Team:Michigan/Project
From 2012.igem.org
(Difference between revisions)
| (22 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
<div class="body"> | <div class="body"> | ||
<div id="basic"> | <div id="basic"> | ||
| - | + | <h3>Abstract</h3> | |
| - | + | Recombinases can be used to create responsive genetic elements in biological systems. Further, one can build complex control circuits using combinations of invertible DNA sequences. We utilized the unidirectional recombinase HbiF to augment an existing recombinase system in <i>Escherichia coli</i> that relied on the unidirectional recombinase FimE. A burst of low level expression of one recombinase is expected to invert the promoter flanked by the recombinase binding sites IRR and IRL, triggering a switch from strong expression of one to another set of proteins made downstream. Expression of the second recombinase is expected to revert the promoter to its original orientation, triggering the original set of protein expression. The inversion will be sustained across cell divisions with little leaky protein expression and negligible performance degradation after repeated inversions. This is a heritable, binary memory system and can be used as a component in more complex systems. | |
<br><br> | <br><br> | ||
| Line 14: | Line 14: | ||
<h2>fimS or synthetic “flip” sequence </h2> | <h2>fimS or synthetic “flip” sequence </h2> | ||
| - | + | The process of fimbriation in <i>Escherichia coli</i> is regulated by the enzyme-catalyzed inversion of a segment of DNA, often labeled <i>fimS</i>, by a series of tyrosine recombinases of which FimE and FimB are the most known. In the absence of ATP or external prosthetic groups, the fim recombinases introduce strand breaks in repeating regions flanking the fimS regulatory sequence and physically reverse the sequence, activating or repressing the expression of fimbriation proteins based on the orientation of a contained promoter (McClain et al., 1991). | |
| - | The process of fimbriation in Escherichia coli is regulated by the enzyme-catalyzed inversion of a segment of DNA, often labeled fimS, by a series of tyrosine recombinases of which FimE and FimB are the most known. In the absence of ATP or external prosthetic groups, the fim recombinases introduce strand breaks in repeating regions flanking the fimS regulatory sequence and physically reverse the sequence, activating or repressing the expression of fimbriation proteins based on the orientation of a contained promoter. | + | <br><br> |
| - | < | + | |
| - | + | Inversion catalyzed by FimE was shown to be primarily unidirectional, inverting <i>fimS</i> from the “on” to “off” orientation in regards to fimbriae production. This stands in contrast to FimB, which is capable of inverting <i>fimS</i> bidirectionally, forming an equilibrium between “off” and “on” states.# An inducible genetic circuit was engineered by Ham et al., 2006 utilizing this property to produce tightly controllable, strong protein expression with relatively low basal expression and a unique mode of activation, requiring only “pulses” of inducer rather than extended exposure. Team Michigan sought to improve on this idea by utilizing two unidirectional recombinases -- FimE and HbiF. | |
| - | Inversion catalyzed by FimE was shown to be primarily unidirectional, inverting fimS from the “on” to “off” orientation | + | |
| - | + | ||
| - | < | + | <br><br> |
| - | + | ||
| - | + | ||
| + | |||
| + | |||
<h2> | <h2> | ||
| - | Orientation of IRL/IRR Components of fimS dictate enzyme specificity | + | Orientation of IRL/IRR Components of <i>fimS</i> dictate enzyme specificity |
</h2> | </h2> | ||
| - | + | The fimS region is composed of two inverted repeats, frequently referred to as inverted repeat left (IRL) and inverted repeat right (IRR). The two flank a region of DNA referred to as the inversion region. Each of the inverted repeats are composed of three sections of DNA, an external half-site, an inverted repeat, and a internal half-site. The external half-sites and inverted repeats are stable and identifiers of the IRL and IRR, while the internal half-sites are mobile, switching between the IRL and IRR, dictating recombinase specificity and the orientation of the invertible region. The internal half-sites are inverted along with the inversion region when the region is acted upon by one of the recombinases (Fig 1). (McCuster et al., 2008) | |
| - | The fimS region is composed of two inverted repeats, frequently referred to as inverted repeat left (IRL) and inverted repeat right (IRR) | + | |
</html> | </html> | ||
| - | [[File: | + | [[File:image00.png|400px|thumb|left|Fig 1. Orientation of fimS inverted repeats. In the “off “state the IRL contains internal half-site A (orange), while the IRR contains internal half-site B (green). When inverted to the “on” state the IRL contains internal half-site B (green), while the IRR contains internal half-site A (orange). <br><br> |
| + | (1) McCuster et al., Molecular Microbiology (2008) 67(1), 171–187]] | ||
<html> | <html> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| + | <br> | ||
<h2>FimE</h2> | <h2>FimE</h2> | ||
| - | + | ||
| - | The FimE enzyme was cloned using the Caltech-made part K137007. A well characterized tyrosine recombinase, the enzyme catalyzes the inversion of a 200-300bp DNA region such that a 3’-facing sequence on the template strand of the region becomes a 3’-facing sequence on the coding strand. This functionality occurs in the lack of general cofactors or ATP, as demonstrated by Ham et al, | + | The FimE enzyme was cloned using the Caltech-made part K137007. A well characterized tyrosine recombinase, the enzyme catalyzes the inversion of a 200-300bp DNA region such that a 3’-facing sequence on the template strand of the region becomes a 3’-facing sequence on the coding strand. This functionality occurs in the lack of general cofactors or ATP, as demonstrated by Ham et al., 2006. |
| - | + | ||
| - | + | <br><br> | |
FimE is dependent on properly oriented inverted repeats (IRs) flanking a region of interest for the reaction to occur. Furthermore, it may only invert a fim sequence in the “on” orientation to the “off” orientation. | FimE is dependent on properly oriented inverted repeats (IRs) flanking a region of interest for the reaction to occur. Furthermore, it may only invert a fim sequence in the “on” orientation to the “off” orientation. | ||
| - | |||
| + | |||
| + | |||
| + | <BR><BR> | ||
<h2>HbiF</h2> | <h2>HbiF</h2> | ||
| - | + | ||
| - | HbiF, a tyrosine recombinase bearing high sequence homology to FimE and other fim recombinases, was discovered in 2006 by Xie et | + | HbiF, a tyrosine recombinase bearing high sequence homology to FimE and other fim recombinases, was discovered in 2006 by Xie et al and also catalyses inversion of the <i>fimS</i> region. Our concept harnesses HbiF to convert the FimE-activatable expression system described by Ham et al., 2006 into a bidirectional switch capable of binary states dependent on the presence and expression of either FimE or HbiF recombinases. Team Michigan’s submission of HbiF, BBa_K880000, possesses a protein fusion standard RFC25 suffix to facilitate circuit regulation or protein quantification. |
| - | < | + | <br> |
| - | + | In contrast to FimE, HbiF catalyzes the rotation of <i>fimS</i> from the “off” to “on” position in vivo. | |
| - | + | <br><br> | |
| - | < | + | |
| - | < | + | <h3>Rationale</h3> |
| - | + | <h2>A Tightly-controlled, stable switch</h2> | |
| - | </ | + | |
| - | < | + | A system utilizing both FimE and HbiF has several advantages beyond available multi-state biological circuits. As the binding site for HbiF becomes a FimE binding site after inversion, the system contains a remarkable number of reusable components. One recombinase can flip the <i>fimS</i> sequence only after the other recombinase has acted. This limitation allows the host to remember its previous state as the inversion region is flipped; it will not respond to duplicates of the same signal and instead allows for all target sequences to accumulate in one state. The switching sequence can also theoretically flip back and forth without generating new component and without being depleted. |
| - | + | <h2>Cheaper Biomanufacturing </h2> | |
| - | + | ||
| - | </ | + | |
| - | + | A system utilizing both FimE and HbiF as actuators would allow for the creation of a genetic circuit that works in a manner analogous to a push-button switch. Current expression systems often rely on inducible promoters systems, such as pBAD and the lac promoter, requiring a constant administration of inducers making them more analogous to a switch that is actively being held in the “ON” position. The constant administration of inducers, such as arabinose and lactose, are a component of the high operating costs of biomanufacturing. FimE and HbiF in concert with separate inducible promoters could be used as a push-button switch where a short exposure to one of the inducers would trigger recombinase expression resulting in inversion of the switch and a new set of protein expression. This “push-button switch” would eliminate the need for constant administration of inducers, resulting in a reduction in operating costs for biomanufacturing. This “push-button switch” would also improve upon the recombinase system described in Ham et al., 2006 as the switch could be readily inverted in either direction, while the Ham et al. system would require re-seeding with uninverted cells everytime protein expression must be reset to the initial condition. | |
| - | + | ||
| - | < | + | <br><br> |
| + | |||
| + | |||
| + | <h2>References</h2> | ||
| + | Ham et al., Biotechnol. Bioeng. (2006), Vol. 94(1) | ||
| + | <br><br> | ||
| + | Ham et al., PLoS ONE (2008), 3(7) | ||
| + | |||
| + | <br><br> | ||
| + | McClain et al., J. Bacteriol. (1991) 173(17) | ||
| + | <br><br> | ||
| + | McCuster et al., Molecular Microbiology (2008), 67(1) | ||
| + | <br><br> | ||
| + | Xie et al. Infection and Immunity (2006) 74(7) | ||
| + | |||
| + | <br><br> | ||
</div> | </div> | ||
| + | </div> | ||
| - | |||
| - | |||
| - | |||
| - | |||
</div> | </div> | ||
Latest revision as of 03:10, 4 October 2012
Abstract
Recombinases can be used to create responsive genetic elements in biological systems. Further, one can build complex control circuits using combinations of invertible DNA sequences. We utilized the unidirectional recombinase HbiF to augment an existing recombinase system in Escherichia coli that relied on the unidirectional recombinase FimE. A burst of low level expression of one recombinase is expected to invert the promoter flanked by the recombinase binding sites IRR and IRL, triggering a switch from strong expression of one to another set of proteins made downstream. Expression of the second recombinase is expected to revert the promoter to its original orientation, triggering the original set of protein expression. The inversion will be sustained across cell divisions with little leaky protein expression and negligible performance degradation after repeated inversions. This is a heritable, binary memory system and can be used as a component in more complex systems.Background
fimS or synthetic “flip” sequence
The process of fimbriation in Escherichia coli is regulated by the enzyme-catalyzed inversion of a segment of DNA, often labeled fimS, by a series of tyrosine recombinases of which FimE and FimB are the most known. In the absence of ATP or external prosthetic groups, the fim recombinases introduce strand breaks in repeating regions flanking the fimS regulatory sequence and physically reverse the sequence, activating or repressing the expression of fimbriation proteins based on the orientation of a contained promoter (McClain et al., 1991).Inversion catalyzed by FimE was shown to be primarily unidirectional, inverting fimS from the “on” to “off” orientation in regards to fimbriae production. This stands in contrast to FimB, which is capable of inverting fimS bidirectionally, forming an equilibrium between “off” and “on” states.# An inducible genetic circuit was engineered by Ham et al., 2006 utilizing this property to produce tightly controllable, strong protein expression with relatively low basal expression and a unique mode of activation, requiring only “pulses” of inducer rather than extended exposure. Team Michigan sought to improve on this idea by utilizing two unidirectional recombinases -- FimE and HbiF.
Orientation of IRL/IRR Components of fimS dictate enzyme specificity
The fimS region is composed of two inverted repeats, frequently referred to as inverted repeat left (IRL) and inverted repeat right (IRR). The two flank a region of DNA referred to as the inversion region. Each of the inverted repeats are composed of three sections of DNA, an external half-site, an inverted repeat, and a internal half-site. The external half-sites and inverted repeats are stable and identifiers of the IRL and IRR, while the internal half-sites are mobile, switching between the IRL and IRR, dictating recombinase specificity and the orientation of the invertible region. The internal half-sites are inverted along with the inversion region when the region is acted upon by one of the recombinases (Fig 1). (McCuster et al., 2008)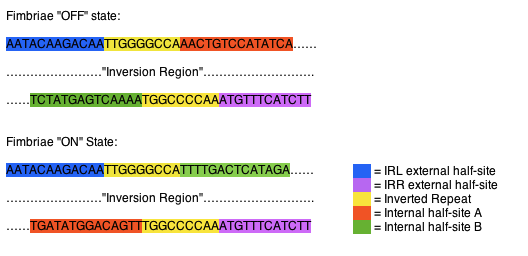
Fig 1. Orientation of fimS inverted repeats. In the “off “state the IRL contains internal half-site A (orange), while the IRR contains internal half-site B (green). When inverted to the “on” state the IRL contains internal half-site B (green), while the IRR contains internal half-site A (orange).
(1) McCuster et al., Molecular Microbiology (2008) 67(1), 171–187
(1) McCuster et al., Molecular Microbiology (2008) 67(1), 171–187
FimE
The FimE enzyme was cloned using the Caltech-made part K137007. A well characterized tyrosine recombinase, the enzyme catalyzes the inversion of a 200-300bp DNA region such that a 3’-facing sequence on the template strand of the region becomes a 3’-facing sequence on the coding strand. This functionality occurs in the lack of general cofactors or ATP, as demonstrated by Ham et al., 2006.FimE is dependent on properly oriented inverted repeats (IRs) flanking a region of interest for the reaction to occur. Furthermore, it may only invert a fim sequence in the “on” orientation to the “off” orientation.
HbiF
HbiF, a tyrosine recombinase bearing high sequence homology to FimE and other fim recombinases, was discovered in 2006 by Xie et al and also catalyses inversion of the fimS region. Our concept harnesses HbiF to convert the FimE-activatable expression system described by Ham et al., 2006 into a bidirectional switch capable of binary states dependent on the presence and expression of either FimE or HbiF recombinases. Team Michigan’s submission of HbiF, BBa_K880000, possesses a protein fusion standard RFC25 suffix to facilitate circuit regulation or protein quantification.In contrast to FimE, HbiF catalyzes the rotation of fimS from the “off” to “on” position in vivo.
Rationale
A Tightly-controlled, stable switch
A system utilizing both FimE and HbiF has several advantages beyond available multi-state biological circuits. As the binding site for HbiF becomes a FimE binding site after inversion, the system contains a remarkable number of reusable components. One recombinase can flip the fimS sequence only after the other recombinase has acted. This limitation allows the host to remember its previous state as the inversion region is flipped; it will not respond to duplicates of the same signal and instead allows for all target sequences to accumulate in one state. The switching sequence can also theoretically flip back and forth without generating new component and without being depleted.Cheaper Biomanufacturing
A system utilizing both FimE and HbiF as actuators would allow for the creation of a genetic circuit that works in a manner analogous to a push-button switch. Current expression systems often rely on inducible promoters systems, such as pBAD and the lac promoter, requiring a constant administration of inducers making them more analogous to a switch that is actively being held in the “ON” position. The constant administration of inducers, such as arabinose and lactose, are a component of the high operating costs of biomanufacturing. FimE and HbiF in concert with separate inducible promoters could be used as a push-button switch where a short exposure to one of the inducers would trigger recombinase expression resulting in inversion of the switch and a new set of protein expression. This “push-button switch” would eliminate the need for constant administration of inducers, resulting in a reduction in operating costs for biomanufacturing. This “push-button switch” would also improve upon the recombinase system described in Ham et al., 2006 as the switch could be readily inverted in either direction, while the Ham et al. system would require re-seeding with uninverted cells everytime protein expression must be reset to the initial condition.References
Ham et al., Biotechnol. Bioeng. (2006), Vol. 94(1)Ham et al., PLoS ONE (2008), 3(7)
McClain et al., J. Bacteriol. (1991) 173(17)
McCuster et al., Molecular Microbiology (2008), 67(1)
Xie et al. Infection and Immunity (2006) 74(7)
 "
"

