Team:TU Darmstadt/Modeling Docking
From 2012.igem.org
(→Results) |
(→Analytics) |
||
| Line 74: | Line 74: | ||
*Flexible Ligand | *Flexible Ligand | ||
===Analytics=== | ===Analytics=== | ||
| - | We extract binding energy from the atomic B-factors and the binding constant from the atomic property derived from all our calculations. We used scatter plots with the binding constant and the inhibition constant in order to compare the plasticizer as well as the PET substrate. | + | We extract binding energy from the atomic B-factors and the binding constant from the atomic property derived from all our calculations. We used scatter plots with the binding constant and the inhibition constant in order to compare the plasticizer as well as the PET substrate.We compared the binding energies as well as the inhibition constant from the best ranked docking runs. Here, OET (dimeric state) became our reference substrate that means if an additive exhibit a higher inhibition constant it may effect our protein. Otherwise, if a binding energies exhibits a higher level than OET we can conclude that an additive may accumulate on our protein surface. Moreover, we compare the binding as well as the inhibition energies of the additives with itself to identify the strongest binder. |
| + | |||
| + | Hence we know the active site of the Fs. Cutinase and the PnB Est. 13 it will be interesting if a plastizicer may docks there. In order to analyze that, we counted the residues which are involved within every docking simulation. Therefore, we measured the distance of every ligand to its corresponding receptor with an appropriate threshold and thus identifying the residues. | ||
===Results=== | ===Results=== | ||
Revision as of 15:07, 23 September 2012
| Homology Modeling | | Gaussian Networks | | Molecular Dynamics | | Information Theory | | Docking Simulation |
|---|
Contents |
Docking
Theory
Goal
While PET.er is degrading PET into its monomers various chemical compounds like polymer plasticizer are released. Since PET is itself a waste material, it will never be in a pure state. Due to this, it is important to study possible interaction of plasticizer as well as PET with the degradation machinery. Hence an experimental design will give a deeper understanding of the interaction of our enzymes with additives.
Methods
Due to the absence of forcefield terms of plasticizer we choose a global docking approach to study the interaction. For the docking simulations we uses the Autodock plugin of Yasara structure. Autodock is one of the ?? mreeost cized and scientific programs for Docking simulations. ??
Docking Protocol
For the docking approach we utilized the standard protocol dock_run.mcr and parallelized it up to six processors. Moreover, we choAse three additives out of the literature acting as the ligand within the simulations.
- OET-dimeric
- S-120
- AO-8
- TPF
Parameter for the Docking Experiment
- Ligands = S120, AO-8, TPF, dimer OET
- Number of docking runs = 200
- ForceField AMBER03
- Flexible Ligand
Analytics
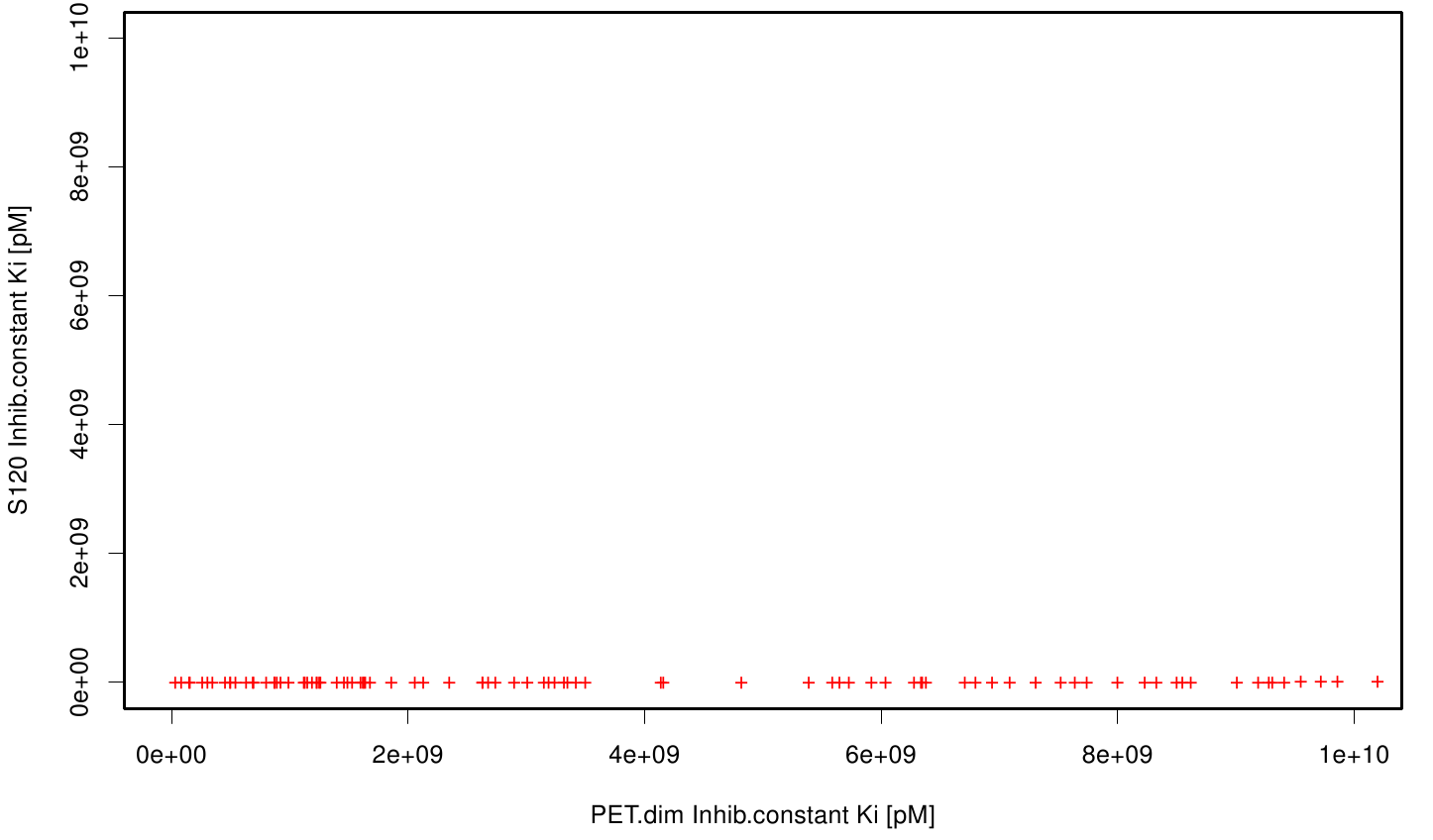
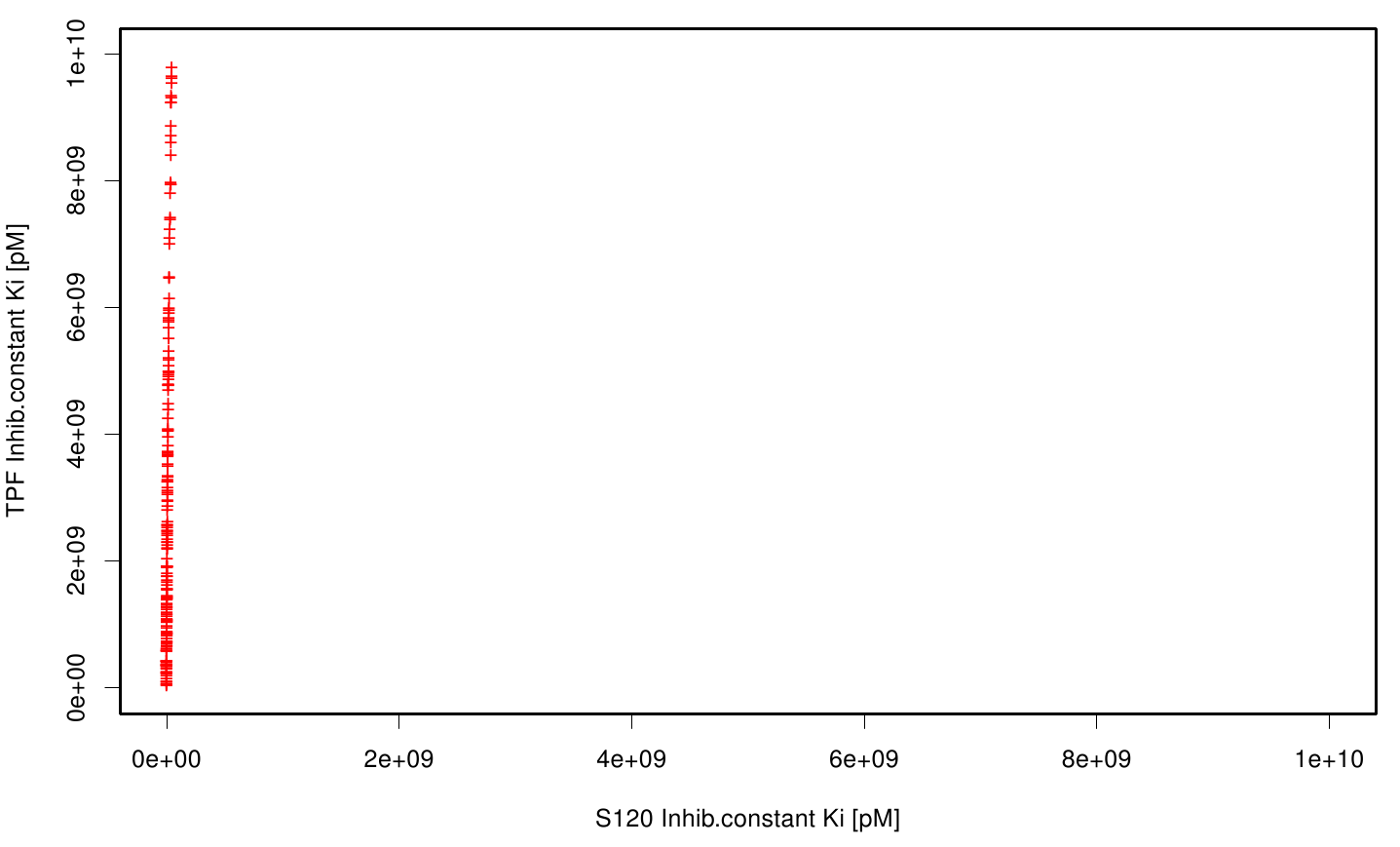
We extract binding energy from the atomic B-factors and the binding constant from the atomic property derived from all our calculations. We used scatter plots with the binding constant and the inhibition constant in order to compare the plasticizer as well as the PET substrate.We compared the binding energies as well as the inhibition constant from the best ranked docking runs. Here, OET (dimeric state) became our reference substrate that means if an additive exhibit a higher inhibition constant it may effect our protein. Otherwise, if a binding energies exhibits a higher level than OET we can conclude that an additive may accumulate on our protein surface. Moreover, we compare the binding as well as the inhibition energies of the additives with itself to identify the strongest binder.
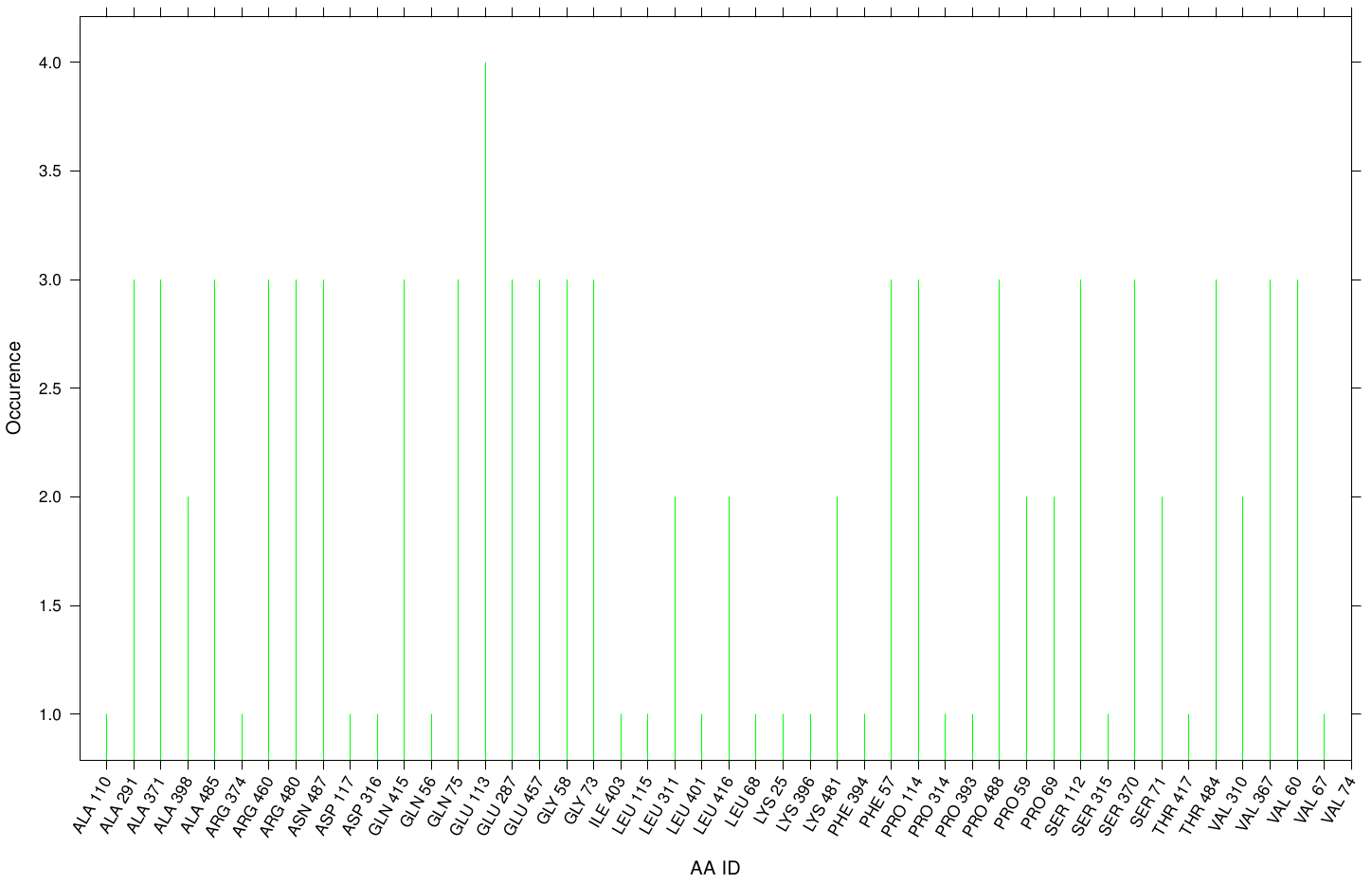
Hence we know the active site of the Fs. Cutinase and the PnB Est. 13 it will be interesting if a plastizicer may docks there. In order to analyze that, we counted the residues which are involved within every docking simulation. Therefore, we measured the distance of every ligand to its corresponding receptor with an appropriate threshold and thus identifying the residues.
 "
"