Team:SJTU-BioX-Shanghai/Project/project2.3
From 2012.igem.org
AleAlejandro (Talk | contribs) (→Desulfurization Pathway) |
AleAlejandro (Talk | contribs) (→Membrane Accelerator - PAH degradation & DBT desulfurization) |
||
| Line 36: | Line 36: | ||
To accelerate degradation of Polycyclic Aromatic Hydrocarbon (PAH) | To accelerate degradation of Polycyclic Aromatic Hydrocarbon (PAH) | ||
| - | To accelerate | + | To accelerate desulfurization of Dibenzothiophene (DBT) |
| Line 46: | Line 46: | ||
{{Template:12SJTU_part_summary_foot}} | {{Template:12SJTU_part_summary_foot}} | ||
| + | ==Biodegradation of Polycyclic Aromatic Hydrocarbon (PAH)== | ||
| - | + | ===Background=== | |
| - | ==Background== | + | |
Polycyclic aromatic hydrocarbons (PAHs), which consist of two or more fused aromatic rings, are widespread in the environment and persist for a very long time. Some PAHs are toxic, mutagenic and carcinogenic and therefore are health hazards. Efforts have been made to screen bacteria strains that could degrade PAH. Moreover, many researchers focused on studying the mechanism of PAH biodegradation. But the rate of natural biodegradation is relatively slow. Team SJTU-BioX-Shanghai is trying to build a Membrane Accelerator to speed up biodegradation rate of PAH biodegradation process. Naphthalene degradation pathway in Pseudomonas species is well studied and thus recruited in our project to build Membrane Accelerator for PAH biodegradation. | Polycyclic aromatic hydrocarbons (PAHs), which consist of two or more fused aromatic rings, are widespread in the environment and persist for a very long time. Some PAHs are toxic, mutagenic and carcinogenic and therefore are health hazards. Efforts have been made to screen bacteria strains that could degrade PAH. Moreover, many researchers focused on studying the mechanism of PAH biodegradation. But the rate of natural biodegradation is relatively slow. Team SJTU-BioX-Shanghai is trying to build a Membrane Accelerator to speed up biodegradation rate of PAH biodegradation process. Naphthalene degradation pathway in Pseudomonas species is well studied and thus recruited in our project to build Membrane Accelerator for PAH biodegradation. | ||
| - | == Degradation Pathway== | + | === Degradation Pathway=== |
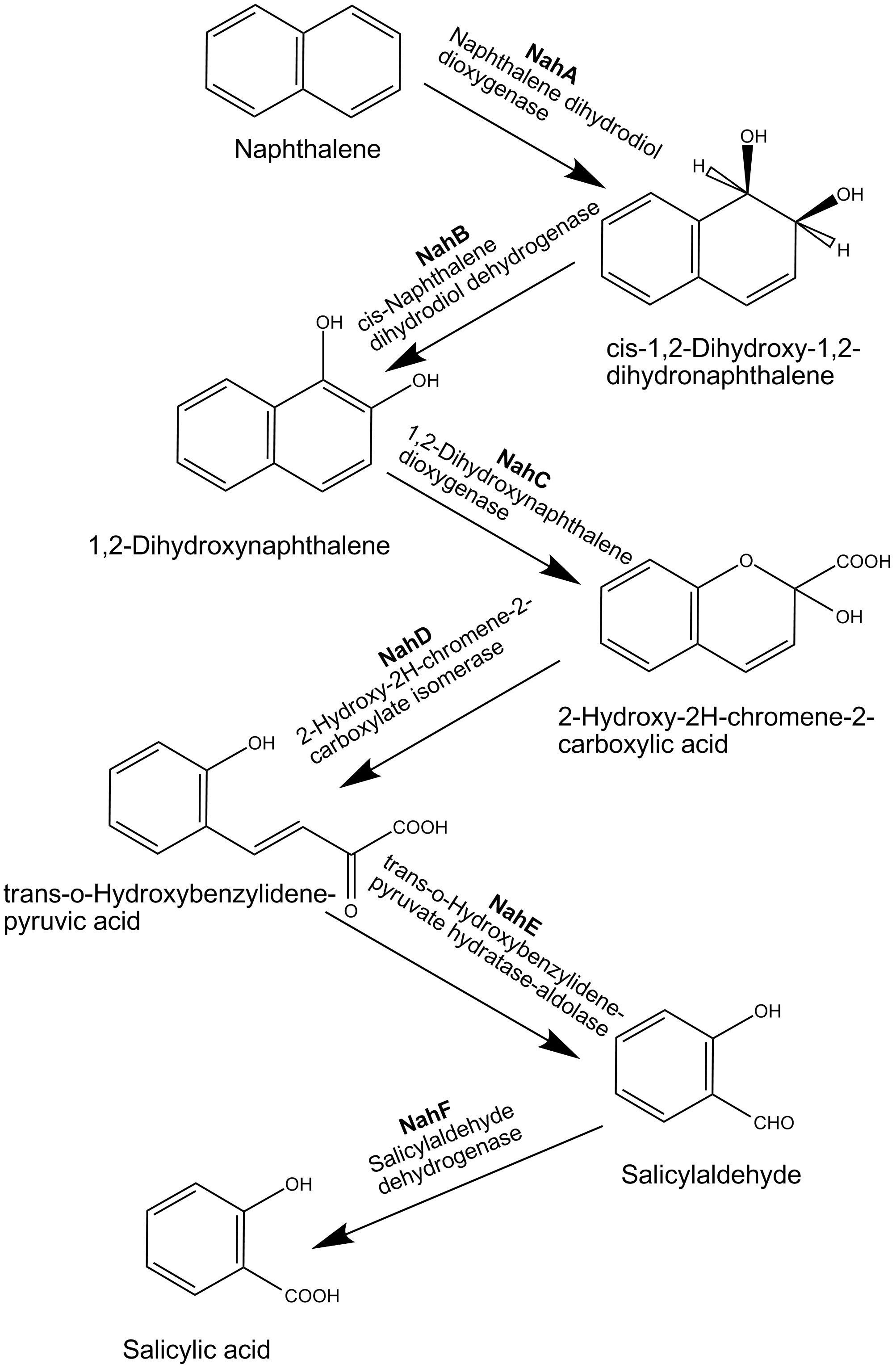
We recruited naphthalene degradation pathway in ''Pseudomonas'' species, which has been well characterized. Six crucial enzymes are involved in naphthalene degradation pathway. | We recruited naphthalene degradation pathway in ''Pseudomonas'' species, which has been well characterized. Six crucial enzymes are involved in naphthalene degradation pathway. | ||
| Line 57: | Line 57: | ||
In the first catabolic step, an oxygen molecule is introduced at the 1,2-position of the aromatic nucleus to produce cis-1,2-dihydroxy-1,2-dihydronaphthalene by naphthalene dihydrodiol dioxygenase. cis-1,2-Dihydroxy-1,2-dihydronaphthalene is then dehydrogenated to 1,2-dihydroxynaphthalene by cis-naphthalene dihydrodiol dehydrogenase. 1,2-Dihydroxynaphthalene is cleaved by 1,2-dihydroxynaphthalene dioxygenase, and the resulting ring-cleavage product spontaneously cyclizes to form 2-hydroxy-2H-chromene-2-carboxylic acid. Enzymatic reactions by an isomerase and a hydratase-aldolase result in the production of salicylaldehyde, which is then transformed to salicylate by salicyladehyde dehydrogenase. | In the first catabolic step, an oxygen molecule is introduced at the 1,2-position of the aromatic nucleus to produce cis-1,2-dihydroxy-1,2-dihydronaphthalene by naphthalene dihydrodiol dioxygenase. cis-1,2-Dihydroxy-1,2-dihydronaphthalene is then dehydrogenated to 1,2-dihydroxynaphthalene by cis-naphthalene dihydrodiol dehydrogenase. 1,2-Dihydroxynaphthalene is cleaved by 1,2-dihydroxynaphthalene dioxygenase, and the resulting ring-cleavage product spontaneously cyclizes to form 2-hydroxy-2H-chromene-2-carboxylic acid. Enzymatic reactions by an isomerase and a hydratase-aldolase result in the production of salicylaldehyde, which is then transformed to salicylate by salicyladehyde dehydrogenase. | ||
| - | == Design== | + | === Design=== |
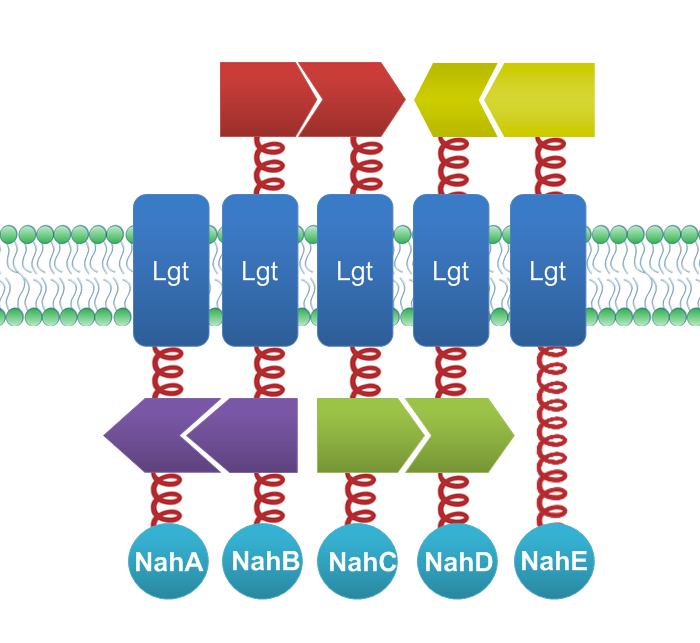
To test whether Membrane Accelerator could accelerate naphthalene biodegradation pathway, we are trying to link six enzymes in this pathway to orderly organized membrane anchor and expressed them in E.coli. E.coli expressing the same type and amount of cytoplasmic enzymes is set as control group. | To test whether Membrane Accelerator could accelerate naphthalene biodegradation pathway, we are trying to link six enzymes in this pathway to orderly organized membrane anchor and expressed them in E.coli. E.coli expressing the same type and amount of cytoplasmic enzymes is set as control group. | ||
| Line 63: | Line 63: | ||
| - | + | ==Biodesulfurization of Dibenzothiophene (DBT) == | |
| - | ==Background== | + | ===Background=== |
We found that many iGEM teams wanted to express enzymes of biodegradation pathway in E.coli. However, it doesn’t have significant superiority compared to traditional strain screening. The power of synthetic biology is not fully demonstrated here. | We found that many iGEM teams wanted to express enzymes of biodegradation pathway in E.coli. However, it doesn’t have significant superiority compared to traditional strain screening. The power of synthetic biology is not fully demonstrated here. | ||
We got inspired by project of 2012 iGEM team Calgary. One of their goals is to achieve desulfuration of dibenzothiophene in E.coli. To improve the system, we aimed to link those enzymes to orderly organized membrane anchors, which would enhance desulfurization pathway remarkably. | We got inspired by project of 2012 iGEM team Calgary. One of their goals is to achieve desulfuration of dibenzothiophene in E.coli. To improve the system, we aimed to link those enzymes to orderly organized membrane anchors, which would enhance desulfurization pathway remarkably. | ||
Sulfur in crude oil could contribute to global warming, acid rain, and various health issues. Biodesulfurization of fossil fuel could help upgrade the quality of fossil fuel and thus the whole environment. | Sulfur in crude oil could contribute to global warming, acid rain, and various health issues. Biodesulfurization of fossil fuel could help upgrade the quality of fossil fuel and thus the whole environment. | ||
| - | ==Desulfurization Pathway== | + | ===Desulfurization Pathway=== |
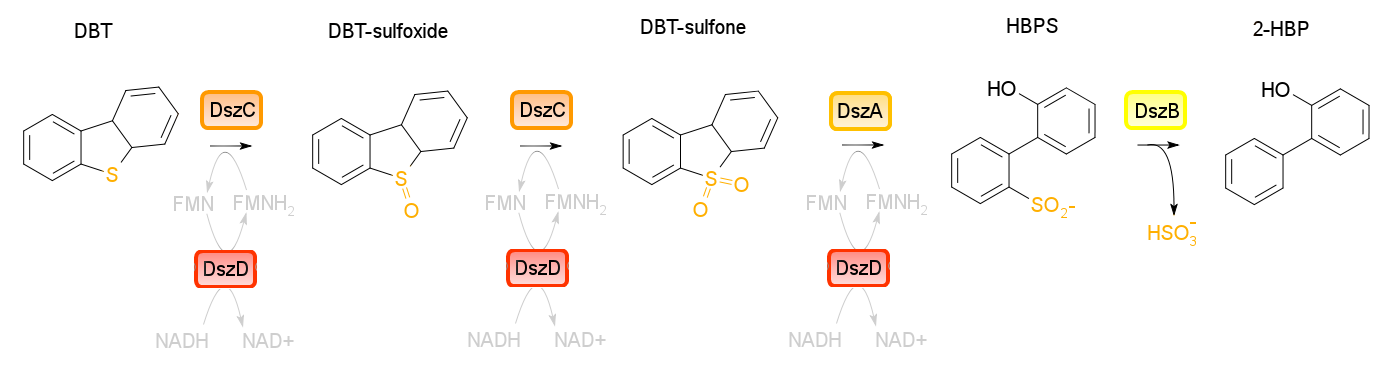
Four enzymes are involved in Desulfurization pathway. Dibenzothiophene monooxygenase (DszC) is responsible for converting DBT to DBT-sulfoxide and finally to DBT-sulfone (DBTO2). DBT-sulfone monooxygenase (DszA) then carries out the next step in the pathway, producing 2-hydroxybiphenyl-2-sulfinic acid (HBPS). HBPS is then converted to the final product by HBPS desulfinase (DszB), producing 2-HBP. The sulfur is released from the hydrocarbon in the form of sulfite. Oxidoreductase (DszD) uses NADH to recycle the FMNH2, allowing the whole reaction to proceed. | Four enzymes are involved in Desulfurization pathway. Dibenzothiophene monooxygenase (DszC) is responsible for converting DBT to DBT-sulfoxide and finally to DBT-sulfone (DBTO2). DBT-sulfone monooxygenase (DszA) then carries out the next step in the pathway, producing 2-hydroxybiphenyl-2-sulfinic acid (HBPS). HBPS is then converted to the final product by HBPS desulfinase (DszB), producing 2-HBP. The sulfur is released from the hydrocarbon in the form of sulfite. Oxidoreductase (DszD) uses NADH to recycle the FMNH2, allowing the whole reaction to proceed. | ||
| Line 77: | Line 77: | ||
For more information, click [https://2012.igem.org/Team:Calgary/Project/OSCAR/Desulfurization Wiki of team Calgary] | For more information, click [https://2012.igem.org/Team:Calgary/Project/OSCAR/Desulfurization Wiki of team Calgary] | ||
| - | == Design== | + | === Design=== |
We are trying to link DszA, DszB, DszC and DszD to orderly organized membrane anchor to accelerate Desulfurization process in E.coli. Due to decreased distance between those enzymes, the proceeding speed could increase sharply. | We are trying to link DszA, DszB, DszC and DszD to orderly organized membrane anchor to accelerate Desulfurization process in E.coli. Due to decreased distance between those enzymes, the proceeding speed could increase sharply. | ||
| - | ==Reference== | + | ==Reference== |
Habe, H. and T. Omori (2003). "Genetics of polycyclic aromatic hydrocarbon metabolism in diverse aerobic bacteria." Bioscience, biotechnology, and biochemistry 67(2): 225-243. | Habe, H. and T. Omori (2003). "Genetics of polycyclic aromatic hydrocarbon metabolism in diverse aerobic bacteria." Bioscience, biotechnology, and biochemistry 67(2): 225-243. | ||
Revision as of 14:40, 22 October 2012
| ||
|
 "
"