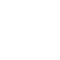Team:Tianjin/Project/OrthogonalSystem
From 2012.igem.org
(→RFP RBS Mutation) |
|||
| (16 intermediate revisions not shown) | |||
| Line 63: | Line 63: | ||
</div> | </div> | ||
<div class="menu_children"> | <div class="menu_children"> | ||
| - | <a href="https://2012.igem.org/Team:Tianjin/Project/ | + | <a href="https://2012.igem.org/Team:Tianjin/Project/Regulation">Logic Metabolism Regulation</a> |
</div> | </div> | ||
<div class="menu_children"> | <div class="menu_children"> | ||
| - | <a href="https://2012.igem.org/Team:Tianjin/Project/ | + | <a href="https://2012.igem.org/Team:Tianjin/Project/Gene">Future Work</a> |
| + | </div> | ||
| + | <div class="menu_children"> | ||
| + | <a href="https://2012.igem.org/Team:Tianjin/Data">BioBrick</a> | ||
| + | </div> | ||
| + | <div class="menu_children"> | ||
| + | <a href="https://2012.igem.org/Team:Tianjin/Project/Technology">Technology</a> | ||
</div> | </div> | ||
</div> | </div> | ||
| Line 139: | Line 145: | ||
[[file:TJU2012-Proj-OS-fig-7.png|thumb|500px|center|'''Figure 5.''' the Operon: RBS/RFP+RBS/RFP, n-RBS. n-RBS binds to the original RBS to translate RFP and GFP. (from TJU iGEM Team 2012)]] | [[file:TJU2012-Proj-OS-fig-7.png|thumb|500px|center|'''Figure 5.''' the Operon: RBS/RFP+RBS/RFP, n-RBS. n-RBS binds to the original RBS to translate RFP and GFP. (from TJU iGEM Team 2012)]] | ||
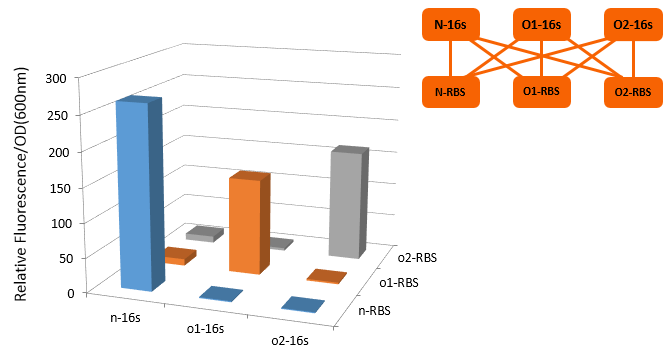
[[file:TJU2012-Proj-OS-fig-12.png|thumb|500px|center|'''Figure 6.''' The figure shows the result of our experiment. (from TJU iGEM Team 2012)]] | [[file:TJU2012-Proj-OS-fig-12.png|thumb|500px|center|'''Figure 6.''' The figure shows the result of our experiment. (from TJU iGEM Team 2012)]] | ||
| - | <span style="font-size:12px"><center>'''Table 1.''' The result of the experiment of the orthogonality verification. (from TJU iGEM Team 2012) | + | <span style="font-size:12px"><center>'''Table 1.''' The result of the experiment of the orthogonality verification. </center> (from TJU iGEM Team 2012)</span> |
<html> | <html> | ||
| Line 148: | Line 154: | ||
<tr> | <tr> | ||
| - | <th style="background:#fbb181;">Group</th> | + | <th style="background:#fbb181;"> Group </th> |
| - | <th style="background:#fbb181;">RFP-Plasmid</th> | + | <th style="background:#fbb181;"> RFP-Plasmid </th> |
<th style="background:#fbb181;">O-16S-Plasmid</th> | <th style="background:#fbb181;">O-16S-Plasmid</th> | ||
| - | <th style="background:#fbb181;">Result</th> | + | <th style="background:#fbb181;"> Result </th> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td style="background:#fdd8c0;">1</td> | + | <td style="background:#fdd8c0;"> 1</td> |
| - | <td style="background:#fdd8c0;">N-RBS</td> | + | <td style="background:#fdd8c0;"> N-RBS</td> |
| - | <td style="background:#fdd8c0;">N/A</td> | + | <td style="background:#fdd8c0;"> N/A</td> |
| - | <td style="background:#fdd8c0;">Expression</td> | + | <td style="background:#fdd8c0;"> Expression</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td>3</td> | + | <td> 3</td> |
| - | <td>O-RBS</td> | + | <td> O-RBS</td> |
| - | <td>Pbad+O-16S(No induced)</td> | + | <td> Pbad+O-16S(No induced) </td> |
| - | <td>N/A</td> | + | <td> N/A</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td style="background:#fdd8c0;">3'</td> | + | <td style="background:#fdd8c0;"> 3'</td> |
| - | <td style="background:#fdd8c0;">O-RBS</td> | + | <td style="background:#fdd8c0;"> O-RBS</td> |
| - | <td style="background:#fdd8c0;">N/A</td> | + | <td style="background:#fdd8c0;"> N/A</td> |
| - | <td style="background:#fdd8c0;">N/A</td> | + | <td style="background:#fdd8c0;"> N/A</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td>4</td> | + | <td> 4</td> |
| - | <td>O-RBS</td> | + | <td> O-RBS</td> |
| - | <td>Pbad+O-16S(No induced)</td> | + | <td> Pbad+O-16S(No induced) </td> |
| - | <td>Expression</td> | + | <td> Expression</td> |
</tr> | </tr> | ||
| Line 203: | Line 209: | ||
====Results==== | ====Results==== | ||
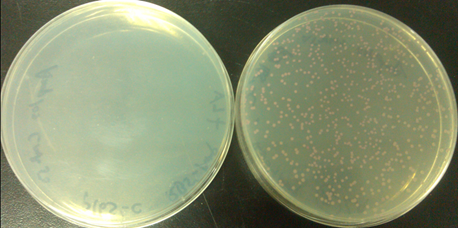
| - | [[file:TJU2012-Proj-SE-fig-3.png|thumb|500px|center|'''Figure | + | [[file:TJU2012-Proj-SE-fig-3.png|thumb|500px|center|'''Figure 8.''' Photo of two plates showing the normal cells (left) and orthogonal cells (right)(from TJU iGEM Team 2012)]] |
| - | + | <span style="font-size:12px"><center>'''Table 2.''' Chart showing the results (from TJU iGEM Team 2012)</center></span> | |
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <table border="0" | ||
| + | cellpadding="5" | ||
| + | style="border:2px solid #7b3101;"> | ||
| + | |||
| + | <tr> | ||
| + | <th style="background:#fbb181;"> Group </th> | ||
| + | <th style="background:#fbb181;"> PSB1A3-RFP </th> | ||
| + | <th style="background:#fbb181;"> O-16S </th> | ||
| + | <th style="background:#fbb181;"> Result </th> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background:#fdd8c0;"> 1(left)</td> | ||
| + | <td style="background:#fdd8c0;"> O-RBS+AmpR</td> | ||
| + | <td style="background:#fdd8c0;"> N/A</td> | ||
| + | <td style="background:#fdd8c0;"> No colony</td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td> 2(right)</td> | ||
| + | <td> O-RBS+AmpR</td> | ||
| + | <td> O-16S</td> | ||
| + | <td> Red colony</td> | ||
| + | </tr> | ||
| + | |||
| + | </table> | ||
| + | </center> | ||
| + | </html> | ||
| + | |||
From the two flats, we could easily find that the left plant plated without Ala,which means cells only containing normal ribosome. There is no colony on the plate. The reason is that without no orthogonal ribosome, Ampr gene could not be transcripted and translated,then cells could not survive on LB flat with Amp. On the other hand, the right flat, added Ala, containing orthogonal ribosome, have red colonies. Existing orthogonal ribosome means that cells could transcript and translate the Amp r gene, survive in Amp situation, the RFP could also be expressed with the help of normal ribosome. | From the two flats, we could easily find that the left plant plated without Ala,which means cells only containing normal ribosome. There is no colony on the plate. The reason is that without no orthogonal ribosome, Ampr gene could not be transcripted and translated,then cells could not survive on LB flat with Amp. On the other hand, the right flat, added Ala, containing orthogonal ribosome, have red colonies. Existing orthogonal ribosome means that cells could transcript and translate the Amp r gene, survive in Amp situation, the RFP could also be expressed with the help of normal ribosome. | ||
| Line 216: | Line 254: | ||
===Gibson Free Energy in the O-Key System=== | ===Gibson Free Energy in the O-Key System=== | ||
The strength of the interaction between SD and ASD sequence is thought to influence translational efficiency as mutations in either the SD or ASD sequence that weaken the interaction reduce the amount of protein made. The mechanism of protein expression is primarily determined by the delta Gibbs free energy of the combination of SD sequence on ribosome and the ASD sequence on RBS of the mRNA. Therefore, it is necessary to investigate the property of ΔG. | The strength of the interaction between SD and ASD sequence is thought to influence translational efficiency as mutations in either the SD or ASD sequence that weaken the interaction reduce the amount of protein made. The mechanism of protein expression is primarily determined by the delta Gibbs free energy of the combination of SD sequence on ribosome and the ASD sequence on RBS of the mRNA. Therefore, it is necessary to investigate the property of ΔG. | ||
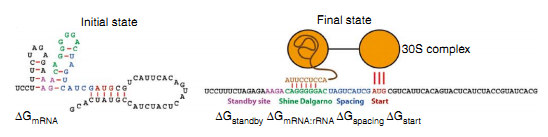
| - | [[file:TJU2012-Proj-OS-fig-4.png|thumb|500px|center|'''Figure | + | [[file:TJU2012-Proj-OS-fig-4.png|thumb|500px|center|'''Figure 9.''' The two states of 16S binding mRNA (from "Automated design of synthetic ribosome binding sites to control protein expression")]] |
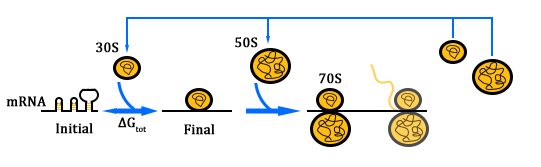
| - | [[file:TJU2012-Proj-OS-fig-6.png|thumb|500px|center|'''Figure | + | [[file:TJU2012-Proj-OS-fig-6.png|thumb|500px|center|'''Figure 10.''' Process of translation (from "Automated design of synthetic ribosome binding sites to control protein expression")]] |
ΔG<sub>tot</sub>=ΔG<sub>mRNA:rRNA</sub>+ΔG<sub>start</sub>+ΔG<sub>spacing</sub>-ΔG<sub>standby</sub>-ΔG<sub>mRNA</sub> | ΔG<sub>tot</sub>=ΔG<sub>mRNA:rRNA</sub>+ΔG<sub>start</sub>+ΔG<sub>spacing</sub>-ΔG<sub>standby</sub>-ΔG<sub>mRNA</sub> | ||
| Line 228: | Line 266: | ||
To calculate ΔG<sub>mRNA:rRNA</sub>, ΔG<sub>start</sub>, ΔG<sub>mRNA</sub> and ΔG<sub>standby</sub>, we use the NUPACK suite of algorithms with the Mfold 3.0 RNA energy parameters. These free energy calculations do not have any additional fitting or training parameters and explicitly depend on the mRNA sequence. In addition, the free energy terms are not orthogonal; changing a single nucleotide can potentially affect multiple energy terms. The relationship between the spacing and the ΔG<sub>spacing</sub> was empirically determined by measuring the protein expression level driven by synthetic RBSs of varying spacing and fitting a quantitative model to this data. | To calculate ΔG<sub>mRNA:rRNA</sub>, ΔG<sub>start</sub>, ΔG<sub>mRNA</sub> and ΔG<sub>standby</sub>, we use the NUPACK suite of algorithms with the Mfold 3.0 RNA energy parameters. These free energy calculations do not have any additional fitting or training parameters and explicitly depend on the mRNA sequence. In addition, the free energy terms are not orthogonal; changing a single nucleotide can potentially affect multiple energy terms. The relationship between the spacing and the ΔG<sub>spacing</sub> was empirically determined by measuring the protein expression level driven by synthetic RBSs of varying spacing and fitting a quantitative model to this data. | ||
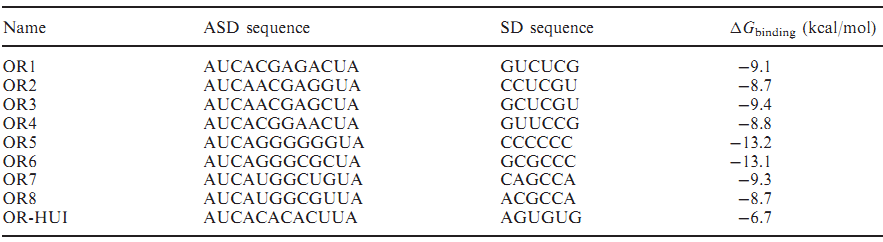
| - | + | <span style="font-size:12px"><center>'''Table 3.''' Orthogonal sequences of different ΔG (from "Computational design of orthogonal ribosomes")</center></span> | |
| + | [[file:TJU2012-Proj-OS-fig-5.png|500px|center]] | ||
In order to design the orthogonality and to predict the protein expression level, we have established the protein expression model which has strong relationship with the wet lab. The model is based on detailed study of the effects of changing Gibbs free energy on the translation process. | In order to design the orthogonality and to predict the protein expression level, we have established the protein expression model which has strong relationship with the wet lab. The model is based on detailed study of the effects of changing Gibbs free energy on the translation process. | ||
| Line 234: | Line 273: | ||
In the process of translation, the initiation rate is the rate determine step. The protein expression amount is consequently direct proportional to the initial translation speed. Furthermore, the initial translation speed is strongly related to the delta Gibbs free energy, which can be calculated out through results of many newly published papers and various commercialized software such as the Vienna RNA Package, RBS-Calculator, and Nucleic Acid Package. | In the process of translation, the initiation rate is the rate determine step. The protein expression amount is consequently direct proportional to the initial translation speed. Furthermore, the initial translation speed is strongly related to the delta Gibbs free energy, which can be calculated out through results of many newly published papers and various commercialized software such as the Vienna RNA Package, RBS-Calculator, and Nucleic Acid Package. | ||
In our model, we calculated the delta Gibbs energy. Ideal results can be obtained from the model. The orthogonality predicted by our model before the wet lab is similar to our prediction. When combined with some wet lab result, we can finally predict the spectroscope curve of RFP and GFP under three designed experimental state. For more details, see our [[Team:Tianjin/Modeling|Model Part]]. | In our model, we calculated the delta Gibbs energy. Ideal results can be obtained from the model. The orthogonality predicted by our model before the wet lab is similar to our prediction. When combined with some wet lab result, we can finally predict the spectroscope curve of RFP and GFP under three designed experimental state. For more details, see our [[Team:Tianjin/Modeling|Model Part]]. | ||
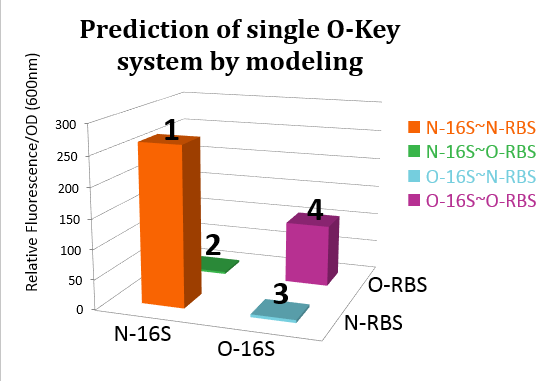
| - | [[file:TJU2012-Proj-OS-fig-14.png|thumb|500px|center|'''Figure | + | [[file:TJU2012-Proj-OS-fig-14.png|thumb|500px|center|'''Figure 11.''' RFP expression level under four pathways (from TJU iGEM Team 2012)]] |
We also want to add two O-Key system in our modeling and predict the result | We also want to add two O-Key system in our modeling and predict the result | ||
| - | [[file:TJU2012-Proj-OS-fig-10.png|thumb|500px|center|'''Figure | + | [[file:TJU2012-Proj-OS-fig-10.png|thumb|500px|center|'''Figure 12.''' The prediction of Two O-Key system by modeling. (from TJU iGEM Team 2012)]] |
=Future Work= | =Future Work= | ||
Latest revision as of 18:55, 26 October 2012

The Orthogonal System of Regulating Translation: the AegiSafe™ O-Key
The synthesis of protein relies on the transcription-translation network. In transcription, mRNA is synthesized through complementary base pairing by RNA polymerase from the DNA template, and is followed by translation. Translation is the third stage of protein biosynthesis. In translation, mRNA produced by transcription is decoded by the ribosome to produce a specific amino acid chain, or polypeptide, that will later fold into an active protein. Therefore, we can say that translation is one of most important of activities in a cell.Due to its importance of protein synthesis, there are intricate and delicate regulation systems. To regulate transcription, cells alter the gene expression levels. This is called transcriptional regulation. Many means of transcriptional regulation, such as various mechanisms as specificity factors, activators, etc., are presented in a cell. There are also inducible and repressible systems, and transcription factor that can determine the initiation rate of transcription.
While a number of genetic tools exist for regulating transcription in cells, far fewer tools exist for translation. Of the tools available in bacteria, the most popular are riboregulators, both cis- and trans-activating, and orthogonal ribosomes (o-ribosomes). In terms of reprograming translation, o-ribosomes are especially powerful as they enable one to partially decouple translation from the native protein synthesis machinery. In particular, o-ribosomes can translate genes with altered Shine-Dalgarno (SD) sequences not recognized by host ribosomes. Therefore, o-ribosomes can be used to explore gene expression dynamics as they potentially provide a method for tuning translation rates. Furthermore, o-ribosomes may have application in synthetic biology as they introduce new functionality within cells.In this year's project, we focus on modifying the Sine-Dalgarno (SD) sequence of the mRNA and the anti-Shine-Dalgarno (ASD) sequence of the 16S rRNA of the ribosome to construct an orthogonal translation system called the AegiSafe™ O-Key. Aegis was originally the shield that was associated with Zeus and Athena, which offers protection. AegiSafe™ means our orthogonal system secure the environment from potential biosafety issue. O-Key stands for orthogonal system that includes orthogonal ribosome (o-ribosome, O-Key) and orthogonal RBS (o-RBS, O-Lock), which indicates that our system can not only shut down contamination, but also using the orthogonal system to selectively open our translation pathway like a key to a lcok. This is a truly amazing system for two independent translation systems co-existing harmoniously and working effectively in one single cell. In the following sections, we will elaborate on the AegiSafe™ O-Key about the orthogonal system and its application.
The Overview of Orthogonal Ribosome (O-Key)
The Canonical Ribosome
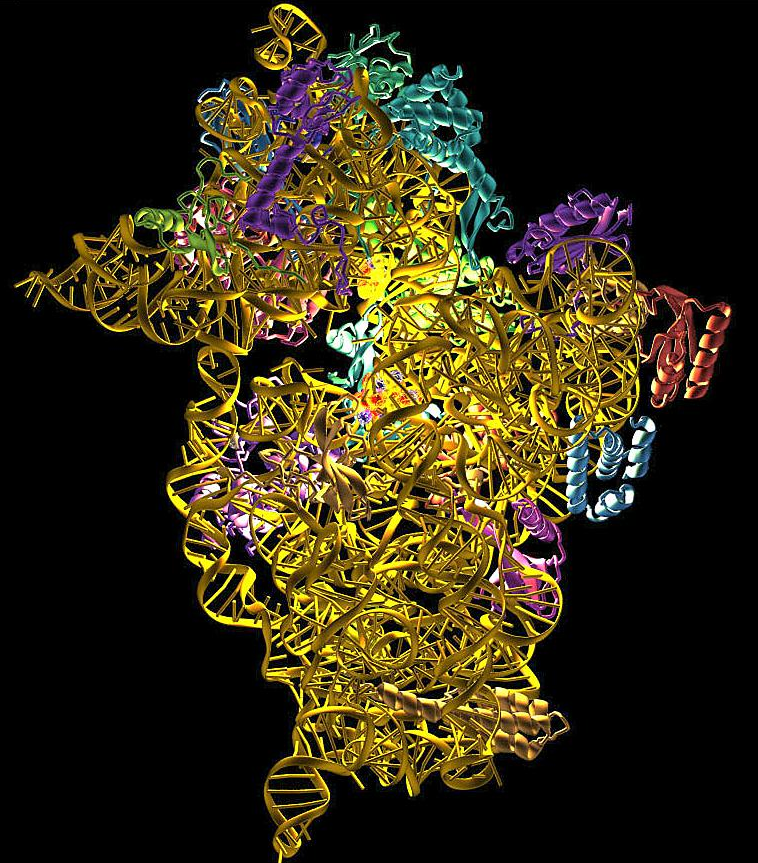
Ribosome is the protein factory of a cell, an organelle without membrane. Its sophisticated structure guarantees the accurate information flow from mRNA to protein. Ribosomes consist of two subunits that fit together and work as one to translate the mRNA into a polypeptide chain during protein synthesis.
Prokaryotes have 70S ribosomes, each consisting of a small (30S) and a large (50S) subunit. The smaller subunit binds to the mRNA pattern, while the larger subunit binds to the tRNA and the amino acids. Their small subunit has a 16S RNA subunit (consisting of 1540 nucleotides) bound to 21 proteins. The large subunit is composed of a 5S RNA subunit (120 nucleotides), a 23S RNA subunit (2900 nucleotides) and 31 proteins. Crystallographic work has shown that there are no ribosomal proteins close to the reaction site for polypeptide synthesis. This suggests that the protein components of ribosomes act as a scaffold that may enhance the ability of rRNA to synthesize protein rather than directly participating in catalysis, while the rRNA reacts with the mRNA and catalyze the synthesis.
Multiple sequences of 16S rRNA can exist within a single bacterium.
It has several functions:
- Like the large (23S) ribosomal RNA, it has a structural role, acting as a scaffold defining the positions of the rebosomal proteins.
- The 3' end contains the anti-Shine-Dalgarno sequence, which binds upstream to the AUG start codon on the mRNA. The 3'-end of 16S RNA binds to the proteins S1 and S21 known to be involved in initiation of protein synthesis
- Interacts with 23S, aiding in the binding of the two ribosomal subunits (50S+30S)
- Stabilizes correct codon-anticodon pairing in the A site, via a hydrogen bond formation between the N1 atom of Adenine (see image of Purine chemical structure) residues 1492 and 1493 and the 2'OH group of the mRNA backbone.
In bacterial cells, ribosomes are synthesized in the cytoplasm through the transcription of multiple ribosome gene operons.
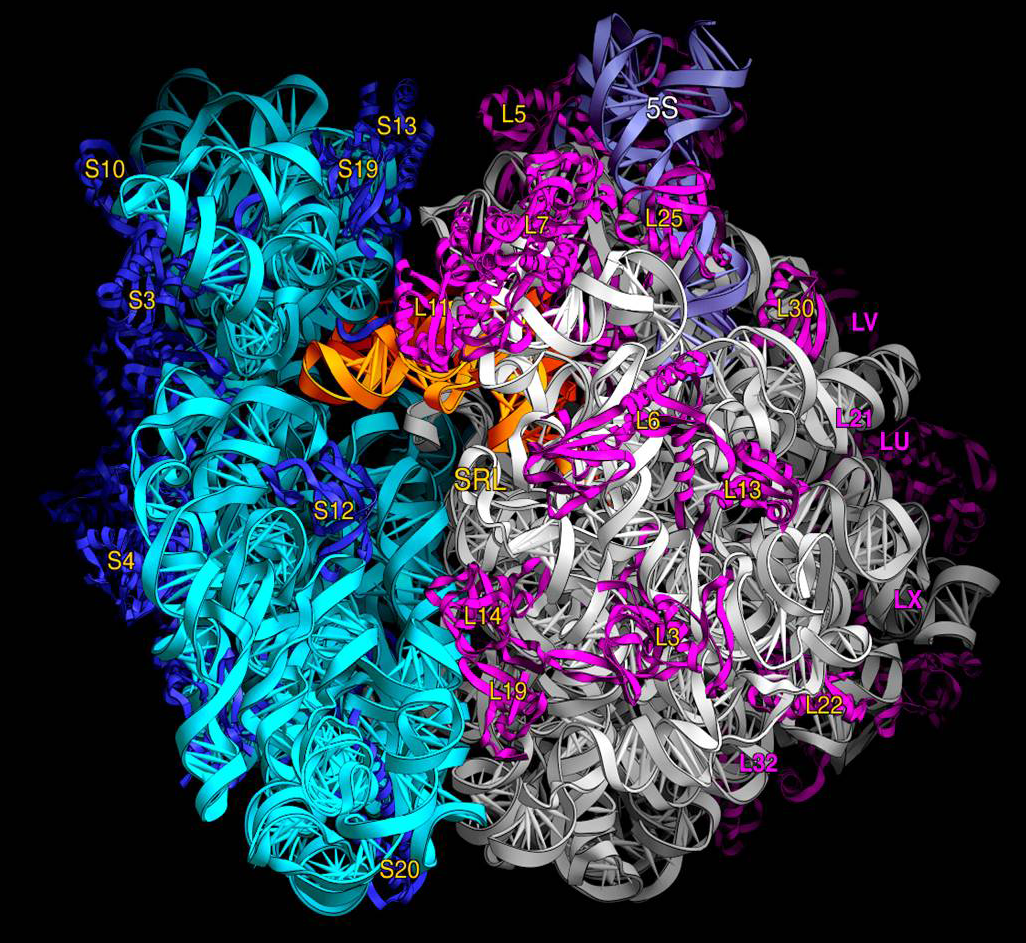
The rRNAs that make up the ribosome are directly translated from the rRNA operon rrnB. The three E. coli rRNA molecules – 23S, 16S, and 5S – are derived from a single 30S rRNA precursor transcript that also includes several tRNAs.
The Orthogonal Ribosome
The previous section described the structure of a canonical ribosome. As for the orthogonal ribosome (O-Key), the primary structure remains the same. The only difference lies in the ASD sequence close the 3' of the 16S rRNA. This altered sequence will not recognize the canonical SD sequence, thus pausing translation, just like a key cannot open a mismatched lock. The O-Key can only pair with its corresponding RBS, i.e. O-Lock. The O-Key (orthogonal) system serves as a powerful tool in regulating inner cellular metabolism. We will elaborate on its application in the following chapter.
Orthogonality Verification Experiment
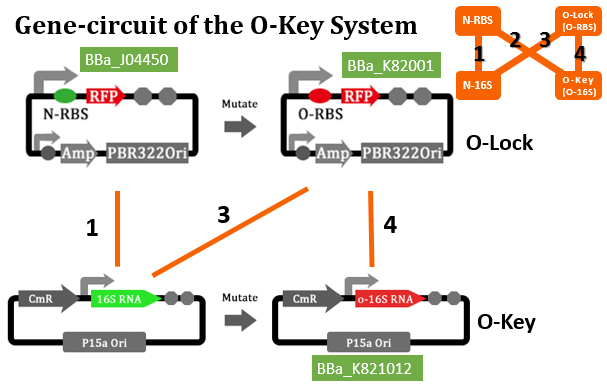
RFP RBS Mutation
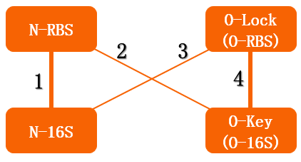
The translation process relies on the positioning function. The function works by the base pairing between the ASD sequence of 30s and the upstream SD sequence of mRNA. If the mutation happens to the SD sequence, its binding Gibson free energy will increase. And the binding will fail when the Gibson free energy decreases to a certain extent. This is equivalent to the installation of a lock on the subsequent gene, i.e. O-Lock. The lock only opens when matched O-Key appears. This is a key-lock system called O-Key system.
From this figure, we can see that in the four conditions, only 1 and 4 can work while the path 2 and 3 can't.
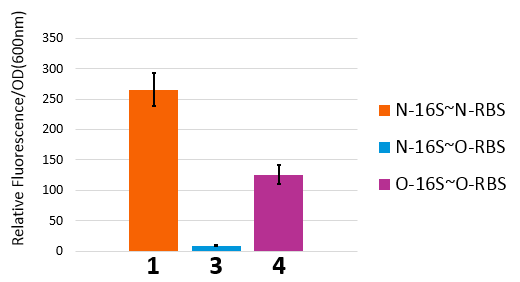
In our real experiment, we regulated and controlled the RFP gene to verify our O-Key system. For the survival of cells must need the 16S RNA, so we can just apply the path 1, 3, 4 to the real experiment.
| Group | RFP-Plasmid | O-16S-Plasmid | Result |
|---|---|---|---|
| 1 | N-RBS | N/A | Expression |
| 3 | O-RBS | Pbad+O-16S(No induced) | N/A |
| 3' | O-RBS | N/A | N/A |
| 4 | O-RBS | Pbad+O-16S(No induced) | Expression |
Ampr RBS Mutation
We used the orthogonal principle described in the previous section to mutate the Ampr RBS, and constructed an orthogonal transcription-translation system that is independent of the existing system.
The SD sequence of Ampr RBS is GAGAAA, and we rationally designed the orthogonal sequence to be GTTCCG. We further mutated the Ampr RBS of the RFP operon, and transformed plasmid into E coli.
Here, we chose two distinct competent cell: the normal competent cells; the orthogonal competent cells containing o-16S rRNA, and plate the cells on Amp LB plates. As we can predict, the normal competent cells would not be able to express the Amp resistance protein, thus cannot survive. On the contrary, the orthogonal cells can resist Amp in the plate and express the RFP.
Process
- Using PCR to mutate 16sRNA on the P15a plasmid to get the mutated P15a with O-16sRNA.
- Changing RBS of Ampr gene on PBR322 to O-RBS.
- Cotransfomating the above two mutated plasmids into cells, plating Amp LB plates, cultivating for 12-24hours.
Both of the plasmids used above contain the RFP gene to display the results and the Pbad promoter induced by Ala; only if Ampr gene could be expressed, cell will reproduce, RFP gene be expressed and the colony turned red.
Results
| Group | PSB1A3-RFP | O-16S | Result |
|---|---|---|---|
| 1(left) | O-RBS+AmpR | N/A | No colony |
| 2(right) | O-RBS+AmpR | O-16S | Red colony |
From the two flats, we could easily find that the left plant plated without Ala,which means cells only containing normal ribosome. There is no colony on the plate. The reason is that without no orthogonal ribosome, Ampr gene could not be transcripted and translated,then cells could not survive on LB flat with Amp. On the other hand, the right flat, added Ala, containing orthogonal ribosome, have red colonies. Existing orthogonal ribosome means that cells could transcript and translate the Amp r gene, survive in Amp situation, the RFP could also be expressed with the help of normal ribosome.
In this part, we have:
- Designed & constructed the O-Key System
- Predicted the orthogonality using modeling
- Characterized the orthogonality in the wet lab
- Installed the O-Lock in front of the Ampr resistance gene
Modeling
Gibson Free Energy in the O-Key System
The strength of the interaction between SD and ASD sequence is thought to influence translational efficiency as mutations in either the SD or ASD sequence that weaken the interaction reduce the amount of protein made. The mechanism of protein expression is primarily determined by the delta Gibbs free energy of the combination of SD sequence on ribosome and the ASD sequence on RBS of the mRNA. Therefore, it is necessary to investigate the property of ΔG.
ΔGtot=ΔGmRNA:rRNA+ΔGstart+ΔGspacing-ΔGstandby-ΔGmRNA
ΔGmRNA:rRNA is the energy released when the last nine nucleotides(nt) of the E. coli 16S rRNA (3′-AUUCCUCCA-5′) hybridizes and co-folds to the mRNA sub-sequence (ΔGmRNA:rRNA<0). Intramolecular folding within the mRNA is allowed. All possible hybridizations between the mRNA and 16S rRNA are considered to find the highest affinity 16S rRNA binding site. The binding site minimizes the sum of the hybridization free energy ΔGmRNA:rRNA and the penalty for nonoptimal spacing, ΔGspacing. Thus, the algorithm can identify the 16S rRNA binding site regardless of its similarity to the consensus Shine-Dalgarno sequence.
- ΔGstart is the energy released when the start codon hybridizes to the initiating tRNA anticodon loop (3'-UAC-5').
- ΔGspacing is the free energy penalty caused by a nonoptimal physical distance between the 16S rRNA binding site and the start codon (ΔGspacing>0). When this distance is increased or decreased from an optimum of 5 nt (or ~17A), the 30S complex becomes distorted, resulting in a decreased translation initiation rate.
- ΔGmRNA is the work required to unfold the mRNA sub-sequence when it folds to its most stable secondary structure, called the minimum free energy structure (ΔGmRNA<0).
- ΔGstandby is the work required to unfold any secondary structures sequestering the standby site (ΔGstandby<0) after the 30S complex assembly. We define the standby site as the four nucleotides upstream of the 16S rRNA binding site, which is its location in a previously studied mRNA.
To calculate ΔGmRNA:rRNA, ΔGstart, ΔGmRNA and ΔGstandby, we use the NUPACK suite of algorithms with the Mfold 3.0 RNA energy parameters. These free energy calculations do not have any additional fitting or training parameters and explicitly depend on the mRNA sequence. In addition, the free energy terms are not orthogonal; changing a single nucleotide can potentially affect multiple energy terms. The relationship between the spacing and the ΔGspacing was empirically determined by measuring the protein expression level driven by synthetic RBSs of varying spacing and fitting a quantitative model to this data.
In order to design the orthogonality and to predict the protein expression level, we have established the protein expression model which has strong relationship with the wet lab. The model is based on detailed study of the effects of changing Gibbs free energy on the translation process.
In the process of translation, the initiation rate is the rate determine step. The protein expression amount is consequently direct proportional to the initial translation speed. Furthermore, the initial translation speed is strongly related to the delta Gibbs free energy, which can be calculated out through results of many newly published papers and various commercialized software such as the Vienna RNA Package, RBS-Calculator, and Nucleic Acid Package. In our model, we calculated the delta Gibbs energy. Ideal results can be obtained from the model. The orthogonality predicted by our model before the wet lab is similar to our prediction. When combined with some wet lab result, we can finally predict the spectroscope curve of RFP and GFP under three designed experimental state. For more details, see our Model Part.
We also want to add two O-Key system in our modeling and predict the result
Future Work
Previous experiments and modeling has described that the orthogonality of our expression system is very tight, but we are still not satisfied with the current state, then we design and do the following experiment. The following experiment includes:
- We are using λRed technology to integrate the O-16s biobrick into the genome for constructing the orthogonal cells, so as to solve the revertants as well as stability problems caused by plasmid instability.
- We are trying to build other groups of different orthogonal (O-16s-RBS) system, so as to achieve the different orthogonality switch effect, and attempt to make them corporate.
- We also consult with some biological gene enterprises to make our results put into production in the life full of modern technology, such as genetic pollution control, gene encryption and metabolic regulation, etc.
References
- http://en.wikipedia.org/wiki/16S_ribosomal_RNA
- Lon M. Chubiz and Christopher V. Rao(2008). Computational design of orthogonal ribosomes, vol.36 no.12
- Howard M Salis, Ethan A Mirsky & Christopher A Voigt(2009).Automated design of synthetic ribosome binding sites to control protein expression, doi:10.1038/nbt.1568
- Wenlin An and Jason W. Chinl(2009).Synthesis of orthogonal transcription-translation networks,vol.106 no.21
- George H.McArthur IV and Stephen S. Fong, Toward Engineering Synthetic Microbial Metabolism
- Kevin Clancy and Christopher A Voigt, Programming cells: towards an automated ‘Genetic Compiler’
- WANG Haisheng, ZHANG Xiaoxia, et al., Recent research progress of bacterial violacein
- Carotenoid Crystal Formation in Arabidopsis and Carrot Roots Caused by Increased Phytoene Synthase Protein Levels
- Guoyin Kai, Pan Liao, Tong Zhang, Wei Zhou, Jing Wang, Hui Xu, Yuanyun Liu, and Lin Zhang, Characterization, Expression Profiling, and Functional Identification of a Gene Encoding Geranylgeranyl Diphosphate Synthase from Salvia miltiorrhiza.
- crtB Masaki Fujisawa,Mio Watanabe,Song-Kang Choi,Maki Teramoto,Kanji Ohyama,and Norihiko Misawa, Enrichment of Carotenoids in Flaxseed (Linum usitatissimum) by Metabolic Engineering with Introduction of Bacterial Phytoene Synthase Gene
- William R. Farmer and James C. Liao, Improving lycopene production in Escherichia coli by engineering metabolic control.
 "
"