Team:Amsterdam/project/applications/main applications
From 2012.igem.org
(Difference between revisions)
| (15 intermediate revisions not shown) | |||
| Line 7: | Line 7: | ||
<div id="sub-menu" class="content-block"> | <div id="sub-menu" class="content-block"> | ||
| - | <h1>Applications</h1> | + | <h1>Facets and Applications</h1> |
| - | + | The Cellular Logbook regards quite unexplored and fundamental based research. Our projects holds a platform for new technology. But there are already some applications that are feasible in the near future. | |
<center>__TOC__</center> | <center>__TOC__</center> | ||
| - | < | + | |
| - | < | + | <h1>Facets</h1> |
| - | < | + | <h4>Combining all sensors: the Cellular Logbook</h4> |
One of the more popular themes in iGEM projects is the creation of a biosensor for a specific product, and as such the iGEM part registry contains many sensors. Every year a lot of newly developed sensors are added. These sensors are very much needed in today’s world where many new threats and problems arise as unexpected dangers. However, most of these biosensors are fundamentally different in design, making it hard to have multiple sensors in one system. On top of that, many previous iGEM teams used fluorescence, pH or electrical conductance as a readout mechanism. | One of the more popular themes in iGEM projects is the creation of a biosensor for a specific product, and as such the iGEM part registry contains many sensors. Every year a lot of newly developed sensors are added. These sensors are very much needed in today’s world where many new threats and problems arise as unexpected dangers. However, most of these biosensors are fundamentally different in design, making it hard to have multiple sensors in one system. On top of that, many previous iGEM teams used fluorescence, pH or electrical conductance as a readout mechanism. | ||
| Line 20: | Line 20: | ||
Our system can be linked to any other sensory system introduced into a microorganism, therefore creating a '''multiple-sensor-microorganism'''. Since the registration of a signal occurs via methylation of a specific DNA sequence called Memory Part (MP), a specific signal can be stored effectively for either a short or a longer period of time, to eventually be read-out in an easy digestion providing a simple yes or no answer. | Our system can be linked to any other sensory system introduced into a microorganism, therefore creating a '''multiple-sensor-microorganism'''. Since the registration of a signal occurs via methylation of a specific DNA sequence called Memory Part (MP), a specific signal can be stored effectively for either a short or a longer period of time, to eventually be read-out in an easy digestion providing a simple yes or no answer. | ||
| - | < | + | <h4>Time Indicator</h4> |
| - | < | + | |
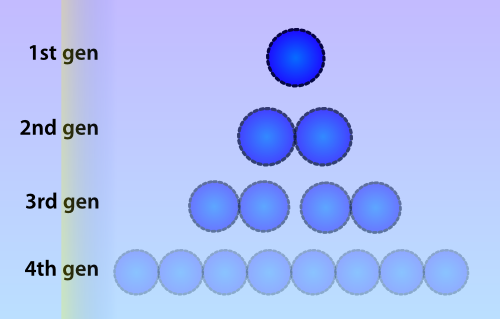
| - | < | + | [[File:Celldivision.png|thumb|right|300px|Due to cell division, the amount of methylated plasmids will be approximately halved during each division cycle. In this picture a lower opacity indicates a lower amount of methylated plasmids]] |
| + | Numerous identical plasmids are often present in single cells and plasmids replicate independently of the bacterial chromosome (Scott 1984). A plasmid copy number (PCN) has been determined for all plasmids in the Parts Registry, which indicates a likely amount of copies of the plasmid to be present in each cell. Unlike eukaryotes, prokaryotes do not copy DNA methylation patterns to the newly synthesized strand during DNA replication. This will lead to a dilution of the amount of ‘written’-plasmids over time, mostly due to cell replication and the ensuing binomial division of the plasmids in the parent cell among the two daughter cells. | ||
| + | |||
| + | The volatilty of this memory design seemed a downside at first, but quickly opened our eyes to a very exciting feature of this system. By analyzing the fraction at the time of memory read-out, the time at which the signal was registered can be inferred. | ||
| + | |||
| + | <h1>Applications</h1> | ||
| + | <h4>Debugger</h4> | ||
Science relies on experimentations. Besides simply measuring or tracking a substance or concentration biologists can encounter unexplainable or vague findings, especially when experimenting with modified or synthetically engineered proteins or genetic networks. Many general questions rise: Is the pathway I’m working with still active? Does my introduced or suspected protein accurately activate my gene? | Science relies on experimentations. Besides simply measuring or tracking a substance or concentration biologists can encounter unexplainable or vague findings, especially when experimenting with modified or synthetically engineered proteins or genetic networks. Many general questions rise: Is the pathway I’m working with still active? Does my introduced or suspected protein accurately activate my gene? | ||
| Line 29: | Line 34: | ||
The system of our Cellular Logbook can help to answer these questions. Our methylation based system only requires a zinc finger and methyltransferase fusion protein (ZnF-Mtase) and our specific Memory Part. Any promoter can be set before our Mtase and can thus be effectively tested. And since the memory plasmid is so expandable not just one but multiple promoters can be tested in the same experiment if a unique zinc finger is linked to a methyltransferase for every sensor. This allows any scientist to simultaneously test all parts of a pathway or '''multiple pathways''' at the same time, creating a fuller understanding of any complex mechanism. | The system of our Cellular Logbook can help to answer these questions. Our methylation based system only requires a zinc finger and methyltransferase fusion protein (ZnF-Mtase) and our specific Memory Part. Any promoter can be set before our Mtase and can thus be effectively tested. And since the memory plasmid is so expandable not just one but multiple promoters can be tested in the same experiment if a unique zinc finger is linked to a methyltransferase for every sensor. This allows any scientist to simultaneously test all parts of a pathway or '''multiple pathways''' at the same time, creating a fuller understanding of any complex mechanism. | ||
| + | <h4>Clean Water Supply Detection</h4> | ||
| + | If we are able to use our bacteria in the environment, our Cellular Logbook can be used to sense toxic signals in the environment. One of the places in need for toxic detection are the water supplies, ponds and rivers in the Netherlands. We had a talk with Ron van der Oost from Waternet [https://www.waternet.nl/about-waternet/]. During this talk we came to the conclusion that a multi sensor that is able to detect 20 different toxic groups, and is able to detect if the concentration has surpassed a specific amount, would be greatly cost reducing compared to the current setup for detecting if a water supply is clean of toxics, which can cost up to 40.000 euros / place. To achieve this we need a system that contains the bacteria, so no bacteria will roam in the environment. The KWR Water Cycle Research Institute[http://www.kwrwater.nl/] has developed a flow-trough sensor that can also serve as a container for our sensor[http://www.kwrwater.nl/uploadedFiles/Website_KWR/Publicaties_%40_Producten/Posters/Development%20of%20a%20water%20toxicity%20sensor%20based%20on%20genetically%20modified%20bacteria.pdf]. | ||
| + | |||
| + | <h4>Compound Emission Detection at Industrial Sites</h4> | ||
| + | The Cellular Logbook can be used at fabrics to measure compound emission. During a talk with Bart van den Burg from the company Biodetection Systems[http://www.bds.nl/], who currently use bioassays to detect compounds in samples, we came to the realization that the Cellular Logbook can be used as a cheap detection system for compound emission of industrial sites. If the Cellular Logbook cells are captured in a system that prevents them from being released in the environment, the Cellular Logbook can be placed at multiple locations, after which they are able tell us where, when and in what concentration a certain chemical has been detected. | ||
| + | |||
| + | <h1>Global Challenges</h1> | ||
| + | We took the global challenges as given by The Millenium Project[http://www.millennium-project.org/millennium/challeng.html] to show what the Cellular Logbook can contribute to worldwide societal problems. As a result we have identified four challenges where the Cellular Logbook is able to significantly contribute to the solution. | ||
| + | |||
| + | <h4>How can everyone have sufficient clean water without conflict?</h4> | ||
| + | One of the main problems in 3rd world countries concerning clean water supplies, apart from the cleaning of the water, is the detection of contaminated water. The Cellular Logbook can serve as a multi-sensor that comprises all the common causes for contaminated water in the 3rd world. Using this multi-sensor we have a cheap and efficient way of detecting contaminated water that is affordable in 3rd world countries. | ||
| - | </ | + | <h4>How can the threat of new and reemerging diseases and immune micro-organisms be reduced?</h4> |
| - | + | Our multi-sensor can be used to detect these diseases but also to monitor places at risk. Since it can be adapted to use any sensor it can be used to effectively scan for several threats. And being inside a live organism our logbook provides a longer measurement instead of just capturing a single moment. Thus giving a much better insight into the situation at hand. | |
| - | < | + | <h4>How can growing energy demands be met safely and efficiently?</h4> |
| + | Using the Cellular logbook adapted to any specific waste or other suspected threats produced when this energy demand is met, it can efficiently provide a means for safer outcome. | ||
| - | + | <h4>How can scientific and technological breakthroughs be accelerated to improve the human condition?</h4> | |
| - | + | By using our Cellular logbook platform as described for the Debugger. Using the Debugger can provide much faster insight when testing pathways or if you just want to find out whether or not your system is working. These fast means of insight can generate a new bundle of information and save precious time for any researcher. | |
</div> | </div> | ||
Latest revision as of 14:49, 24 September 2012
 "
"