Team:UC Chile/Cyanolux/Project short
From 2012.igem.org
(Difference between revisions)
Juan Alamos (Talk | contribs) |
|||
| Line 8: | Line 8: | ||
The importance of Biological context in Synthetic Biology has been largely underestimated. We have addressed this issue by centering our project on enhancing functionality of a previously characterized Biobrick, LuxBrick, by placing it in a context which allows new features. | The importance of Biological context in Synthetic Biology has been largely underestimated. We have addressed this issue by centering our project on enhancing functionality of a previously characterized Biobrick, LuxBrick, by placing it in a context which allows new features. | ||
| - | <html><center><img src="https://static.igem.org/mediawiki/2012/3/3f/Rationaleschemechile.jpg" align="middle" width=" | + | <html><center><img src="https://static.igem.org/mediawiki/2012/3/3f/Rationaleschemechile.jpg" align="middle" width="996"></center></html> |
<h3>Bioluminescence</h3> | <h3>Bioluminescence</h3> | ||
| Line 15: | Line 15: | ||
In 2010 the Cambridge iGEM team Biobricked the LuxBrick, a collection of genes from the Lux operon that incorporate both the Luciferase and the substrate production enzymes without regulation, allowing endogenous bioluminescence on E. coli. | In 2010 the Cambridge iGEM team Biobricked the LuxBrick, a collection of genes from the Lux operon that incorporate both the Luciferase and the substrate production enzymes without regulation, allowing endogenous bioluminescence on E. coli. | ||
| - | <html><center><img src="https://static.igem.org/mediawiki/2012/f/fc/Biolumilulilu.jpg" align="middle" width=" | + | <html><center><img src="https://static.igem.org/mediawiki/2012/f/fc/Biolumilulilu.jpg" align="middle" width="720"></center></html> |
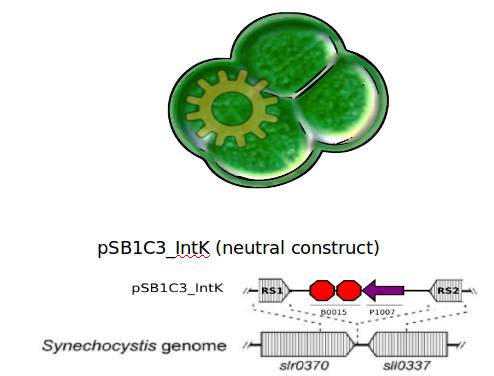
<h3>Chassis</h3> | <h3>Chassis</h3> | ||
| Line 22: | Line 22: | ||
Coupling the endogenous circadian rhythms of this organism to the expression of the Lux genes will enable high-level functionality, through an automatically switching system that turns on bioluminescence only when needed. | Coupling the endogenous circadian rhythms of this organism to the expression of the Lux genes will enable high-level functionality, through an automatically switching system that turns on bioluminescence only when needed. | ||
| - | <html><center><img src="https://static.igem.org/mediawiki/2012/d/da/Synefeatures2.jpg" align="middle" width=" | + | <html><center><img src="https://static.igem.org/mediawiki/2012/d/da/Synefeatures2.jpg" align="middle" width="690"></center></html> |
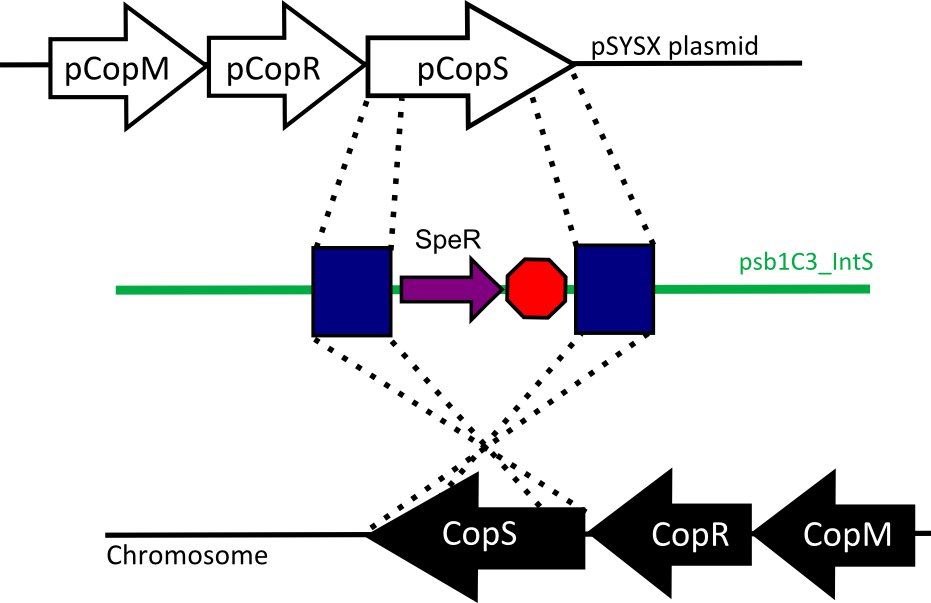
<h2>Strategy</h2> | <h2>Strategy</h2> | ||
According to literature (CITA!), the limiting step for the bacterial bioluminescent reaction is the substrate (n-decanal) concentration, therefore, to control light emission over time we decided to control it´s abundance in the cells, which in our model is a function of the substrates generation (by Lux C, D, E and G enzymes) and consumption (by the LuxAB luciferase). | According to literature (CITA!), the limiting step for the bacterial bioluminescent reaction is the substrate (n-decanal) concentration, therefore, to control light emission over time we decided to control it´s abundance in the cells, which in our model is a function of the substrates generation (by Lux C, D, E and G enzymes) and consumption (by the LuxAB luciferase). | ||
| - | <html><center><img src="https://static.igem.org/mediawiki/2012/f/fc/Dospromotorusi.jpg" align="middle" width=" | + | <html><center><img src="https://static.igem.org/mediawiki/2012/f/fc/Dospromotorusi.jpg" align="middle" width="850"></center></html> |
| Line 37: | Line 37: | ||
It assumes that the metabolite's production is controlled by enzymes under the control of promoter 1 and it´s degradation by enzymes under promoter 2. | It assumes that the metabolite's production is controlled by enzymes under the control of promoter 1 and it´s degradation by enzymes under promoter 2. | ||
For more details please check [2012.igem.org/Team:UC_Chile/Cyanolux/Modelling here] | For more details please check [2012.igem.org/Team:UC_Chile/Cyanolux/Modelling here] | ||
| - | <html><center><img src="https://static.igem.org/mediawiki/2012/d/db/Blackbox.2.jpg" align="middle" width=" | + | <html><center><img src="https://static.igem.org/mediawiki/2012/d/db/Blackbox.2.jpg" align="middle" width="660"></center></html> |
| Line 78: | Line 78: | ||
With the relevance of context in mind, a biomimetic biolamp structure was designed that resembles the organ in which Vibrio fischeri -the bacteria from which the lux genes were biobricked- lives. | With the relevance of context in mind, a biomimetic biolamp structure was designed that resembles the organ in which Vibrio fischeri -the bacteria from which the lux genes were biobricked- lives. | ||
| + | <html><center><img src="https://static.igem.org/mediawiki/2012/b/b3/Biomimetic.jpg" align="middle" width="860"></center></html> | ||
| + | |||
[https://2012.igem.org/Team:UC_Chile/Cyanolux/Biolamp Full description of the device here] | [https://2012.igem.org/Team:UC_Chile/Cyanolux/Biolamp Full description of the device here] | ||
Revision as of 04:02, 26 October 2012
 "
"