|
|
| (39 intermediate revisions not shown) |
| Line 1: |
Line 1: |
| - | <!-- *** What falls between these lines is the Alert Box! You can remove it from your pages once you have read and understood the alert *** -->
| |
| - |
| |
| | <html> | | <html> |
| - | <div id="box" style="width: 700px; margin-left: 137px; padding: 5px; border: 3px solid #000; background-color: #fe2b33;"> | + | <style type=text/css> |
| - | <div id="template" style="text-align: center; font-weight: bold; font-size: large; color: #f6f6f6; padding: 5px;">
| + | |
| - | This is a template page. READ THESE INSTRUCTIONS.
| + | body, #content { |
| - | </div>
| + | background: #FFFFFF |
| - | <div id="instructions" style="text-align: center; font-weight: normal; font-size: small; color: #f6f6f6; padding: 5px;">
| + | } |
| - | You are provided with this team page template with which to start the iGEM season. You may choose to personalize it to fit your team but keep the same "look." Or you may choose to take your team wiki to a different level and design your own wiki. You can find some examples <a href="https://2008.igem.org/Help:Template/Examples">HERE</a>.
| + | |
| - | </div>
| + | body { |
| - | <div id="warning" style="text-align: center; font-weight: bold; font-size: small; color: #f6f6f6; padding: 5px;">
| + | margin: 10px 0 0 0; |
| - | You <strong>MUST</strong> have all of the pages listed in the menu below with the names specified. PLEASE keep all of your pages within your teams namespace.
| + | font-family:arial; |
| - | </div>
| + | } |
| - | </div> | + | |
| | + | #content { |
| | + | padding:10px auto; |
| | + | margin:0 auto; |
| | + | color: black; |
| | + | position: relative; |
| | + | border: none; |
| | + | } |
| | + | |
| | + | #whole_wrapper { |
| | + | margin:100px 0 0 0; |
| | + | } |
| | + | |
| | + | #right_wrapper2 { |
| | + | padding: 0 0 0 72px; |
| | + | } |
| | + | |
| | + | h1 { |
| | + | border-bottom: 1px solid #AAA; |
| | + | padding: 0 0 10px 0; |
| | + | } |
| | + | |
| | + | #title { |
| | + | width:85%; |
| | + | padding-left:20px; |
| | + | color:crimson; |
| | + | border-bottom: 1px solid #AAA; |
| | + | padding: 0 0 10px 0; |
| | + | } |
| | + | |
| | + | |
| | + | |
| | + | table { |
| | + | margin: 0 0 30px 0; |
| | + | } |
| | + | |
| | + | td { |
| | + | padding: 5px; |
| | + | } |
| | + | |
| | + | #top-section { |
| | + | width: 965px; |
| | + | height: 0; |
| | + | margin: 0 auto; |
| | + | padding: 0; |
| | + | border: none; |
| | + | } |
| | + | |
| | + | .left-menu:hover { |
| | + | background-color: #FFFFFF; |
| | + | } |
| | + | #menubar li a { |
| | + | background-color: #FFFFFF; |
| | + | } |
| | + | #menubar:hover { |
| | + | color: black; |
| | + | } |
| | + | #menubar li a { |
| | + | color: white; |
| | + | } |
| | + | #menubar:hover li a { |
| | + | color: black; |
| | + | } |
| | + | #menubar > ul > li:last-child { |
| | + | display:none; |
| | + | } |
| | + | |
| | + | dt {width:90%;padding-left:25px;} |
| | + | |
| | + | dd {width:85%;padding-left:20px;} |
| | + | |
| | + | /*hide default igem banner and reformat style into blank slate*/ |
| | + | |
| | + | #p-logo {display:none;} |
| | + | |
| | + | #top-section {border:none;} |
| | + | |
| | + | #menubar a {color:#000000;} |
| | + | #menubar a:hover{text-decoration:none; color:#52749C;} |
| | + | |
| | + | |
| | + | .printfooter {display:none;} |
| | + | #footer-box {border:none;} |
| | + | #catlinks {display:none;} |
| | + | .firstHeading {display:none;} |
| | + | |
| | + | </style> |
| | </html> | | </html> |
| | | | |
| - | <!-- *** End of the alert box *** --> | + | <html> |
| | + | <head> |
| | + | </head> |
| | + | <body style="background-color:#000000;"> |
| | + | </body> |
| | + | <html dir="ltr" lang="en"> |
| | + | <head> |
| | + | <meta charset="UTF-8" /> |
| | + | <title>Our BioBricks « E. musici</title> |
| | + | <link rel="profile" href="http://gmpg.org/xfn/11" /> |
| | + | <link rel="stylesheet" href="http://s1.wp.com/wp-content/themes/pub/triton-lite/style.css?m=1346172069g" type="text/css" media="screen" /> |
| | + | <link rel="pingback" href="http://uscigem2012.wordpress.com/xmlrpc.php" /> |
| | + | <link rel="alternate" type="application/rss+xml" title="E. musici » Feed" href="http://uscigem2012.wordpress.com/feed/" /> |
| | + | <link rel="alternate" type="application/rss+xml" title="E. musici » Comments Feed" href="http://uscigem2012.wordpress.com/comments/feed/" /> |
| | + | <link rel="alternate" type="application/rss+xml" title="E. musici » Our BioBricks Comments Feed" href="http://uscigem2012.wordpress.com/our-biobricks/feed/" /> |
| | + | <script type="text/javascript"> |
| | + | /* <![CDATA[ */ |
| | + | function addLoadEvent(func){var oldonload=window.onload;if(typeof window.onload!='function'){window.onload=func;}else{window.onload=function(){oldonload();func();}}} |
| | + | /* ]]> */ |
| | + | </script> |
| | + | <link rel='stylesheet' id='all-css-0' href='http://s1.wp.com/_static/??/wp-content/blog-plugins/loggedout-follow/widget.css,/wp-content/mu-plugins/post-flair/style.css,/wp-content/mu-plugins/post-flair/sharing/sharing.css,/wp-content/themes/h4/global.css?m=1345177132j' type='text/css' media='all' /> |
| | + | <link rel='stylesheet' id='print-css-0' href='http://s0.wp.com/wp-content/mu-plugins/global-print/global-print.css?m=1335386953g' type='text/css' media='print' /> |
| | + | <script type='text/javascript'> |
| | + | /* <![CDATA[ */ |
| | + | var LoggedOutFollow = {"invalid_email":"Your subscription did not succeed, please try again with a valid email address."}; |
| | + | /* ]]> */ |
| | + | </script> |
| | + | <script type='text/javascript' src='http://s2.wp.com/_static/??/wp-includes/js/jquery/jquery.js,/wp-content/blog-plugins/loggedout-follow/widget.js,/wp-includes/js/comment-reply.js?m=1333391137j'></script> |
| | + | <link rel='stylesheet' id='all-css-0' href='http://s1.wp.com/wp-content/mu-plugins/highlander-comments/style.css?m=1343991657g' type='text/css' media='all' /> |
| | + | <!--[if lt IE 8]> |
| | + | <link rel='stylesheet' id='highlander-comments-ie7-css' href='http://s2.wp.com/wp-content/mu-plugins/highlander-comments/style-ie7.css?m=1307172283g&ver=20110606' type='text/css' media='all' /> |
| | + | <![endif]--> |
| | + | <link rel="EditURI" type="application/rsd+xml" title="RSD" href="http://uscigem2012.wordpress.com/xmlrpc.php?rsd" /> |
| | + | <link rel="wlwmanifest" type="application/wlwmanifest+xml" href="http://uscigem2012.wordpress.com/wp-includes/wlwmanifest.xml" /> |
| | + | <link rel='prev' title='Project' href='http://uscigem2012.wordpress.com/project/' /> |
| | + | <link rel='next' title='Lab Notebook' href='http://uscigem2012.wordpress.com/lab-notebook/' /> |
| | + | <meta name="generator" content="WordPress.com" /> |
| | + | <link rel='canonical' href='http://uscigem2012.wordpress.com/our-biobricks/' /> |
| | + | <link rel='shortlink' href='http://wp.me/P2CDGD-c' /> |
| | + | <link rel="alternate" type="application/json+oembed" href="http://public-api.wordpress.com/oembed/1.0/?format=json&url=http%3A%2F%2Fuscigem2012.wordpress.com%2Four-biobricks%2F&for=wpcom-auto-discovery" /><link rel="alternate" type="application/xml+oembed" href="http://public-api.wordpress.com/oembed/1.0/?format=xml&url=http%3A%2F%2Fuscigem2012.wordpress.com%2Four-biobricks%2F&for=wpcom-auto-discovery" /><meta property="og:type" content="article" /> |
| | + | <meta property="og:title" content="Our BioBricks" /> |
| | + | <meta property="og:url" content="http://uscigem2012.wordpress.com/our-biobricks/" /> |
| | + | <meta property="og:description" content="Below are the BioBrick parts our team created and submitted to the iGEM Registry. Click images to enlarge. Featured BioBricks are highlighted in yellow. BBa_K842006 cheA Description and Function ..." /> |
| | + | <meta property="og:site_name" content="E. musici" /> |
| | + | <meta property="og:image" content="http://uscigem2012.files.wordpress.com/2012/07/flia.gif?w=200" /> |
| | + | <meta property="og:image" content="http://uscigem2012.files.wordpress.com/2012/07/flhd.gif?w=200" /> |
| | + | <meta property="og:image" content="http://uscigem2012.files.wordpress.com/2012/07/flgj.gif?w=200" /> |
| | + | <meta property="og:image" content="http://uscigem2012.files.wordpress.com/2012/07/t7_rbs_chez.gif?w=200" /> |
| | + | <meta property="og:image" content="http://uscigem2012.files.wordpress.com/2012/07/t7_rbs_chey.gif?w=200" /> |
| | + | <meta name="twitter:image" content="http://uscigem2012.files.wordpress.com/2012/07/flia.gif" /> |
| | + | <meta name="twitter:card" content="summary" /> |
| | + | <meta property="fb:app_id" content="249643311490" /> |
| | + | <link rel="shortcut icon" type="image/x-icon" href="http://s2.wp.com/i/favicon.ico?m=1311976025g" sizes="16x16 24x24 32x32 48x48" /> |
| | + | <link rel="icon" type="image/x-icon" href="http://s2.wp.com/i/favicon.ico?m=1311976025g" sizes="16x16 24x24 32x32 48x48" /> |
| | + | <link rel="apple-touch-icon-precomposed" href="http://s0.wp.com/i/webclip.png?m=1311618091g" /> |
| | + | <link rel='openid.server' href='http://uscigem2012.wordpress.com/?openidserver=1' /> |
| | + | <link rel='openid.delegate' href='http://uscigem2012.wordpress.com/' /> |
| | + | <link rel="search" type="application/opensearchdescription+xml" href="http://uscigem2012.wordpress.com/osd.xml" title="E. musici" /> |
| | + | <link rel="search" type="application/opensearchdescription+xml" href="http://wordpress.com/opensearch.xml" title="WordPress.com" /> |
| | + | <style> |
| | + | /* <![CDATA[ */ |
| | + | /* Block: reblog */ |
| | + | |
| | + | .reblog-from img { margin: 0 10px 0 0; vertical-align: middle; padding: 0; border: 0; } |
| | + | .reblogger-note img.avatar { float: left; padding: 0; border: 0; } |
| | + | .reblogger-note-content { margin: 0 0 20px; } |
| | + | .reblog-post .wpcom-enhanced-excerpt-content { border-left: 3px solid #eee; padding-left: 15px; } |
| | + | .reblog-post ul.thumb-list { display: block; list-style: none; margin: 2px 0; padding: 0; clear: both; } |
| | + | .reblog-post ul.thumb-list li { display: inline; margin: 0; padding: 0 1px; border: 0; } |
| | + | .reblog-post ul.thumb-list li a { margin: 0; padding: 0; border: 0; } |
| | + | .reblog-post ul.thumb-list li img { margin: 0; padding: 0; border: 0; } |
| | + | |
| | + | .reblog-post .wpcom-enhanced-excerpt { clear: both; } |
| | + | |
| | + | .reblog-post .wpcom-enhanced-excerpt address, |
| | + | .reblog-post .wpcom-enhanced-excerpt li, |
| | + | .reblog-post .wpcom-enhanced-excerpt h1, |
| | + | .reblog-post .wpcom-enhanced-excerpt h2, |
| | + | .reblog-post .wpcom-enhanced-excerpt h3, |
| | + | .reblog-post .wpcom-enhanced-excerpt h4, |
| | + | .reblog-post .wpcom-enhanced-excerpt h5, |
| | + | .reblog-post .wpcom-enhanced-excerpt h6, |
| | + | .reblog-post .wpcom-enhanced-excerpt p { font-size: 100% !important; } |
| | + | |
| | + | .reblog-post .wpcom-enhanced-excerpt blockquote, |
| | + | .reblog-post .wpcom-enhanced-excerpt pre, |
| | + | .reblog-post .wpcom-enhanced-excerpt code, |
| | + | .reblog-post .wpcom-enhanced-excerpt q { font-size: 98% !important; } |
| | + | |
| | | | |
| - | {| style="color:#1b2c8a;background-color:#0c6;" cellpadding="3" cellspacing="1" border="1" bordercolor="#fff" width="62%" align="center"
| + | /* ]]> */ |
| - | !align="center"|[[Team:USC|Home]]
| + | </style> |
| - | !align="center"|[[Team:USC/Team|Team]]
| + | |
| - | !align="center"|[https://igem.org/Team.cgi?year=2012&team_name=USC Official Team Profile]
| + | <style type="text/css"> |
| - | !align="center"|[[Team:USC/Project|Project]]
| + | /* <![CDATA[ */ |
| - | !align="center"|[[Team:USC/Parts|Parts Submitted to the Registry]]
| + | /* |
| - | !align="center"|[[Team:USC/Modeling|Modeling]]
| + | * Link color |
| - | !align="center"|[[Team:USC/Notebook|Notebook]]
| + | */ |
| - | !align="center"|[[Team:USC/Safety|Safety]]
| + | a, |
| - | !align="center"|[[Team:USC/Attributions|Attributions]]
| + | .post-wrap a, |
| - | |}
| + | .by-author a, |
| | + | #posts .post-content .post-foot a { |
| | + | color: #e1c647; |
| | + | } |
| | + | |
| | + | #masthead, |
| | + | #access ul ul, |
| | + | #comments .form-submit input, |
| | + | #comments .form-submit input:hover, |
| | + | #footer { |
| | + | background-color: #bf1d2c; |
| | + | } |
| | + | #access .current-menu-item > a, |
| | + | #access .current-menu-ancestor > a, |
| | + | #access .current_page_item > a, |
| | + | #access .current_page_ancestor > a { |
| | + | background: rgba( 0, 0, 0, 0.3 ); |
| | + | } |
| | + | #access a, |
| | + | #access ul ul a, |
| | + | #comments .form-submit input, |
| | + | #footer { |
| | + | color: rgba( 255, 255, 255, 0.7 ); |
| | + | } |
| | + | #access ul ul a { |
| | + | border-bottom: 1px solid rgba( 0, 0, 0, 0.2 ); |
| | + | } |
| | + | #access ul ul a:hover, |
| | + | #access ul ul :hover > a { |
| | + | background: rgba( 0, 0, 0, 0.3 ); |
| | + | color: rgba( 255, 255, 255, 1 ); |
| | + | } |
| | + | #access .current-menu-item > a, |
| | + | #access .current-menu-ancestor > a, |
| | + | #access .current_page_item > a, |
| | + | #access .current_page_ancestor > a, |
| | + | #footer a { /* Top-level current links */ |
| | + | color: rgba( 255, 255, 255, 1 ); |
| | + | } |
| | + | #access ul ul .current-menu-item > a, |
| | + | #access ul ul .current-menu-ancestor > a, |
| | + | #access ul ul .current_page_item > a, |
| | + | #access ul ul .current_page_ancestor > a { /* Sub-level current links */ |
| | + | background: rgba( 0, 0, 0, 0.3 ); |
| | + | color: rgba( 255, 255, 255, 1 ); |
| | + | } |
| | + | #access ul ul .current-menu-ancestor > a, |
| | + | #access ul ul .current_page_ancestor > a { |
| | + | background: rgba( 0, 0, 0, 0.3 ); |
| | + | } |
| | + | #access ul ul .current-menu-ancestor > a:hover, |
| | + | #access ul ul .current_page_ancestor > a:hover { |
| | + | background: rgba( 0, 0, 0, 0.3 ); |
| | + | } |
| | + | /* ]]> */ |
| | + | </style> |
| | + | <meta name="application-name" content="E. musici" /><meta name="msapplication-window" content="width=device-width;height=device-height" /><meta name="msapplication-tooltip" content="University of Southern California iGEM 2012" /><meta name="msapplication-task" content="name=Subscribe;action-uri=http://uscigem2012.wordpress.com/feed/;icon-uri=http://s2.wp.com/i/favicon.ico" /><meta name="msapplication-task" content="name=Sign up for a free blog;action-uri=http://wordpress.com/signup/;icon-uri=http://s2.wp.com/i/favicon.ico" /><meta name="msapplication-task" content="name=WordPress.com Support;action-uri=http://support.wordpress.com/;icon-uri=http://s2.wp.com/i/favicon.ico" /><meta name="msapplication-task" content="name=WordPress.com Forums;action-uri=http://forums.wordpress.com/;icon-uri=http://s2.wp.com/i/favicon.ico" /><meta name="title" content="Our BioBricks | E. musici on WordPress.com" /> |
| | + | <meta name="description" content="Below are the BioBrick parts our team created and submitted to the iGEM Registry. Click images to enlarge. Featured BioBricks are highlighted in yellow. BBa_K842006 cheA Description and Function cheA is a… Read More →" /> |
| | + | <style type="text/css"> |
| | + | #header-image { |
| | + | background: url( 'http://uscigem2012.files.wordpress.com/2012/10/igem-header.gif' ) no-repeat; |
| | + | float: left; |
| | + | margin: 0 0 20px; |
| | + | width: 960px; |
| | + | height: 270px; |
| | + | } |
| | + | #header-image a { |
| | + | display: block; |
| | + | text-indent: -9999px; |
| | + | width: 100%; |
| | + | height: 100%; |
| | + | } |
| | + | </style> |
| | + | |
| | + | <style type="text/css"> |
| | + | #logo { |
| | + | display: none; |
| | + | } |
| | | | |
| | + | </style> |
| | + | |
| | + | <style type="text/css" id="custom-background-css"> |
| | + | body.custom-background { background-color: #ffffff; } |
| | + | </style> |
| | + | </head> |
| | + | <body class="page page-id-12 page-template-default custom-background single-author one-column gecko highlander-enabled highlander-light"> |
| | | | |
| | + | <div id="masthead"> |
| | + | <div class="container"> |
| | + | <div id="access"> |
| | + | <div class="menu"><ul><li class="page_item page-item-2"><a href="https://2012.igem.org/Team:USC">E. Musici</a></li><li class="page_item page-item-5"><a href="https://2012.igem.org/Team:USC/Team">Team</a></li><li class="page_item page-item-10"><a href="https://2012.igem.org/Team:USC/Project">Project</a></li><li class="page_item page-item-12 current_page_item"><a href="https://2012.igem.org/Team:USC/Parts">Our BioBricks</a></li><li class="page_item page-item-14"><a href="https://2012.igem.org/Team:USC/Notebook">Lab Notebook</a></li><li class="page_item page-item-16"><a href="https://2012.igem.org/Team:USC/HumanPractices">Human Practices</a></li><li class="page_item page-item-45"><a href="https://2012.igem.org/Team:USC/Safety">Safety</a></li></ul></div> |
| | + | </div><!-- #access --> |
| | + | </div><!-- .container --> |
| | + | </div><!-- #masthead --> |
| | | | |
| - | An important aspect of the iGEM competition is the use and creation of standard biological parts. Each team will make new parts during iGEM and will place them in the [http://partsregistry.org Registry of Standard Biological Parts]. The iGEM software provides an easy way to present the parts your team has created . The "groupparts" tag will generate a table with all of the parts that your team adds to your team sandbox. Note that if you want to document a part you need to document it on the [http://partsregistry.org Registry], not on your team wiki.
| + | <div id="header"> |
| | + | <div class="container"> |
| | + | <div id="header-image"> |
| | + | <a href="https://2012.igem.org/Team:USC" title="E. musici" rel="home">E. musici</a> |
| | + | </div><!-- #header-image --> |
| | + | <div id="logo"> |
| | + | <h1> |
| | + | <a href="http://uscigem2012.wordpress.com/">E. musici</a> |
| | + | </h1> |
| | + | <div class="desc"> |
| | + | University of Southern California iGEM 2012 </div><!-- .desc --> |
| | + | </div><!-- #logo --> |
| | + | </div><!-- .container --> |
| | + | </div><!-- #header --> |
| | | | |
| - | Remember that the goal of proper part documentation is to describe and define a part such that it can be used without a need to refer to the primary literature. The next iGEM team should be able to read your documentation and be able to use the part successfully. Also, you should provide proper references to acknowledge previous authors and to provide for users who wish to know more.
| + | |
| | + | <div id="post-12" class="post-12 page type-page status-publish hentry"> |
| | + | <div class="post-content"> |
| | + | <h2 class="postitle"> |
| | + | Our BioBricks </h2><!-- .postitle --> |
| | | | |
| | + | <p style="text-align:center;"><span style="color:#000000;">Below are the BioBrick parts our team created and submitted to the iGEM Registry.</span></p> |
| | + | <p style="text-align:center;"><span style="color:#000000;">Click images to enlarge. Featured BioBricks are <SPAN style="BACKGROUND-COLOR: #ffff00">highlighted</SPAN> in yellow.</span></p> |
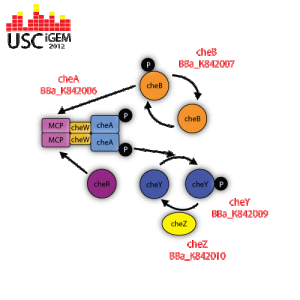
| | + | <p><span style="color:#000000;"><span style="color:#000000;"><a href="http://uscigem2012.files.wordpress.com/2012/07/biobrick-gene-pathway.gif"><img class="aligncenter" title="BioBrick-Gene-Pathway" src="http://uscigem2012.files.wordpress.com/2012/07/biobrick-gene-pathway.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></span></p> |
| | + | <table width="918" border="0" cellspacing="0" cellpadding="0"> |
| | + | <tbody> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/chea1.gif"><img class="aligncenter size-medium wp-image-205" title="cheA" src="http://uscigem2012.files.wordpress.com/2012/07/chea1.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842006</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>cheA</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>cheA</em> is a regulatory protein that becomes activated when the methyl-accepting chemotaxis protein (MCP) is not bound to any attractant. The activated <em>cheA</em> self-phosphorylates itself by hydrolyzing ATP in order to gain a phosphate group. <em>cheA</em> then donates its phosphate group to activate <em>cheB</em> or <em>cheA</em>.</span></p> |
| | + | <p><span style="color:#000000;">Used for researcher control of signaling mechanisms that influence flagella function.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842006"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842006</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used with other chemotaxis genes to form a complete regulatory pathway that controls the response system in MCP and the direction of rotation in flagella.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Chemotaxis protein <em>cheA</em> – Escherichia coli (strain K12). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P07363</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/cheb1.gif"><img class="aligncenter size-medium wp-image-206" title="cheB" src="http://uscigem2012.files.wordpress.com/2012/07/cheb1.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842007</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>cheB</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>CheB</em> removes the methyl groups that are attached to the gamma-glutamyl methyl ester residues located in the methyl-accepting chemotaxis protein (MCP). This allows for the bacteria to remain alert to the changes in the environment.</span></p> |
| | + | <p><span style="color:#000000;">Used for researcher control of signaling mechanisms that influence flagella function.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842007"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842007</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used with other chemotaxis genes to form a complete regulatory pathway that controls the response system in MCP and the direction of rotation in flagella. </span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Chemotaxis response regulator protein-glutamate methylesterase – Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P04042</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/chey1.gif"><img class="aligncenter size-medium wp-image-208" title="cheY" src="http://uscigem2012.files.wordpress.com/2012/07/chey1.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842009</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>cheY</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>CheY</em> is a regulatory protein that reverses the direction of flagella rotation from counter-clockwise to clockwise. When the methyl-accepting chemotaxis protein (MCP) is not bound to any attractant, <em>cheA</em> is activated and told to self-phosphorylate itself by hydrolyzing ATP. Once the <em>cheA</em> is phosphorylated, it transfers its phosphate group to <em>cheY</em>, therefore activating it. <em>CheY</em> then moves to the cytoplasmic side of the flagella apparatus and binds to it. When flagella is not bound to <em>cheY</em>, it rotates in a counter-clockwise direction, which allows the bacteria to move in a “forward” direction. Flagella that is bound to phosphorylated <em>cheY</em> switches its rotation to clockwise, which induces tumbling.</span></p> |
| | + | <p><span style="color:#000000;">Used for researcher control of signaling mechanisms that influence flagella function.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842009"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842009</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used with other chemotaxis genes to form a complete regulatory pathway that controls the response system in MCP and the direction of rotation in flagella.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Chemotaxis protein <em>CheY</em> – Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P0A2D5</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/chez1.gif"><img class="aligncenter size-medium wp-image-209" title="cheZ" src="http://uscigem2012.files.wordpress.com/2012/07/chez1.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842010</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>cheZ</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>CheZ</em> is a regulatory protein that constantly dephosphorylates <em>cheY</em> that is attached to a flagellum. The incessant dephosphorylation is what causes bacteria to run, stop, and tumble when in an environment that has a concentration gradient and remain responsive to any changes in chemical concentration.</span></p> |
| | + | <p><span style="color:#000000;">Used for researcher control of signaling mechanisms that influence flagella function.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842010"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842010</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used with other chemotaxis genes to form a complete regulatory pathway that controls the response system in MCP and the direction of rotation in flagella.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Protein phosphatase <em>CheZ</em> – Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P07800</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/mota1.gif"><img class="aligncenter size-medium wp-image-214" title="motA" src="http://uscigem2012.files.wordpress.com/2012/07/mota1.gif?w=214&h=300" alt="" width="214" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842011</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>motA</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>motA</em> is a structural protein for the stationary element of the flagella motor. The protein works in conjunction with <em>motB</em> to form this structure, which then in turn controls the flagella’s rotation.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842011"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842011</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. The protein works in conjunction with <em>motB</em> to form the stationary base of the flagella motor, which then in turn controls the flagella’s rotation. This part can be used in the complete recreation of an <em>E. coli’s</em> genetic flagellar pathway.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Motility protein A – Escherichia coli (strain K12). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P09348</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/motb1.gif"><img class="aligncenter size-medium wp-image-215" title="motB" src="http://uscigem2012.files.wordpress.com/2012/07/motb1.gif?w=214&h=300" alt="" width="214" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842012</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>motB</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>motB</em> is a structural protein for the stationary element of the flagella motor. The protein works in conjunction with <em>motA</em> to form this structure, which then in turn controls the flagella’s rotation. It is also believed that <em>motB</em> is responsible for forming the structure that connects the flagella motor to the cell wall.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842012"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842012</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. The protein works in conjunction with <em>motA</em> to form the stationary base of the flagella motor, which then in turn controls the flagella’s rotation. This part can be used in the complete recreation of an <em>E. coli’s</em> genetic flagellar pathway.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Motility protein B – Escherichia coli (strain K12). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P0AF06</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/t7_rbs_chez.gif"><img class="aligncenter size-medium wp-image-217" title="T7_RBS_CheZ" src="http://uscigem2012.files.wordpress.com/2012/07/t7_rbs_chez.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842014</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>T7 RBS cheZ</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;">T7 serves as a promoter that is stimulated by IPTG. RBS is a sequence in DNA located upstream of the start codon. It affects the rate at which the open reading frame is translated. <em>CheZ</em> is a regulatory protein that constantly dephosphorylates <em>cheY</em> that is attached to a flagellum. The incessant dephosphorylation is what causes bacteria to run, stop, and tumble when in an environment that has a concentration gradient and remain responsive to any changes in chemical concentration. Together, <em>cheZ</em> is induced by the presence of IPTG.</span></p> |
| | + | <p><span style="color:#000000;">Used for researcher control of signaling mechanisms that influence flagella function.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842014"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842014</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used with other chemotaxis genes to form a complete regulatory pathway that controls the response system in MCP and the direction of rotation in flagella.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Protein phosphatase <em>CheZ</em> – Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P07800</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/chey1.gif"><img class="aligncenter size-medium wp-image-208" title="cheY" src="http://uscigem2012.files.wordpress.com/2012/07/chey1.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842015</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>T7 RBS cheY</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;">T7 serves as a promoter that is stimulated by IPTG. RBS is a sequence in DNA located upstream of the start codon. It affects the rate at which the open reading frame is translated. <em>CheY</em> is regulatory protein that reverses the direction of flagella rotation from counter-clockwise to clockwise. When the methyl-accepting chemotaxis protein (MCP) is not bound to any attractant, <em>cheA</em> is activated and told to self-phosphorylate itself by hydrolyzing ATP. Once the <em>cheA</em> is phosphorylated, it transfers its phosphate group to <em>cheY</em>, therefore activating it. <em>CheY</em> then moves to the cytoplasmic side of the flagella apparatus and binds to it. When flagella are not bound to <em>cheY</em>, it rotates in a counter-clockwise direction, which allows the bacteria to move in a “forward” direction. Flagella that are bound to phosphorylated <em>cheY</em> switches its rotation to clockwise, which induces tumbling.</span></p> |
| | + | <p><span style="color:#000000;">Used for researcher control of signaling mechanisms that influence flagella function.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842015"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842015</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used with other chemotaxis genes to form a complete regulatory pathway that controls the response system in MCP and the direction of rotation in flagella.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Chemotaxis protein <em>CheY</em> – Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P0A2D5</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/flgj.gif"><img class="aligncenter size-medium wp-image-220" title="flgJ" src="http://uscigem2012.files.wordpress.com/2012/07/flgj.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842002</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>flgJ</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
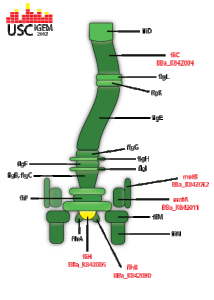
| | + | <p><span style="color:#000000;"><em>flgJ</em> transcribes a flagellum specific muramidase which hydrolyzes the peptidoglycan layer in the cell membrane. This creates a small hole in the side of the bacterium in which the flagellar rod is formed. It is most commonly believed that <em>flgJ</em> is exported via the flagellum specific exportation system, making it a class 3 flagella gene.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842002"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842002</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used in conjunction with the entire flagella synthesis genetic pathway to reproduce a complete flagella apparatus. Additionally, if this part is used separate from the flagellar genetic pathway and connected to a gene promoter, it will continually react with the cell wall until the peptidoglycan layer of the cell membrane can no longer sustain the structure of the bacterium.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Nambu, T., Minamino, T., Macnab, R., & Kutsukake, K. (1999). Peptidoglycan-Hydrolyzing Activity of the <em>FlgJ</em> Protein, Essential for Flagellar Rod Formation in Salmonella typhimurium. <em>Journal of Bacteriology</em>, <em>181</em>(5), 1555-1561. Retrieved October 1, 2012, from http://jb.asm.org/content/181/5/1555</span></p> |
| | + | <p><span style="color:#000000;">Peptidoglycan hydrolase <em>flgJ</em> – Escherichia coli (strain K12). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P75942</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/flia.gif"><img class="aligncenter size-medium wp-image-222" title="fliA" src="http://uscigem2012.files.wordpress.com/2012/07/flia.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842003</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>fliA</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>fliA</em> is an alternate sigma factor for the class 3 flagella operons. When transcribed, it forms a component of RNA polymerase sigma 28.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842003"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842003</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used in conjunction with the entire flagella synthesis genetic pathway to reproduce a complete flagella apparatus.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>fliA</em>:Quickview – EcoliWiki. (n.d.). <em>EcoliWiki</em>. Retrieved October 1, 2012, from http://ecoliwiki.net/colipedia/index.php/fliA</span></p> |
| | + | <p><span style="color:#000000;">Ide, N., & Kutsukake, K. (n.d.). Identification of a novel Escherichia coli gene whose e… [Gene. 1997] – PubMed – NCBI. <em>National Center for Biotechnology Information</em>. Retrieved October 1, 2012, from http://www.ncbi.nlm.nih.gov/pubmed/9358034</span></p> |
| | + | <p><span style="color:#000000;">Claret, L., Miquel, S., Vieille, N., Ryjenkov, D., Gomelsky, M., & Darfeuille-Michaud, A. A. (2007). The Flagellar Sigma Factor <em>FliA</em> Regulates Adhesion and Invasion of Crohn Disease-associated Escherichia coli via a Cyclic Dimeric GMP-dependent Pathway. <em>The Journal Of Biological Chemistry</em>, <em>282</em>(46), 33275-33283. Retrieved October 1, 2012, from http://www.jbc.org/content/282/46/33275.full.pdf</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/flih1.gif"><img class="aligncenter size-medium wp-image-213" title="fliH" src="http://uscigem2012.files.wordpress.com/2012/07/flih1.gif?w=214&h=300" alt="" width="214" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842005</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>fliH</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>FliH</em> functions as part of the flagella specific export system and is necessary for flagellar growth and assembly. It is believed to be part of a two-component system consisting of <em>fliH</em> and <em>fliL</em> that hydrolyses ATP until construction of the flagella apparatus is completed.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842005"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842005</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is to be used in conjunction with the entire flagella synthesis genetic pathway to reproduce a complete flagella apparatus. </span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Minamino, T., & MacNab, R. (n.d.). <em>FliH</em>, a soluble component of the type III flag… [Mol Microbiol. 2000] – PubMed – NCBI. <em>National Center for Biotechnology Information</em>. Retrieved October 1, 2012, from http://www.ncbi.nlm.nih.gov/pubmed/10998179</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/flic_gfp.gif"><img class="aligncenter size-medium wp-image-212" title="fliC_GFP" src="http://uscigem2012.files.wordpress.com/2012/07/flic_gfp.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842013</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>fliC-GFP</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;">When transcribed, <em>RBS-fliC-GFP</em> generates the flagellum protein bonded with a green fluorescent protein. This structure would then be exported to the growing flagella apparatus and used in the construction of the filament. When strongly expressed, this structure would directly result in the flagella filament's containing GFP, allowing for the flagella to be detectable. Unfortunately we were unable to promote the sequence enough to test whether this construct actually functions in this manner.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842013"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842013</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">The <em>fliC</em> gene was cloned via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. Due to the fact that <em>fliC</em> has multiple S and P restriction enzyme sites located within its DNA sequence, it was generated with XbaI sites on the forward and reverse primer. This is so as to allow for the placement of a gene promoter in the E-X sites upstream of the construct in an iGEM biobrick vector. It is important to note though that the current structure consists of the following so as to avoid the accidental loss of a gene during the cloning procedure.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;">EcoRI-XbaI-RBS-<em>fliC</em>-XbaI-GFP-SpeI-PstI</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>fliC</em>:Quickview – EcoliWiki. (n.d.). <em>EcoliWiki</em>. Retrieved October 1, 2012, from http://ecoliwiki.net/colipedia/index.php/fliC</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/flhb1.gif"><img class="aligncenter size-medium wp-image-210" title="flhB" src="http://uscigem2012.files.wordpress.com/2012/07/flhb1.gif?w=214&h=300" alt="" width="214" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842000</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>flhB</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;">Class 2 flagella operon. When transcribed, <em>flhB</em> is one of three flagella genes responsible for the generation of proteins that export flagellin to the flagella apparatus. The flagellin is then directly used in the synthesis of the structure's basal body. Additionally, <em>flhB</em> directly transcribes the operon <em>flhBAE</em>.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842000"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842000</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>flhB</em> was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. It is directly downstream of the class 1 flagella operons <em>flhD</em> and <em>flhC</em>. It also works in conjunction with <em>fliL</em> and <em>fliH</em> to make up the flagellin export system. This part can be used either to generate a flagella in an <em>E. coli</em> strain in which they are not already present, or to promote further formation of pre-existing flagella.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Flagellar biosynthetic protein <em>flhB</em> – Escherichia coli (strain K12). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P76299</span></p> |
| | + | <p><span style="color:#000000;"><em>flhB</em>:Quickview – EcoliWiki. (n.d.). <em>EcoliWiki</em>. Retrieved October 1, 2012, from http://ecoliwiki.net/colipedia/index.php/flhB:Quickview</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td rowspan="3" valign="top" width="377"><span style="color:#000000;"> <a href="http://uscigem2012.files.wordpress.com/2012/07/flhd.gif"><img class="aligncenter size-medium wp-image-221" title="flhD" src="http://uscigem2012.files.wordpress.com/2012/07/flhd.gif?w=300&h=300" alt="" width="300" height="300" /></a></span></td> |
| | + | <td rowspan="3" valign="top" width="120"><span style="color:#000000;"><strong>BBa_K842001</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>flhD</em></span></td> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Description and Function</strong></span></p> |
| | + | <p><span style="color:#000000;"><em>flhD</em> along with <em>flhC</em> form the master regulation operon for the synthesis of flagella in<em> E. coli. </em> They are responsible for the downstream transcription of the class 2 flagella genes. <em>flhD</em> is directly induced by several separate systems; among these is the two-component quorum sensing response system, qseB and qseC.</span></p> |
| | + | <p><span style="color:#000000;">Used for reconstitution of flagella apparatus in <em>E. coli</em> B strains.</span></p> |
| | + | <p><span style="color:#000000;"><a href="http://partsregistry.org/wiki/index.php?title=Part:BBa_K842001"><span style="color:#0000FF;">http://partsregistry.org/wiki/index.php?title=Part:BBa_K842001</span></a></span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Cloning and Uses</strong></span></p> |
| | + | <p><span style="color:#000000;">This part was generated via PCR from the <em>E. coli</em> strain DH5α and cloned into the vector pSB1C3. <em>FlhD</em> is to be used in conjunction with <em>flhC</em> in order to form the transcriptional operon for the production of flagella. This part can be used either to generate a flagella in an <em>E. coli</em> strain in which they are not already present, or to promote further formation of pre-existing flagella.</span></td> |
| | + | </tr> |
| | + | <tr> |
| | + | <td valign="top" width="420"><span style="color:#000000;"><strong>Sources</strong></span></p> |
| | + | <p><span style="color:#000000;">Flagellar transcriptional regulator <em>FlhD</em> – Escherichia coli (strain K12). (n.d.). <em>UniProt</em>. Retrieved October 1, 2012, from http://www.uniprot.org/uniprot/P0A8S9</span></td> |
| | + | </tr> |
| | + | </tbody> |
| | + | </table> |
| | + | <p><strong><span style="color:#000000;">New Constructs Being Tested:</span></strong></p> |
| | + | <p><span style="color:#000000;">Because <em>cheY</em> and <em>cheZ</em> have opposing functions, we wanted to test the expression of both in a single E.coli. To ensure equal transcription from this construct, we utilized the commercially availible pCOLA-Duet and pCDF-Duet vectors from Novagen, which have two RBS sites flanking two unique multiple cloning sites, and are all under the control of a single T7 promoter.</span></p> |
| | + | <p><span style="color:#000000;">These new constructs worked perfectly, and we are currently designing and constructing a similar system in the pSBC1C3 backbone as a new BioBrick.</span></p> |
| | + | <p><span style="color:#000000;">Although we have not included these in the registry yet, as they have not been fully tested, we have generated the following constructs:</span></p> |
| | + | <p><span style="color:#000000;">pCOLA-Duet-<em>CheY</em></span></p> |
| | + | <p><span style="color:#000000;">pCOLA-Duet-<em>CheZ</em></span></p> |
| | + | <p><span style="color:#000000;">pCDF-Duet-<em>CheY</em></span></p> |
| | + | <p><span style="color:#000000;">pCDF-Duet-<em>CheZ</em></span></p> |
| | + | <div id="jp-post-flair" class="sharedaddy sd-like-enabled sd-sharing-enabled"></div> </div><!-- .post-content --> |
| | + | </div><!-- #post-12 --> |
| | | | |
| | + | </body> |
| | + | </html> |
| | + | <center> |
| | <groupparts>iGEM012 USC</groupparts> | | <groupparts>iGEM012 USC</groupparts> |
| | + | </center> |











 "
"
