 BBa_K892010]) on a high copy Chloramphenicol resistance backbone and cultured it in the turbidostat. The turbidostat was operated overnight and the above data illustrates that the transformed MG1655 grew using ethylene glycol as its sole carbon source, which untransformed MG1655 cannot do. This is the case because the optical density (OD)/cell density rose from an initial arbitrary OD of 0.18 (time of inoculation) to an OD of 0.3, which was maintained throughout the time of the experiment (15 hours). The turbidostat maintained this OD by constantly varying the amount of new media put into the culture vessel. The amount added was reduced whenever the OD surpassed 0.3, which was quite frequently. Although time constraints did not allow us to transform MG1655 with our fucO-aldA operon and characterize it, it can be seen from the data above that when fucO and aldA genes are individually transformed and over expressed in MG1655 <i>E. coli</i>, <b><i>E. coli</i> gains the ability to utilize ethylene glycol as its sole carbon source.</b> The average time (based off of a 10mL culture volume and an average dilution rate of 30µL/min) for <i>E. coli</i> to undergo binary fission was 333 minutes, faster than the literature value of 360 minutes [10]. | ||
| + | |||
| + | |||
---- | ---- | ||
<h1 id='Future'>Future Directions <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | <h1 id='Future'>Future Directions <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | ||
| + | |||
| + | ===Putting it All together=== | ||
| + | The parts as they are right now have not been put into one cell strain and tested as a system. The obvious next step would be to test the efficacy of the entire plastic degrading construct against polyurethane. Using the turbidostat, we have the luxury of having an optimized and continual growth environment for our system to mutate <i>E. coli</i> towards improved functionality. Better cell strains would be periodically saved and tested in harsher selective conditions until eventually the strain is able to survive and reproduce off of polyurethane as its sole carbon source. We recognize that this may take many iterations to complete but the potential end product could be a cell that is able to consume plastic much faster, safer and more economically than traditional recycling methods. | ||
| + | |||
| + | ===Trash to Treasure=== | ||
| + | After stable growth off of plastic is achieved the next step would be to clone in a third plasmid that would produce some valuable commodity. Washington 2011 demonstrated that diesel fuel can be synthesized from central metabolites using their Petrobrick platform. The addition of the Petrobrick to the system would allow <i>E. coli</i> to process plastic waste and turn it into biofuels. | ||
| + | |||
| + | |||
---- | ---- | ||
<h1 id='Parts'>Parts Submitted <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | <h1 id='Parts'>Parts Submitted <html><a href="#Background"><font size="3">[Top]</font></a></html></h1> | ||
| + | |||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K892008 BBa_K892008 '''osmY'''] | ||
| + | |||
| + | The coding sequence for osmotically induced protein Y, a protein that when fused to another protein, gets exported out of <i>E. coli</i> cells. | ||
| + | |||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K892009 BBa_K892009 '''fucO'''] | ||
| + | |||
| + | The gene fucO, which codes for glycolaldehyde reductase, is one of the genes required for <i>E. coli</i> to utilize ethylene glycol as a food source. The coding sequence is put behind a strong biofab promoter and RBS. | ||
| + | |||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K892010 BBa_K892010 '''aldA'''] | ||
| + | |||
| + | The gene aldA, which codes for glycolaldehyde dehydrogenase, is the other gene required for <i>E. coli</i> to utilize ethylene glycol as a food source. The coding sequence is put behind a strong biofab promoter and RBS. | ||
| + | |||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K892011 BBa_K892011 '''sfGFP-osmY'''] | ||
| + | |||
| + | A composite part for osmotically induced protein Y fused downstream to sfGFP using a glycine - serine linker regulated by the lacI promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_R0011 BBa_R0011]) and the standard Elowitz RBS ([http://partsregistry.org/wiki/index.php?title=Part:BBa_B0034 BBa_B0034]). | ||
| + | |||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K892012 BBa_K892012 '''PUR Esterase'''] | ||
| + | |||
| + | An enzyme that breaks down polyurethane plastic behind the control of the lacI promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_R0011 BBa_R0011]) and the standard Elowitz RBS ([http://partsregistry.org/wiki/index.php?title=Part:BBa_B0034 BBa_B0034]). | ||
| + | |||
| + | [http://partsregistry.org/wiki/index.php?title=Part:BBa_K892013 BBa_K892013 '''fucO-aldA'''] | ||
| + | |||
| + | The combination of BBa_K892009 and BBa_K892010 behind a strong biofab promoter. | ||
| + | |||
| + | |||
---- | ---- | ||
| + | <html><a name="Sources"></a></html> | ||
<h1 id='Parts'>Sources <html><a href="#Sources"><font size="3">[Top]</font></a></html></h1> | <h1 id='Parts'>Sources <html><a href="#Sources"><font size="3">[Top]</font></a></html></h1> | ||
<html> | <html> | ||
| Line 56: | Line 134: | ||
<li>Gregory, M. R. "Environmental Implications of Plastic Debris in Marine Settings--entanglement, Ingestion, Smothering, Hangers-on, Hitch-hiking and Alien Invasions." Philosophical Transactions of the Royal Society B: Biological Sciences 364.1526 (2009): 2013-025. Print.</li> | <li>Gregory, M. R. "Environmental Implications of Plastic Debris in Marine Settings--entanglement, Ingestion, Smothering, Hangers-on, Hitch-hiking and Alien Invasions." Philosophical Transactions of the Royal Society B: Biological Sciences 364.1526 (2009): 2013-025. Print.</li> | ||
| - | <li>"Global Polyurethane Market to Reach 9.6 Mln Tons by 2015." Plastemart.com. N.p., 30 Aug. 2011. Web. <http://www.plastemart.com/Plastic-Technical-Article.asp?LiteratureID=1674></li> | + | <li>"Global Polyurethane Market to Reach 9.6 Mln Tons by 2015." Plastemart.com. N.p., 30 Aug. 2011. Web. <a href="http://www.plastemart.com/Plastic-Technical-Article.asp?LiteratureID=1674">http://www.plastemart.com/Plastic-Technical-Article.asp?LiteratureID=1674</a> </li> |
| - | <li>"Polyurethane Recycling." Polyurethanes. American Chemistry Council, n.d. Web. <http://polyurethane.americanchemistry.com/Sustainability/Recycling>. </li> | + | <li>"Polyurethane Recycling." Polyurethanes. American Chemistry Council, n.d. Web. <a href="http://polyurethane.americanchemistry.com/Sustainability/Recycling">http://polyurethane.americanchemistry.com/Sustainability/Recycling</a> </li> |
| - | <li>"Frequently Asked Questions on Polyurethanes." Polyurethanes.org. European Diisocyanate and Polyol Producers Association, n.d. Web. <http://www.polyurethanes.org/index.php?page=faqs>. </li> | + | <li>"Frequently Asked Questions on Polyurethanes." Polyurethanes.org. European Diisocyanate and Polyol Producers Association, n.d. Web. <a href="http://www.polyurethanes.org/index.php?page=faqs">http://www.polyurethanes.org/index.php?page=faqs</a> </li> |
<li>Takasuga, T., N. Umetsu, T. Makino, K. Tsubota, KS Sajwan, and KS Kumar. "Role of Temperature and Hydrochloric Acid on the Formation of Chlorinated Hydrocarbons and Polycyclic Aromatic Hydrocarbons during Combustion of Paraffin Powder, Polymers, and Newspaper." Archives of Environmental Contamination and Toxicology (2007): 8-21. Print. </li> | <li>Takasuga, T., N. Umetsu, T. Makino, K. Tsubota, KS Sajwan, and KS Kumar. "Role of Temperature and Hydrochloric Acid on the Formation of Chlorinated Hydrocarbons and Polycyclic Aromatic Hydrocarbons during Combustion of Paraffin Powder, Polymers, and Newspaper." Archives of Environmental Contamination and Toxicology (2007): 8-21. Print. </li> | ||
| Line 70: | Line 148: | ||
<li>Bokinsky, Gregory, Et. Al. "Synthesis of Three Advanced Biofuels from Ionic Liquid-penetreated Switchgrass Using Engineered Escherichia Coli." Proceedings of the National Academy of Sciences of the United States of America 108.50 (2011): 19949-9954. Print. </li> | <li>Bokinsky, Gregory, Et. Al. "Synthesis of Three Advanced Biofuels from Ionic Liquid-penetreated Switchgrass Using Engineered Escherichia Coli." Proceedings of the National Academy of Sciences of the United States of America 108.50 (2011): 19949-9954. Print. </li> | ||
| - | <li> Boronat, Albert, Estrella Caballero, and Juan Aguilar. "Experimental Evolution of a Metabolic Pathway for Ethylene Glycol Utilization by Escherichia Coli." Journal of Bacteriology Jan. (1983): 134-39. Web. </li> | + | <li> Boronat, Albert, Estrella Caballero, and Juan Aguilar. "Experimental Evolution of a Metabolic Pathway for Ethylene Glycol Utilization by Escherichia Coli." Journal of Bacteriology Jan. (1983): 134-39. Web. <a href="http://www.ncbi.nlm.nih.gov/pmc/articles/PMC217350/">http://www.ncbi.nlm.nih.gov/pmc/articles/PMC217350/</a> </li> |
| + | |||
| + | <li> E. Toprak, A. Veres, J. B. Michel, R. Chait, D. L. Hartl, R. Kishony, “Evolutionary paths to antibiotic resistance under dynamically sustained drug selection,” Nature Genetics, vol. 44, no. 1, pp. 101-105, Jan. 2012. </li> | ||
</ol> | </ol> | ||
</html> | </html> | ||
Latest revision as of 17:38, 15 November 2012
“Plastics: made to last forever, designed to throw away”--5gyres.org
Background
Playing an integral role in our modern world, plastics account for over a third of products made today. They have a relatively low cost of production, serve to promote the development of industry, lower the cost of consumer goods, and are easy to produce on a large scale. Plastics, regardless of disposability, compose 10% of the waste generated in the world and an even higher percentage is carelessly thrown into our environment[1]. Plastic pollution discharges debris into the ocean that are ingested by wildlife, often resulting in injury or death[2].
The PUR Problem
A commonly used plastic, Polyurethane (PUR), is used to create a wide range of products including dense solids, rigid insulating foam, and thermoset elastics. Because of its extreme versatility, the world consumption of PUR is continually increasing. According to current market analysis, the global production of PUR is estimated at 27 billion pounds per year and is expected to increase to 36 billion pounds by the year 2016[3]. The leading method of recycling polyurethane is to mechanically grind the waste and rebind it into carpet cushioning, accounting for about 4% of waste polyurethane[4]. However, due to the high costs of transportation and chemical degradation, most PURs are incinerated to recapture some of the energy used to make them[5].
Current Means of Disposal are Undesirable
Non-biodegradable plastics are often disposed of through waste incineration as it is the most efficient method of degradation. However, a major issue with this is that the products of plastic burning, which include polychlorinated di-benzo-p-dioxins/furans and carbon dioxide, are known to be carcinogenic[6]. Upon incineration, these gaseous products are released into the atmosphere and have the potential to cause future problems to human health as well as contribute to global warming. As an additional concern, plastics constitute large volumes of space as they are habitually disposed of in landfills. It has been shown that plasticizers and plastic additives leak from the these plastics and contaminate aquatic environments[7]. Thus, because of the environmental damage that current disposal methods incur, there is a large need for safer methods of plastic degradation.
Engineering Microbes to Degrade PUR
To solve the problem of PUR recycling, we propose a bacteria that is both able to degrade PUR and subsist off of the products of degradation as its sole carbon source. To this end, we propose a two plasmid system. The first plasmid would have two genes. The first gene, polyeurethane esterase, will encode an enzyme that is able to break down the PUR polymer structure into two molecules, one of which, ethylene glycol, can diffuse across the membrane of the bacterium[8]. The second gene, osmotic inducible protein Y (osmY), would encode a protein that fuses to PUR esterase and exports the enzyme through the cell membrane and into the supernatant[9]. The second plasmid would have an operon composed of, glycolaldehyde reductase (fucO) and glycoaldehyde dehydrogenase (aldA), that allows the bacterium to use ethylene glycol as its central metabolite[10]. This system would allow the bacteria to turn the plastics plaguing our landfills into bacterial biomass which would in turn degrade more PUR.
Our App: The Turbidostat[Top]

To characterize and optimize growth of specific parts of our overall polyurethane degradation pathway we developed a homemade software application (App). However, devices that can accomplish this are not commercially available. Additionally, details for such tools are often sparsely detailed and frequently require exotic reagents and specialized training. To circumvent these issues, we instead decided to use our expertise in electrical, biological, and computer engineering to create an application that could control cheaply and readily available hardware to achieve the desired directed evolution and characterization functionality.
Our App uniquely controls an assortment of hardware devices easily found online, creating a fully functional turbidostat. A turbidostat is a closed-loop continuous culture device that maintains a desired constant cell density and chemical environment. The constant environment allows for continuous log phase growth and constant selective pressures (in our case polyurethane or ethylene glycol), which enables proper characterization (seen through media input per dilution cycle) and reduces evolutionary trajectory drift. By maintaining constant selective pressures, the rate of evolution is also dramatically improved[11]. By creating this turbidostat App (written in Python), given sufficient time, we are able to properly assess and optimize our individual parts as well as the entire polyurethane degrading circuit in a controlled and reliable manner. Please click the link for complete instructions on how to build your own turbidostat.
Methods [Top]
PUR Esterase Assay
Polyurethane esterase degrades polyurethane by cleaving the ester bonds to produce one of a variety of monomers with isocyanate groups on either end and an ethylene glycol. For our circuit, we have chosen to use the esterase estCS2 ([http://partsregistry.org/Part:BBa_K892012 BBa_K892012]). Originally isolated and characterized by Kang et al, the sequence of estCS2 is available at [http://www.ncbi.nlm.nih.gov/nuccore/283462288 GenBank (GU256649)][8]. Since their DNA constructs were unavailable and the authors did not respond to requests, we synthetically constructed the entire 1.7kb estCS2 gene from four 500-base gBlocks provided by IDT using Gibson cloning.
To assay esterase activity we made one 50mL TB + kanamycin overnight culture for the cells containing the esterase plasmid and a 50mL TB overnight culture with untransformed cells. After incubating the cultures for a day, we spun them down to form a pellet. The supernatant was then poured out and then the remaining pellets were resuspended the in enough water for each culture to be equalized at an arbitrary OD of 1.4. Three milliliters of each resuspended culture was aliquoted out into three microcentrifuge tubes (1 mL of one culture in each tube, 6 total microcentrifuge tubes). These tubes, along with 6 microcentrifuge tubes that contained only 1mL of LB were then sonicated at an amplitude of 20 with 1 second pulses on and off for 30 seconds. After sonication we poured each lysate and LB aliquots into 25mL culture tubes containing 1mL of LB and a preweighed sample of foam. This gave us a total of 12, 25mL culture tubes containing a pre weighed foam piece with LB+lysate or LB+sonicated LB. These culture tubes were left on the bench top at room temperature overnight. In the morning we washed the foam with DI water and dried them in a 65º C incubator. After drying, we weighed the samples to see if there was any change in the mass of the foam samples.
Exporter Protein OsmY Assay
Since polyurethane is a large polymer, larger than the cell itself, it cannot be imported for degradation and therefore our esterase must be exported. It has been shown in Bolkinsky, Gregory et al that a protein, osmotically inducible protein Y (osmY, [http://partsregistry.org/Part:BBa_K892008 BBa_K892008]), is naturally exported by E. coli [9]. Furthermore this paper shows that osmY can be used as an export tag in E. coli when it is translationally fused to a protein.
To assay the exportation system we created a fusion protein. The fusion protein consisted of super folder GFP (sfGFP) fused to osmY ([http://partsregistry.org/Part:BBa_K892008 BBa_K892011]), which allowed for visual assessment of whether or not osmY was correctly exported into the supernatant. For the control, sfGFP fused to mamI ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K590016 BBa_K590016]), a protein that binds to the cell membrane, was used. The sfGFP-osmY and sfGFP-mamI plasmids were transformed into BL21 and then were grown up overnight (after picking a colony of each from their transformation plates) in M9 1% glucose media. To quantify whether or not osmY properly exported proteins outside of the cell membrane, 4, 500µL aliquots of each overnight culture was made. From each of these aliquots, 200µL of the culture was pipetted into a 96 well plate and then the remaining culture was spun down in a centrifuge at 3000g for 3 minutes. Carefully, from each of the spun down aliquots, 200µL of the supernatant was pipetted into the 96 well plate. The remaining supernatant for each aliquot was disposed of and the cells were resuspended with 300 µL of fresh M9 1% glucose media. From the each of the resuspended aliquots, 200µL was pipetted into the 96 well plate. The 96 well plate was then read with a plate reader; each well was excited with a 480+/-20 nm light and then a detector quantified how much 509 +/-20 nm light was give off.
Using the information from the plate reader we could determine if osmY properly exports the fused protein. If the supernatant from the cells expressing the osmY construct had high fluorescence with respect to cells+supernatant while the supernatant of the control sfGFP-mamI had low fluorescence compared to its overall cells+supernatant fluorescence than it would be concluded that the protein was properly exported when fused to osmY. But if there was little to no supernatant fluorescence of the sfGFP-osmY construct or sfGFP-mamI and sfGFP-osmY illustrated the same ratio of supernatant fluorescence to cells+supernatant fluorescence then it would be concluded that osmY did not export proteins outside of the cell or our designed fusion between sfGFP and osmY disrupted sfGFP/osmY function.
The Turbidostat Assay
The media used for the turbidostat assay was comprised exclusively of M9 salts and ethylene glycol. Because of this choice of media, the only carbon source available to E. coli was ethylene glycol and because this cannot be digested by normal lab strains of E. coli, only E. coli that can digest ethylene glycol will survive[10]. The turbidostat App tracks cell growth by first blanking (setting optical density (OD) to 0) and then measuring and recording OD values after inoculation. Because OD and cell density are both linearly related, OD is a proxy for cell density measurements. After blanking on the M9 30mM ethylene glycol media, MG1655 cells that were transformed with fucO ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892009 BBa_K892009]) and aldA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892010 BBa_K892010]), which were grown in TB to log phase and then washed with PBS, were added to the turbidostat's culture vessel to an OD of 0.2. This allowed cells still in log phase to divide without exceeding the OD threshold of 0.3 (our desired constant cell density). After inoculation, the turbidostat was left to run overnight. Cell density and amount of media added per dilution cycle was captured in the morning. With this data, an understanding for how well the transformed E. coli subsists using ethylene glycol as its sole carbon source was gained.
The next assay ran was to transform MG1655 with a fucO-aldA operon plasmid ([http://partsregistry.org/Part:BBa_K892013 BBa_K892013]). This operon consisted of fucO and aldA being on a single plasmid that contains a single promoter. With this present, transformed MG1655 cells would be theoretically more viable in ethylene glycol since the cells would not have to use energy to create two antibiotic resistance genes (although no antibiotics were used in the the M9 ethylene glycol media, the cells still transcribe and translate the genes). By running the turbidostat in the same way as the dual transformation experiment, an idea of how well the transformed operon faired versus how well the two plasmid transformation could be gained.
Results Summary [Top]
PUR Esterase Assay Results
We ran the PUR esterase assay and we found that the foam we used did not degrade. We quantified this by measuring the weight of the foam pieces to the nearest milligram before and after we ran the assay. We actually ended up leaving the lysate with the foam for three days to improve degradation but this had no effect on the end result. The reason we think the foam was not degraded was because we did not know the full composition of the foam, it could of had other polymers mixed in with the polyurethane that prevented our enzyme from properly degrading it. Further testing will need to be done to properly assess the functionality of our PUR esterase.
Exporter Protein OsmY Assay Results
When running the osmY assay described in the methods section, we saw that sfGFP fused to osmY ([http://partsregistry.org/Part:BBa_K892008 BBa_K892011]) resulted in a much higher fluorescence ratio of cells+supernatant to supernatant than the fluorescence ratio of cells+supernatant to supernatant of sfGFP fused to mamI. Even though the cells only fluorescence of sfGFP-osmY plasmid exhibited nearly the same level of fluorescence as supernatant only, it does not go against the argument that osmY is properly exporting sfGFP into the supernatant. sfGFP-osmY is produced within the cell and it cannot be expected that all osmY will be secreted from the cell at a given moment. Thus from the above data, it can be gathered that a protein fused to osmY will be exported into the supernatant.
Turbidostat Assay Results
Following the turbidostat assay protocol, we used transformed MG1655 with fucO ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892009 BBa_K892009]) on a medium copy Kanamycin resistance backbone and aldA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K892010 BBa_K892010]) on a high copy Chloramphenicol resistance backbone and cultured it in the turbidostat. The turbidostat was operated overnight and the above data illustrates that the transformed MG1655 grew using ethylene glycol as its sole carbon source, which untransformed MG1655 cannot do. This is the case because the optical density (OD)/cell density rose from an initial arbitrary OD of 0.18 (time of inoculation) to an OD of 0.3, which was maintained throughout the time of the experiment (15 hours). The turbidostat maintained this OD by constantly varying the amount of new media put into the culture vessel. The amount added was reduced whenever the OD surpassed 0.3, which was quite frequently. Although time constraints did not allow us to transform MG1655 with our fucO-aldA operon and characterize it, it can be seen from the data above that when fucO and aldA genes are individually transformed and over expressed in MG1655 E. coli, E. coli gains the ability to utilize ethylene glycol as its sole carbon source. The average time (based off of a 10mL culture volume and an average dilution rate of 30µL/min) for E. coli to undergo binary fission was 333 minutes, faster than the literature value of 360 minutes [10].
Future Directions [Top]
Putting it All together
The parts as they are right now have not been put into one cell strain and tested as a system. The obvious next step would be to test the efficacy of the entire plastic degrading construct against polyurethane. Using the turbidostat, we have the luxury of having an optimized and continual growth environment for our system to mutate E. coli towards improved functionality. Better cell strains would be periodically saved and tested in harsher selective conditions until eventually the strain is able to survive and reproduce off of polyurethane as its sole carbon source. We recognize that this may take many iterations to complete but the potential end product could be a cell that is able to consume plastic much faster, safer and more economically than traditional recycling methods.
Trash to Treasure
After stable growth off of plastic is achieved the next step would be to clone in a third plasmid that would produce some valuable commodity. Washington 2011 demonstrated that diesel fuel can be synthesized from central metabolites using their Petrobrick platform. The addition of the Petrobrick to the system would allow E. coli to process plastic waste and turn it into biofuels.
Parts Submitted [Top]
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K892008 BBa_K892008 osmY]
The coding sequence for osmotically induced protein Y, a protein that when fused to another protein, gets exported out of E. coli cells.
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K892009 BBa_K892009 fucO]
The gene fucO, which codes for glycolaldehyde reductase, is one of the genes required for E. coli to utilize ethylene glycol as a food source. The coding sequence is put behind a strong biofab promoter and RBS.
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K892010 BBa_K892010 aldA]
The gene aldA, which codes for glycolaldehyde dehydrogenase, is the other gene required for E. coli to utilize ethylene glycol as a food source. The coding sequence is put behind a strong biofab promoter and RBS.
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K892011 BBa_K892011 sfGFP-osmY]
A composite part for osmotically induced protein Y fused downstream to sfGFP using a glycine - serine linker regulated by the lacI promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_R0011 BBa_R0011]) and the standard Elowitz RBS ([http://partsregistry.org/wiki/index.php?title=Part:BBa_B0034 BBa_B0034]).
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K892012 BBa_K892012 PUR Esterase]
An enzyme that breaks down polyurethane plastic behind the control of the lacI promoter ([http://partsregistry.org/wiki/index.php?title=Part:BBa_R0011 BBa_R0011]) and the standard Elowitz RBS ([http://partsregistry.org/wiki/index.php?title=Part:BBa_B0034 BBa_B0034]).
[http://partsregistry.org/wiki/index.php?title=Part:BBa_K892013 BBa_K892013 fucO-aldA]
The combination of BBa_K892009 and BBa_K892010 behind a strong biofab promoter.
Sources [Top]
- Barnes, D. K. A., F. Galgani, R. C. Thompson, and M. Barlaz. "Accumulation and Fragmentation of Plastic Debris in Global Environments." Philosophical Transactions of the Royal Society B: Biological Sciences 364.1526 (2009): 1985-998. Print.
- Gregory, M. R. "Environmental Implications of Plastic Debris in Marine Settings--entanglement, Ingestion, Smothering, Hangers-on, Hitch-hiking and Alien Invasions." Philosophical Transactions of the Royal Society B: Biological Sciences 364.1526 (2009): 2013-025. Print.
- "Global Polyurethane Market to Reach 9.6 Mln Tons by 2015." Plastemart.com. N.p., 30 Aug. 2011. Web. http://www.plastemart.com/Plastic-Technical-Article.asp?LiteratureID=1674
- "Polyurethane Recycling." Polyurethanes. American Chemistry Council, n.d. Web. http://polyurethane.americanchemistry.com/Sustainability/Recycling
- "Frequently Asked Questions on Polyurethanes." Polyurethanes.org. European Diisocyanate and Polyol Producers Association, n.d. Web. http://www.polyurethanes.org/index.php?page=faqs
- Takasuga, T., N. Umetsu, T. Makino, K. Tsubota, KS Sajwan, and KS Kumar. "Role of Temperature and Hydrochloric Acid on the Formation of Chlorinated Hydrocarbons and Polycyclic Aromatic Hydrocarbons during Combustion of Paraffin Powder, Polymers, and Newspaper." Archives of Environmental Contamination and Toxicology (2007): 8-21. Print.
- Teuten, E. L., J. M. Saquing, D. R. U. Knappe, M. A. Barlaz, S. Jonsson, A. Bjorn, S. J. Rowland, R. C. Thompson, T. S. Galloway, R. Yamashita, D. Ochi, Y. Watanuki, C. Moore, P. H. Viet, T. S. Tana, M. Prudente, R. Boonyatumanond, M. P. Zakaria, K. Akkhavong, Y. Ogata, H. Hirai, S. Iwasa, K. Mizukawa, Y. Hagino, A. Imamura, M. Saha, and H. Takada. "Transport and Release of Chemicals from Plastics to the Environment and to Wildlife." Philosophical Transactions of the Royal Society B: Biological Sciences 364.1526 (2009): 2027-045. Print.
- Kang, Chul-Hyung. "A Novel Family VII Esterase with Industrial Potential from Compost Metagenomic Library." Microbial Cell Factories 10.41 (2011): n. pag. Print.
- Bokinsky, Gregory, Et. Al. "Synthesis of Three Advanced Biofuels from Ionic Liquid-penetreated Switchgrass Using Engineered Escherichia Coli." Proceedings of the National Academy of Sciences of the United States of America 108.50 (2011): 19949-9954. Print.
- Boronat, Albert, Estrella Caballero, and Juan Aguilar. "Experimental Evolution of a Metabolic Pathway for Ethylene Glycol Utilization by Escherichia Coli." Journal of Bacteriology Jan. (1983): 134-39. Web. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC217350/
- E. Toprak, A. Veres, J. B. Michel, R. Chait, D. L. Hartl, R. Kishony, “Evolutionary paths to antibiotic resistance under dynamically sustained drug selection,” Nature Genetics, vol. 44, no. 1, pp. 101-105, Jan. 2012.
 "
"