Team:LMU-Munich/Bacillus BioBricks
From 2012.igem.org
| Line 206: | Line 206: | ||
| - | The concentrations of the antibiotics and | + | The concentrations of the antibiotics for selection of ''B.subtilis'' transformation and tests for plasmid insertion can be found in our [https://2012.igem.org/Team:LMU-Munich/Lab_Notebook/Protocols Protocol] section. |
| - | See [[Team:LMU-Munich/Bacillus_BioBricks/vector_use| here]] to find out how to use ''B. subtilis'' vectors. In this [[Team:LMU-Munich/Bacillus_BioBricks/integration|overview]], the mechanism of | + | See [[Team:LMU-Munich/Bacillus_BioBricks/vector_use| here]] to find out how to use ''B. subtilis'' vectors. In this [[Team:LMU-Munich/Bacillus_BioBricks/integration|overview]], the integration mechanism of ''B. subtilis'' vectors is described. |
The design and special use of our Sporovector can be found [[Team:LMU-Munich/Bacillus_BioBricks/Sporovector |here]]. | The design and special use of our Sporovector can be found [[Team:LMU-Munich/Bacillus_BioBricks/Sporovector |here]]. | ||
| - | For results, check our | + | For results, check our Data page: [[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Data/Vectors]] |
| Line 315: | Line 315: | ||
{| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
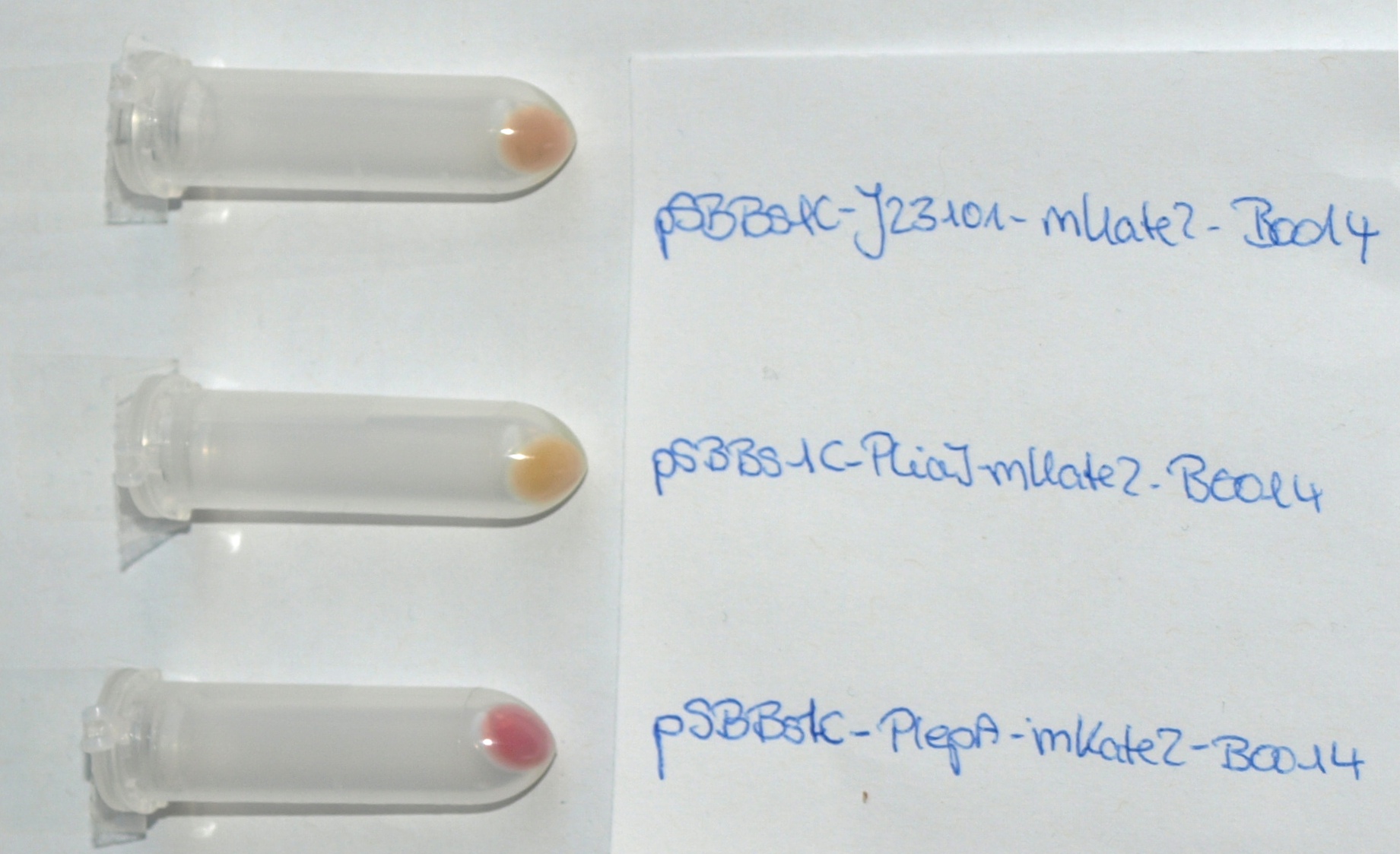
| - | <font color="#000000"; size="2"><p align="justify">'''''mKate2'' fused to the terminator B0014 under the control of the Anderson promoter J23101 (up), P<sub>''liaI''</sub> (middle) and P<sub>''lepA''</sub> (down) in pSB<sub>''Bs''</sub>1C.''' Pellets are ''Escherichia coli'' cells which contain the plasmid with the | + | <font color="#000000"; size="2"><p align="justify">'''''mKate2'' fused to the terminator B0014 under the control of the Anderson promoter J23101 (up), P<sub>''liaI''</sub> (middle) and P<sub>''lepA''</sub> (down) in pSB<sub>''Bs''</sub>1C.''' Pellets are ''Escherichia coli'' cells which contain the plasmid with the insert labelled to the right.</p></font> |
|} | |} | ||
|} | |} | ||
| Line 326: | Line 326: | ||
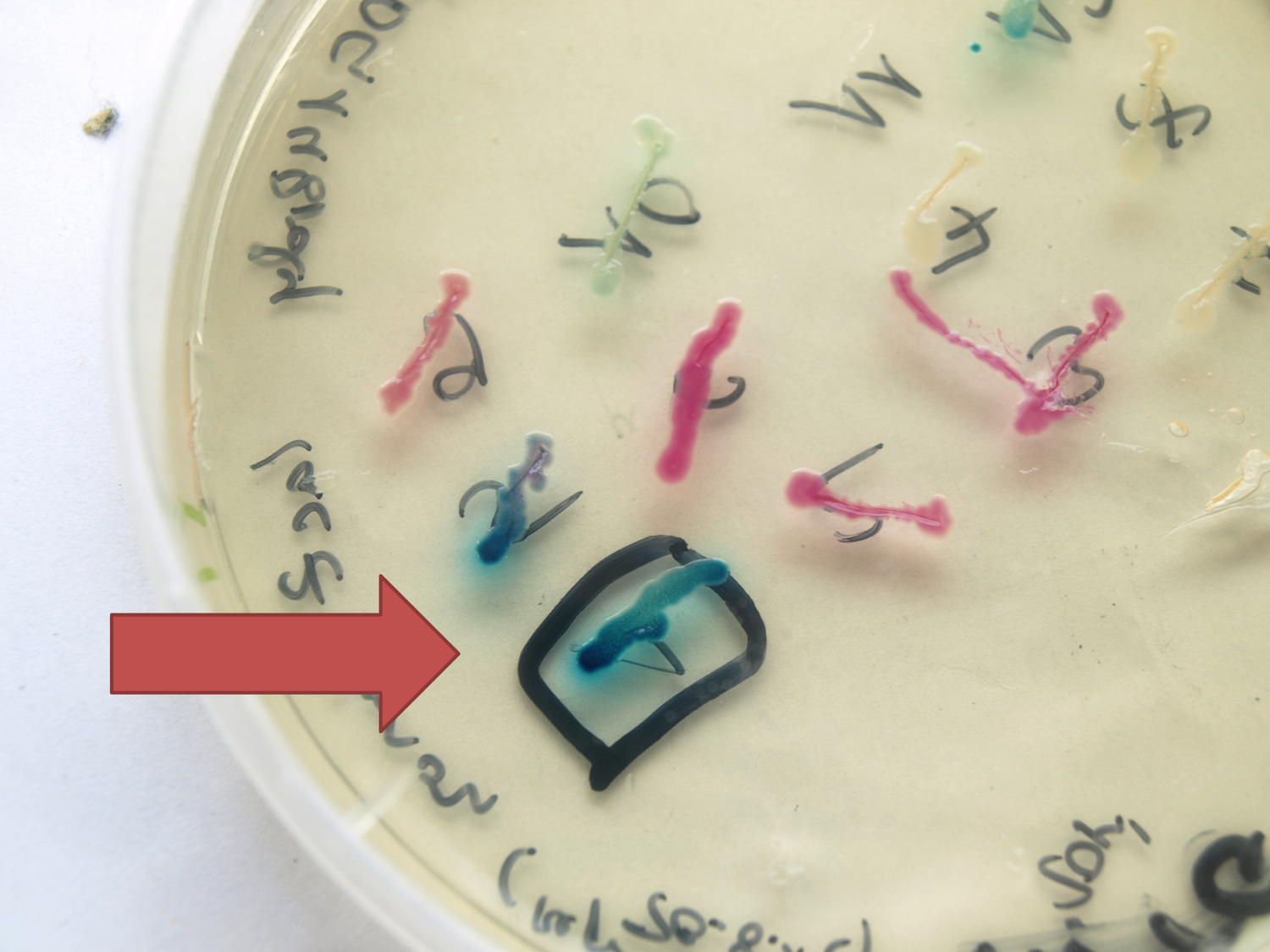
[[File:LacZ plate.png|<p align="justify">''lacZ'' fused to the terminator B0014 under the control of P<sub>''spac''</sub> in the expression vector pSB<sub>Bs</sub>0K-P<sub>''spac''</sub>. P<sub>''spac''</sub> induced with IPTG</p>|thumb|300px|left]] | [[File:LacZ plate.png|<p align="justify">''lacZ'' fused to the terminator B0014 under the control of P<sub>''spac''</sub> in the expression vector pSB<sub>Bs</sub>0K-P<sub>''spac''</sub>. P<sub>''spac''</sub> induced with IPTG</p>|thumb|300px|left]] | ||
| - | <p align="justify">This ''lacZ'' gene is derived from the ''Bacillus'' reporter vector pAC6. It is constructed in the Freiburg Standard (Assembly 25) for in-frame fusion proteins. It also includes a ribosome binding site optimized for ''Bacillus subtilis'' translation. This ''lacZ'' BioBrick was tested in the expression vector pSB<sub>Bs</sub>0K-P<sub>''spac''</sub>. This construct | + | <p align="justify">This ''lacZ'' gene is derived from the ''Bacillus'' reporter vector pAC6. It is constructed in the Freiburg Standard (Assembly 25) for in-frame fusion proteins. It also includes a ribosome binding site optimized for ''Bacillus subtilis'' translation. This ''lacZ'' BioBrick was tested in the expression vector pSB<sub>Bs</sub>0K-P<sub>''spac''</sub>. This construct gives output, so the reporter gene is functional. See [https://2012.igem.org/Team:LMU-Munich/Data/Vectors Data] in the vector evaluation section of pSB<sub>Bs</sub>0K-P<sub>''spac''</sub>.</p> |
| Line 351: | Line 351: | ||
<p align="justify"> | <p align="justify"> | ||
| - | All our tags have been synthesized by gene art. They are designed in Freiburg standard with an included optimized ribosome binding site. We have not yet tested our tags. </p> | + | All our tags have been synthesized by [http://de-de.invitrogen.com/site/de/de/home/Products-and-Services/Applications/Cloning/gene-synthesis.html?CID=fl-genesynthesis gene art]. They are designed in Freiburg standard with an included optimized ribosome binding site. We have not yet tested our tags. </p> |
prefix: GAATTCCGCGGCCGCTTCTAGATAAGGAGGAACTACTATGGCCGGC | prefix: GAATTCCGCGGCCGCTTCTAGATAAGGAGGAACTACTATGGCCGGC | ||
| Line 373: | Line 373: | ||
<p align="justify"> | <p align="justify"> | ||
| - | The cMyc-tag is a tag derived from the cMyc gene product. Antibodies were derived from the immunisation with synthetic peptides from the cMyc sequence ([http://mcb.asm.org/content/5/12/3610.short Mol. Cell. Biol. 5 | + | The cMyc-tag is a tag derived from the cMyc gene product. Antibodies were derived from the immunisation with synthetic peptides from the cMyc sequence ([http://mcb.asm.org/content/5/12/3610.short Mol. Cell. Biol. 5]). The aminoacid sequence is <b>EQKLISEEDL</b>.</p> |
| Line 379: | Line 379: | ||
<p align="justify"> | <p align="justify"> | ||
| - | The His-tag is a metal chelating peptide ([http://www.nature.com/nbt/journal/v6/n11/full/nbt1188-1321.html Hochuli, E.; Bannwarth, W.; Döbeli, H.; Gentz, R.; Stüber, D. (1988)]) consisting of at least 6 histidins. It can therefore be used for protein purification by metal 2+ (mostly nickel or cobalt) | + | The His-tag is a metal chelating peptide ([http://www.nature.com/nbt/journal/v6/n11/full/nbt1188-1321.html Hochuli, E.; Bannwarth, W.; Döbeli, H.; Gentz, R.; Stüber, D. (1988)]) consisting of at least 6 histidins. It can therefore be used for protein purification by metal<sup>2+</sup>-ion containing columns (mostly nickel or cobalt). There are also antibodies against this tag, or nickel/cobalt containing fluorescent probes can be used for detection. Also a immobilization is possible in nickel/cobalt coated plastikware. |
The aminoacid sequence is:<b>HHHHHHHHHH</b></p> | The aminoacid sequence is:<b>HHHHHHHHHH</b></p> | ||
Revision as of 15:51, 25 September 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]


B4 - 22 core parts for Bacillus subtilis
Since Bacillus subtilis is not a common organism used in iGEM, please check out our Introduction.
We created a toolbox of Bacillus BioBricks to contribute to the registry to make it accessible to many more future iGEM-Teams!
This Bacillus BioBrickBox (B4) contains the following Bacillus specific parts:
| Vectors | Promoters | Reporters | Affinity tags |

| 
| 
| |
|
pSBBs1C |
Anderson |
gfp |
Flag |
Bacillus Vectors 
We have generated a suit of BioBrick-compatible vectors: three empty ones with different antibiotic resistances and integration loci, two reporter and two expression vectors. Here is a list of all the vectors we cloned and used.
For the use of our vectors please see our Protocols page. A general introduction to Bacillus subtilis and its integrative vectors can be found here. All vectors have ampicillin as Escherichia coli resistance and RFP in the multiple cloning site as selection marker.
| Vector Name | Resistance | Insertion | Description | Vector origin | ||
|---|---|---|---|---|---|---|
| BioBrick | Eco | Bsu | locus | Name | Reference | |
| pSBBs1C [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823023 (BBa_K823023)] | Amp | Cm | amyE | empty | pDG1662 | [http://www.ncbi.nlm.nih.gov/pubmed/8973347 Guérout-Fleury] |
| pSBBs4S [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823022 (BBa_K823022)] | Amp | Spec | thrC | empty | pDG1731 | [http://www.ncbi.nlm.nih.gov/pubmed/8973347 Guérout-Fleury] |
| pSBBs2E [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823027 (BBa_K823027)] | Amp | MLS | lacA | empty | pAX01 | [http://www.ncbi.nlm.nih.gov/pubmed/11274134 Härtl] |
| pSBBs1C-lacZ [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823021 (BBa_K823021) ] | Amp | Cm | amyE | lacZ reporter | pAC6 | [http://www.ncbi.nlm.nih.gov/pubmed/11902727 Stülke] |
| pSBBs3C-luxABCDE [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823025 (BBa_K823025)] | Amp | Cm | sacA | luxABCDE reporter | pAH328 | [http://www.ncbi.nlm.nih.gov/pubmed/20709900 Schmalisch] |
| pSBBs4S-PXyl [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823024 (BBa_K823024)] | Amp | Spec | thrC | Xylose-promoter | pXT | [http://www.ncbi.nlm.nih.gov/pubmed/11069659 Derré] |
| pSBBs0K-Pspac [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823026 (BBa_K823026)] | Amp | Kan | replicative | IPTG-promoter | pDG148 | [http://www.ncbi.nlm.nih.gov/pubmed/11728721 Joseph] |
| Sporovector [http://partsregistry.org/wiki/index.php?title=Part:BBa_K823054 (BBa_K823054)] | Amp | Spec | thrC | to create Sporobeads | pSBBs4S | Sporovector |
Here you can find the corresponding vector maps:

| 
|

| 
|

| 
|

| 
|
The number in the vector's name codes for the insertion locus and the following letter for the Bacillus subtilis resistance gene according to the following table:
| Number | Insertion locus | Letter | Resistance |
|---|---|---|---|
| 0 | replicative | C | Chloramphenicol (Cm) |
| 1 | amyE (amylase) | E | MLS (Erythromycin + Lincomycin) |
| 2 | lacA (β-galactosidase) | K | Kanamycin (Kan) |
| 3 | sacA (sucrase) | S | Spectinomycin (Spec) |
| 4 | thrC (threonine synthase) |
The concentrations of the antibiotics for selection of B.subtilis transformation and tests for plasmid insertion can be found in our Protocol section.
See here to find out how to use B. subtilis vectors. In this overview, the integration mechanism of B. subtilis vectors is described.
The design and special use of our Sporovector can be found here.
For results, check our Data page: ![]()
Bacillus Promoters 
To provide a set of promoters with different strength we characterized several promoters in Bacillus subtilis. They can be divided in three different groups: the constitutive promoters from the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] from the Partsregistry, the constitutive promoters PliaG, Pveg and PlepA from B. subtilis, and the inducible promoters PliaI and Pxyl-xylR from B. subtilis. For the characterization of the different promoters we used the lux operon ![]() where promoter activity leads to expression of the luciferase and to the formation of luminescence. For this promoter evaluation the reporter vector pSBBs3C-luxABCDE was used which was not fully in BioBrickStandard at this time because of one last forbidden restriction site. We also used the reporter gene lacZ
where promoter activity leads to expression of the luciferase and to the formation of luminescence. For this promoter evaluation the reporter vector pSBBs3C-luxABCDE was used which was not fully in BioBrickStandard at this time because of one last forbidden restriction site. We also used the reporter gene lacZ ![]() . Here, promoter activation results in expression of a β-galactosidase, whose activity can be measured by β-galactosidase assays. Therefore we used the reporter vector pSBBs1C-lacZ. See this page for an overview and background information of all evaluated promoters and see the Data page for more details.
. Here, promoter activation results in expression of a β-galactosidase, whose activity can be measured by β-galactosidase assays. Therefore we used the reporter vector pSBBs1C-lacZ. See this page for an overview and background information of all evaluated promoters and see the Data page for more details.
Overview of all evaluated promoters
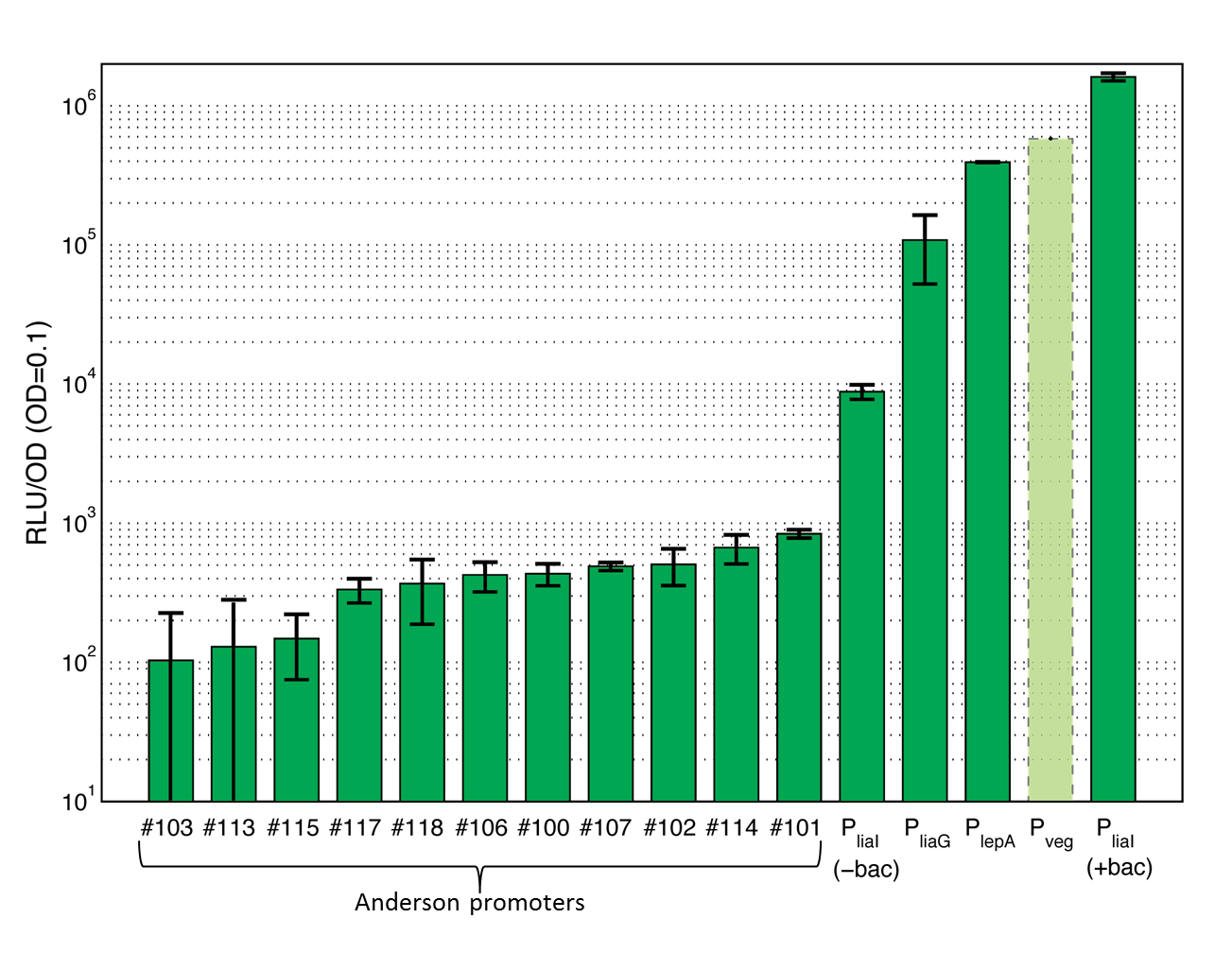
This section gives an overview of all evaluated promoters which cover a large range of activity. For more details and informations of the experiments see the Data page of the promoters. Note that Pveg was not evaluated with luminescence measurements and this bar is just projected from the results of the beta-galactosidase assay.
|
Anderson promoters
The first group of promoters evaluated are the promoters of the [http://partsregistry.org/Part:BBa_J23100 Anderson collection] ("Anderson promoters"). They have already been measured in Escherichia coli where they all showed a constitutive behavior with different strength. In this project, eleven Anderson promoters were characterized in B. subtilis with the lux operon as a reporter. In B. subtilis these promoters show quiet low activity (see Data Anderson promoters ![]() ).
To confirm these results some Anderson promoters were also evaluated with the reporter gene lacZ by doing β-galactosidase assays (see Data Anderson promoters
).
To confirm these results some Anderson promoters were also evaluated with the reporter gene lacZ by doing β-galactosidase assays (see Data Anderson promoters ![]() ).
).
- J23100 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823004 BioBrick:BBa_K823004])
- J23101 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823005 BioBrick:BBa_K823005])
- J23102 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823006 BioBrick:BBa_K823006])
- J23103 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823007 BioBrick:BBa_K823007])
- J23106 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823008 BioBrick:BBa_K823008])
- J23107 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823009 BioBrick:BBa_K823009])
- J23113 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823010 BioBrick:BBa_K823010])
- J23114 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823011 BioBrick:BBa_K823011])
- J23115 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823012 BioBrick:BBa_K823012])
- J23117 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823013 BioBrick:BBa_K823013])
- J23118 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823014 BioBrick:BBa_K823014])
Constitutive promoters from B. subtilis
The second group of promoters are the constitutive promoters from B. subtilis. We evaluated the promoters PliaG, Pveg and PlepA. Therefore, we used the lux operon as well as the lacZ gene as reporters.
- PliaG ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823000 BioBrick:BBa_K823000])
PliaG is a weak, constitutive promoter from B. subtilis. It is responsible for the transcription of the last four genes of the liaIHGFSR locus and therefore for the production of the components of the LiaRS system, which is important for the detection of cell wall antibiotics [http://www.ncbi.nlm.nih.gov/pubmed?term=Journal%20of%20Bacteriology%2C%20188%20%2814%29%3A%205153%E2%80%935166: (Jordan et al., 2006)]. PliaG was evaluated with the lux operon (see Data constitutive promoters ![]() ) as well as the lacZ (see Data constitutive promoters
) as well as the lacZ (see Data constitutive promoters ![]() ) as reporter. This promoter showed a much higher activity than the Anderson promoters, but was still weak in comparison to other evaluated Bacillus promoters.
) as reporter. This promoter showed a much higher activity than the Anderson promoters, but was still weak in comparison to other evaluated Bacillus promoters.
- Pveg ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823003 BioBrick:BBa_K823003])
Pveg is known to show a strong constitutive activity during the vegetative growth phase and sporulation. This promoter is important for the transcription of the veg gene, which plays a role during sporulation [http://www.ncbi.nlm.nih.gov/pubmed?term=J.%20Biochem.%2C%20133%20%284%29%3A%20475%E2%80%93483: (Fukushima et al., 2003)]. Pveg was measured by using the reporter gene lacZ (see Data constitutive promoters ![]() ). This promoter was the strongest of our evaluation.
). This promoter was the strongest of our evaluation.
- PlepA ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823002 BioBrick:BBa_K823002])
PlepA is constitutive promoter which is important for the transcription of the bicistronic operon. One of the expressed proteins is the protein PlepA [http://www.ncbi.nlm.nih.gov/pubmed?term=Microbiology%2C%20142%3A%201641%E2%80%931649: (Homuth et al., 1996)]. This protein plays an important role during the translation as it can move the mRNA-tRNA complex one step back in the ribosome which is expected to improve the fidelity of translation [http://www.ncbi.nlm.nih.gov/pubmed?term=Cell%2C%20127%20%284%29%3A%20721%E2%80%93733: (Qin et al., 2006)]. This promoter was evaluated with the lux operon as a reporter (see Data constitutive promoters ![]() ). The activity of this promoter is between the activity of the strongest Bacillus promoter Pveg and the weak PliaG.
). The activity of this promoter is between the activity of the strongest Bacillus promoter Pveg and the weak PliaG.
Inducible promoters from B. subtilis
The last group of promoters consists of two inducible promoters of B. subtilis , PliaI and Pxyl-XylR. They are useful to decide when to turn on gene expression because these promoters need an inducer to start transcription. They are evaluated with the reporters lux and lacZ.
- PliaI ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823001 BioBrick:BBa_K823001])
PliaI is an inducible promoter from B. subtilis which is induced by antibiotics that interfere with integrity and biosynthesis of the cell wall. In the presence of a stimulus, the two-component system LiaRS is activated. The activated response regulator LiaR binds to the operator of the promoter and induces the transcription of the lia locus. When the promoter is turned on the two proteins LiaI and LiaH are expressed which play an important role in the stress response [http://www.ncbi.nlm.nih.gov/pubmed?term=Journal%20of%20Bacteriology%2C%20188%20%2814%29%3A%205153%E2%80%935166: (Jordan et al., 2006)]. This promoter is evaluated with the reporter lux (see Data inducible promoters ![]() ) as well as lacZ (see Data inducible promoters
) as well as lacZ (see Data inducible promoters ![]() ). The induction was measured with different concentrations of bacitracin. By induction with different bacitracin concentrations we were able to cover a large range of promoter activity.
). The induction was measured with different concentrations of bacitracin. By induction with different bacitracin concentrations we were able to cover a large range of promoter activity.
- PXyl-xylR ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K8230015 BioBrick:BBa_K823015])
PXyl-xylR is a xylose-inducible promoter from B. subtilis. XylR is the repressor of PXyl in the absence of the sugar Xylose. In the presence of Xylose, XylR dissociates from the operator and PXyl is active (see e.g. [http://www.ncbi.nlm.nih.gov/pubmed/2544559 Kreuzer et al.]). For this promoter we have not yet suceeded to clone it in a reporter vector to evaluate the activity. So far, there is no data for this promoter.
Bacillus Reporters 
We designed some reporters that are commonly used in B. subtilis or are codon optimized versions of popular reporter genes. All reporters have a modified iGEM Freiburg standard ([http://partsregistry.org/Help:Assembly_standard_25 RCF 25]) pre- and suffix for assembly of in-frame fusion proteins. Our prefix also includes the B. subitlis optimized ribosome binding site.
prefix: GAATTCCGCGGCCGCTTCTAGATAAGGAGGAACTACTATGGCCGGC
suffix: ACCGGTTAATACTAGTAGCGGCCGCTGCAGT
- GFP ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823039 BioBrick:BBa_K823039])
- mKate2 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823029 BioBrick:BBa_K823029])
|
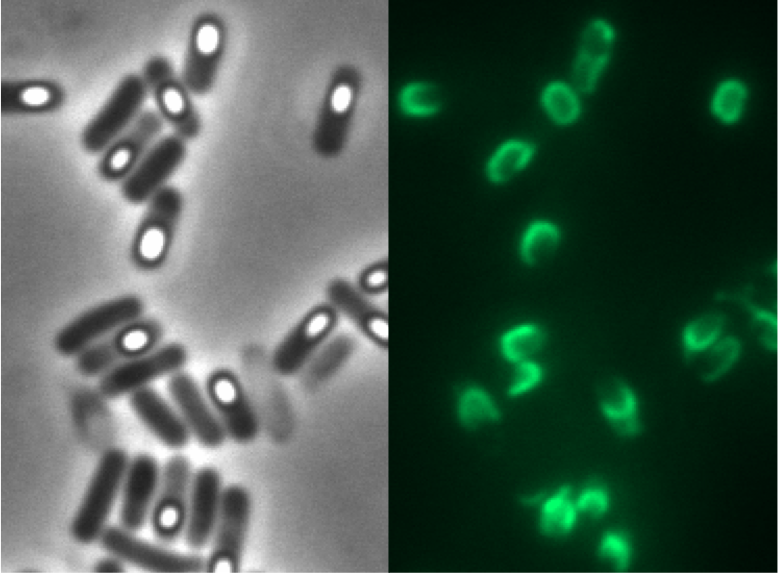
We synthesized this monomeric far-red fluorescence protein with a codon-optimized version for the use in B. subtilis with pre- and suffix of the Freiburg standard. We cloned this reporter in front of the terminator B0014. For the evaluation this reporter was successfully combined with the promoters PliaI, PlepA and the Anderson promoter J23101 in the empty Bacillus vector pSBBs1C from our BacillusBioBrickBox. At the moment we have the right construct which is integrated into the chromosome of B. subtilis. Unfortunately we have no equipment to measure this reporter. Neither our plate reader nor the fluorescent microscope have the required filters.
- LacZ ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823019 BioBrick:BBa_K823019])
This lacZ gene is derived from the Bacillus reporter vector pAC6. It is constructed in the Freiburg Standard (Assembly 25) for in-frame fusion proteins. It also includes a ribosome binding site optimized for Bacillus subtilis translation. This lacZ BioBrick was tested in the expression vector pSBBs0K-Pspac. This construct gives output, so the reporter gene is functional. See Data in the vector evaluation section of pSBBs0K-Pspac.
- luc ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K823028 BioBrick:BBa_K823028])
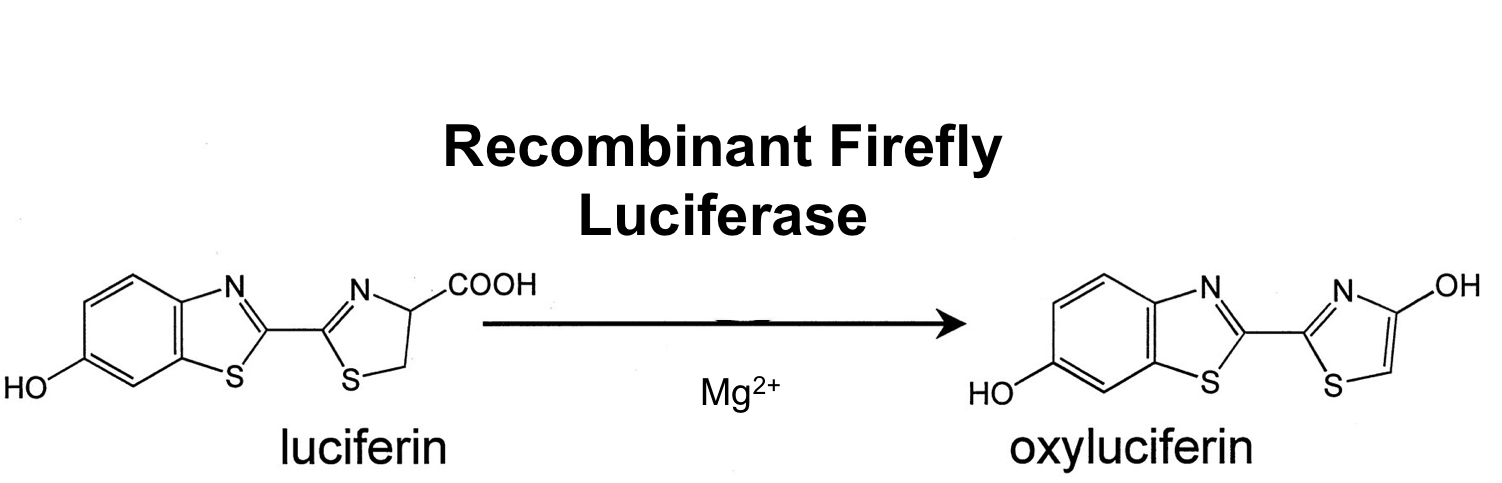
This is the luc monomeric firefly luciferase which needs a special substrate to produce luminescence. With Ribosome binding site included, codon optimized for Bacillus subtilis and synthesized by gene art. It was used in B. subtilis before ([http://www.ncbi.nlm.nih.gov/pubmed/21552330 Mirouze et al. 2011]).
Firefly luciferase is by far the most commonly used bioluminescent reporter. This monomeric enzyme of 61kDa catalyzes a two-step oxidation reaction to yield light, usually in the green to yellow region, typically 550–570nm . The first step is activation of the luciferyl carboxylate by ATP to yield a reactive mixed anhydride. In the second step, this activated intermediate reacts with oxygen to create a transient dioxetane that breaks down to the oxidized products, oxyluciferin and CO2. Upon mixing with substrates, firefly luciferase produces an initial burst of light that decays over about 15 seconds to a low level of sustained luminescence. This kinetic profile reflects the slow release of the enzymatic product, thus limiting catalytic turnover after the initial reaction.
The popularity of native firefly luciferase as a genetic reporter is due to the sensitivity and convenience of the enzyme assay and tight coupling of protein synthesis with enzyme activity. Firefly luciferase, which is encoded by the luc gene, is a monomer that does not require any post-translational modifications; it is available as a mature enzyme directly upon translation of its mRNA. Catalytic competence is attained immediately after release from the ribosome. Also, luciferase has a very short half-life in cells (approximately 3 hours). Combined, these properties make luciferase an extremely responsive reporter, far more so than other commonly used reporters.
Reference:[http://www.promega.com/resources/product-guides-and-selectors/protocols-and-applications-guide/bioluminescent-reporters/ Promega]
Affinity Tags 
All our tags have been synthesized by [http://de-de.invitrogen.com/site/de/de/home/Products-and-Services/Applications/Cloning/gene-synthesis.html?CID=fl-genesynthesis gene art]. They are designed in Freiburg standard with an included optimized ribosome binding site. We have not yet tested our tags.
prefix: GAATTCCGCGGCCGCTTCTAGATAAGGAGGAACTACTATGGCCGGC
suffix: ACCGGTTAATACTAGTAGCGGCCGCTGCAGT
- 3x Flag - tag [http://partsregistry.org/Part:BBa_K823034 (BioBrick:BBa_K823034)]
The Flag-tag was the first epitope tag to be published ([http://www.nature.com/nbt/journal/v6/n10/full/nbt1088-1204.html T.P. Hopp, K.S. Prickett et al. (1988)]). It consists of eight hydrophobic aminoacids: DYKDDDDK and the 3x Flag tag is: DYKDHDGDYKDHDIDYKDDDDK. There are a variety of monoclonal antibodies against this tag, N-terminal as well as position insensitive.
- HA - tag [http://partsregistry.org/Part:BBa_K823035 (BioBrick:BBa_K823035)]
The HA-tag is an epitope derived from the HA-virus. There was first an antibody against it and then the epitope was characterized ([http://www.ncbi.nlm.nih.gov/pubmed/6204768 Wilson, I.A. et al. (1984)]). It was then furthermore used as a tag for protein purification and recognition ([http://www.ncbi.nlm.nih.gov/pubmed/2455217 Field, J. et al. (1988)]). The aminoacid sequence is: YPYDVPDYA.
- cMyc - tag [http://partsregistry.org/Part:BBa_K823036 (BioBrick:BBa_K823036)]
The cMyc-tag is a tag derived from the cMyc gene product. Antibodies were derived from the immunisation with synthetic peptides from the cMyc sequence ([http://mcb.asm.org/content/5/12/3610.short Mol. Cell. Biol. 5]). The aminoacid sequence is EQKLISEEDL.
- His - tag [http://partsregistry.org/Part:BBa_K823037 (BioBrick:BBa_K823037)]
The His-tag is a metal chelating peptide ([http://www.nature.com/nbt/journal/v6/n11/full/nbt1188-1321.html Hochuli, E.; Bannwarth, W.; Döbeli, H.; Gentz, R.; Stüber, D. (1988)]) consisting of at least 6 histidins. It can therefore be used for protein purification by metal2+-ion containing columns (mostly nickel or cobalt). There are also antibodies against this tag, or nickel/cobalt containing fluorescent probes can be used for detection. Also a immobilization is possible in nickel/cobalt coated plastikware. The aminoacid sequence is:HHHHHHHHHH
- Strep - tag [http://partsregistry.org/Part:BBa_K823038 (BioBrick:BBa_K823038)]
The Strep-tag is a mimicry peptide of biotin which binds to Streptavidin ([http://www.sciencedirect.com/science/article/pii/S1050386299000339 Skerra, A. and Schmidt, T.G.M. (1999)]). Its sequence is WSHPQFEK. It can be used for protein purification, immobilisation with Streptavidin or Strep-tactin ([http://www.ncbi.nlm.nih.gov/pubmed/9415448 Voss, S. and Skerra, A. (1997)]) or detection with Strep-tactin or antibodies.

| 
| 
| 
|
| Bacillus Intro | Bacillus BioBrickBox | Sporobeads | Germination STOP |
 "
"