Team:SJTU-BioX-Shanghai/Project/project3.2
From 2012.igem.org
Huanan1991 (Talk | contribs) |
Huanan1991 (Talk | contribs) (→Switch Modeling - Overview) |
||
| Line 25: | Line 25: | ||
__NOTOC__ | __NOTOC__ | ||
| - | + | =Switch Modeling= | |
| + | {{Template:12SJTU_part_summary_head}} | ||
| + | |||
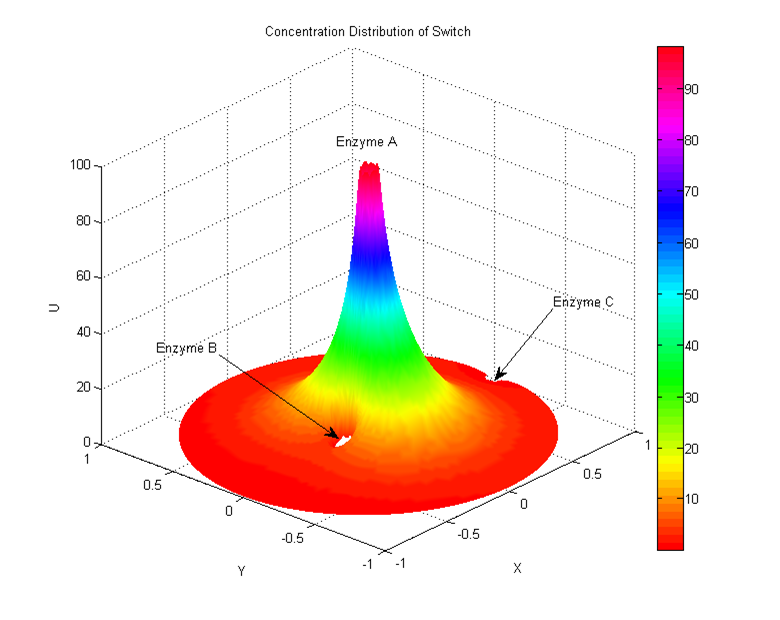
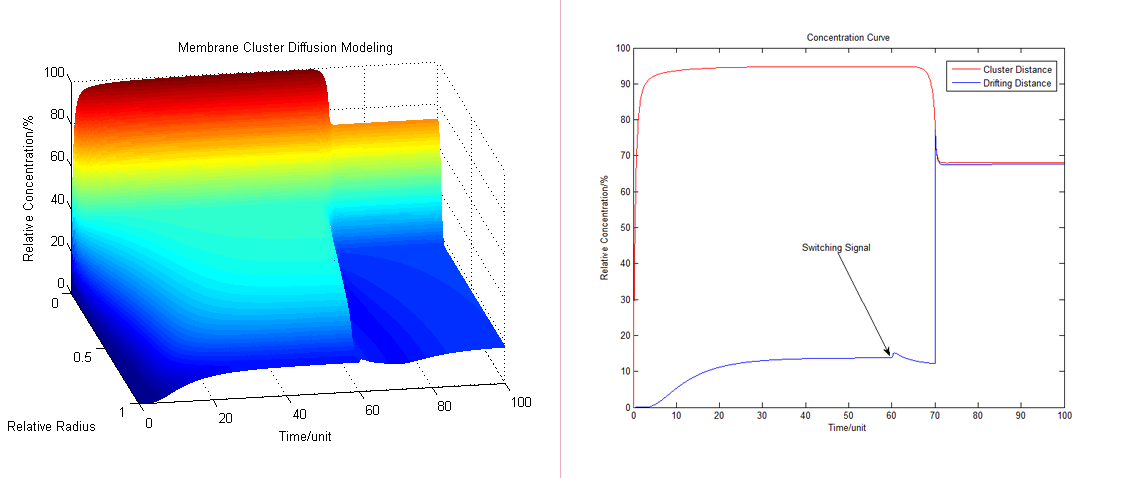
In our system either inner or outer signals can result in a rearrangement of our protein clusters. These interactions activated by signals can change the distances between enzymes, adjust the reaction rate and then finally alter the production amount. We assume that the random diffusion distance on the membrane is about 50-100 times the distance of our cluster interaction radius, and according to PDE model the concentration deceases with radius. | In our system either inner or outer signals can result in a rearrangement of our protein clusters. These interactions activated by signals can change the distances between enzymes, adjust the reaction rate and then finally alter the production amount. We assume that the random diffusion distance on the membrane is about 50-100 times the distance of our cluster interaction radius, and according to PDE model the concentration deceases with radius. | ||
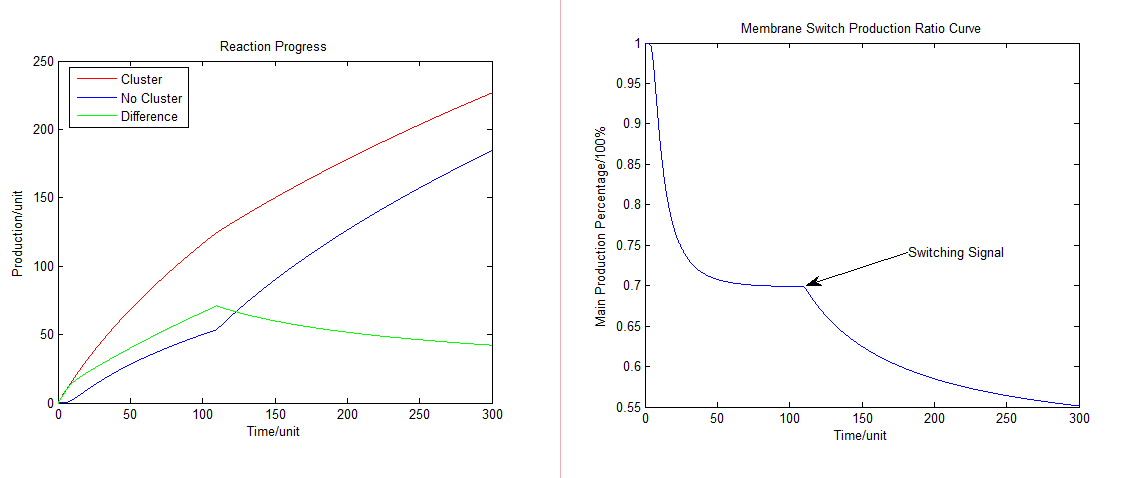
Working with a chain reaction which has different path ways, after getting a signal, the system will soon recruits our membrane protein parts and alters the dynamic equilibrium of products. In other words, alters the amount and ratio of our final products. | Working with a chain reaction which has different path ways, after getting a signal, the system will soon recruits our membrane protein parts and alters the dynamic equilibrium of products. In other words, alters the amount and ratio of our final products. | ||
| + | {{Template:12SJTU_part_summary_foot}} | ||
==Membrane Switch Model== | ==Membrane Switch Model== | ||
Revision as of 17:29, 26 September 2012
| ||
|
 "
"