Team:University College London/LabBook/Week14
From 2012.igem.org



Contents |
Monday 10.09.12
Aim: Day 2 of Generating competent cells. The aim is to incubate cells in the presence of CaCl2 to prepare the cell wall, such that it becomes permeable to DNA
Method: https://2012.igem.org/Team:University_College_London/Protocols/Competency2
Tuesday 11.09.12
Aim: Day 3 of Generating competent cells
Method: https://2012.igem.org/Team:University_College_London/Protocols/Competency3

14-2

Tuesday 11.09.12
Step 1 - Aim:
To generate enough plasmid backbone (PSB1C3) to be used in a 3A ligation. This is followed by a PCR clean-up.
Methods:
https://2012.igem.org/Team:University_College_London/LabBook/Protocols/PCR#Transformation_Protocol_2
Step 2 - Aim:
To check whether PCR is successful. After the PCR reaction, we can detect the presence of the correct products by running a 1% gel electrophoresis.
Methods:
https://2012.igem.org/Team:University_College_London/Protocols/Electrophoresis
Results:
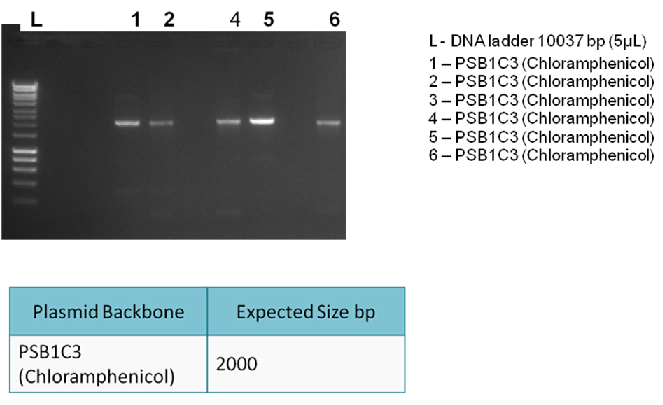
The following gel results indicate whether PCR was successful.
Conclusion:
Since band sizes were as expected, the PCR was considered to be successful. Hence, the backbone was purified and used in the 3A ligation.

Thursday 13.09.12
Aim:
To generate enough linker (BBa_J23119 + BBa_B0034) to be used in a 3A ligation. This is followed by a PCR clean-up.
Methods: https://2012.igem.org/Team:University_College_London/LabBook/Protocols/PCR#Transformation_Protocol_2
Step 2 - Aim:
To check whether PCR is successful. After the PCR reaction, we can detect the presence of the correct products by running a 1% gel electrophoresis.
Methods:
https://2012.igem.org/Team:University_College_London/Protocols/Electrophoresis
Results:
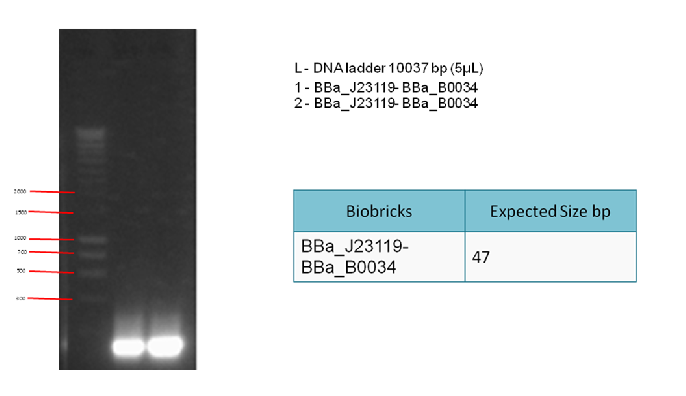
The following gel results indicate whether PCR was successful.
Conclusion:
Since we obtained the expected bands in the gel, the PCR was considered successful and the purified DNA used in the 3A ligation.

Friday 14.09.13
Aim: To characterise nuclease activity BL21 cells
Method:
To add protocol
Results:
The following table shows readings of the diameters and OD at different time points:
| Date | Time | Colony diameter/mm | Halo diameter/mm | Absorbance at 600 OD |
|---|---|---|---|---|
| Friday | 12.30 | 0 | 0 | 0 |
| Friday | 16.30 | 0 | 0 | 0 |
| Friday | 19.30 | 0 | 0 | 0 |
| Friday | 22.30 | 0 | 0 | 0 |
| Saturday | 01.30 | 0.5 | 1.5 | 0.068 |
| Saturday | 04.30 | 1 | 2 | 0.098 |
| Saturday | 07.30 | 1.5 | 3 | 0.159 |
| Saturday | 10.30 | 1.5 | 3.5 | 0.192 |
| Saturday | 12.30pm | 2 | 4 | 0.203 |
| Saturday | 14.30 | 2 | 4 | 0.209 |
| Saturday | 16.30 | 2.5 | 5 | 0.215 |
The average depth of the colonies was noted to be 0.5 - 1mm
Conclusion:
It can be seen that nuclease works as expected in BL21 cells. However, nuclease activity is less efficient when compared to nuclease activity in WnU cells. This is in line with expectations, since nuclease is found naturally in WnU cells but not in BL21 cells.
 "
"