Team:SJTU-BioX-Shanghai/Project/project1.2
From 2012.igem.org
AleAlejandro (Talk | contribs) (→Fluorescence Complementation Assay) |
AleAlejandro (Talk | contribs) (→Membrane Accelerator) |
||
| Line 30: | Line 30: | ||
As there is no compartment in prokaryotic cells, enzymes involved in a biochemical pathway diffuse all over the cytoplasm. Intermediates produced from one enzyme cannot be passed efficiently to the next due to spatial obstacles. | As there is no compartment in prokaryotic cells, enzymes involved in a biochemical pathway diffuse all over the cytoplasm. Intermediates produced from one enzyme cannot be passed efficiently to the next due to spatial obstacles. | ||
| - | Rationally designed assemblies of enzymes could help substrates flow by decreasing distance that intermediates have to travel between enzymes, and thus increase yields of sequential biological reactions. If we attach those enzymes to engineered membrane proteins which constitutively assemble together, the reactions are going to proceed much faster. | + | Rationally designed assemblies of enzymes could help substrates flow by decreasing distance that intermediates have to travel between enzymes, and thus increase yields of sequential biological reactions. If we attach those enzymes to engineered membrane proteins which constitutively assemble together, the reactions are going to proceed much faster. Based on this theory, we tried to build a ''Membrane Accelerator'' which organizes enzymes involved in sequential reactions in order on ''E.coli'''s inner membrane. |
{{Template:12SJTU_part_summary_foot}} | {{Template:12SJTU_part_summary_foot}} | ||
| Line 36: | Line 36: | ||
==Construction== | ==Construction== | ||
| - | As demonstrated before, we | + | As demonstrated before, we based our device on ''E.coli'' inner membrane protein Lgt. SRP-dependent signaling sequence of DsbA is fused to N-terminus to localize fusion proteins to membrane. |
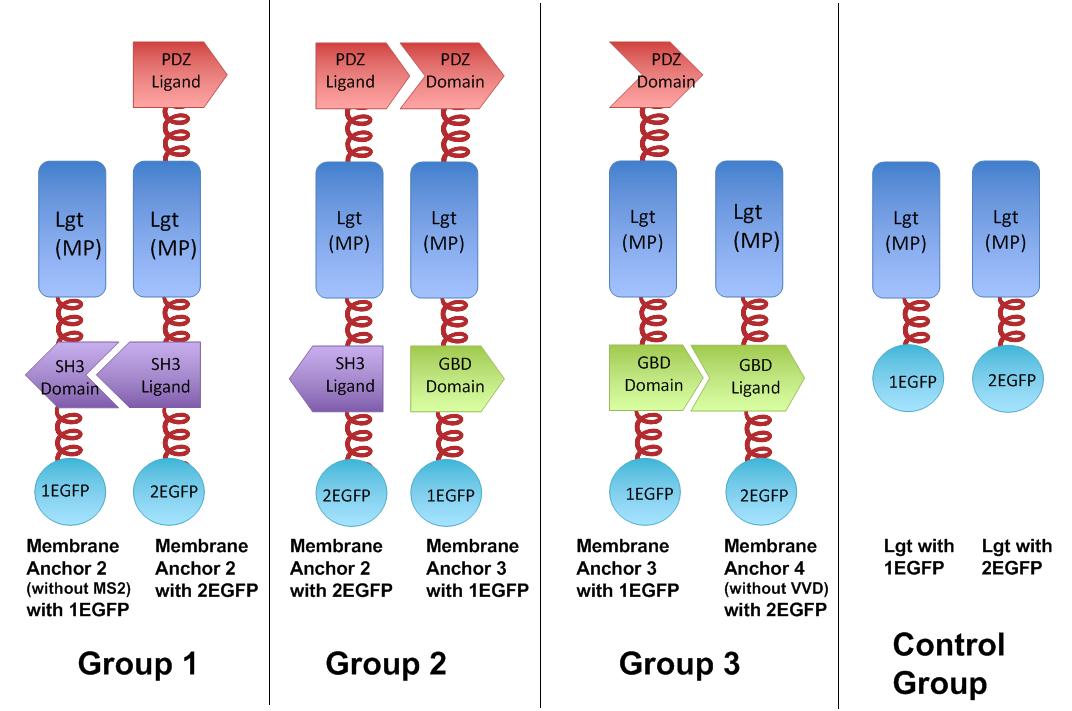
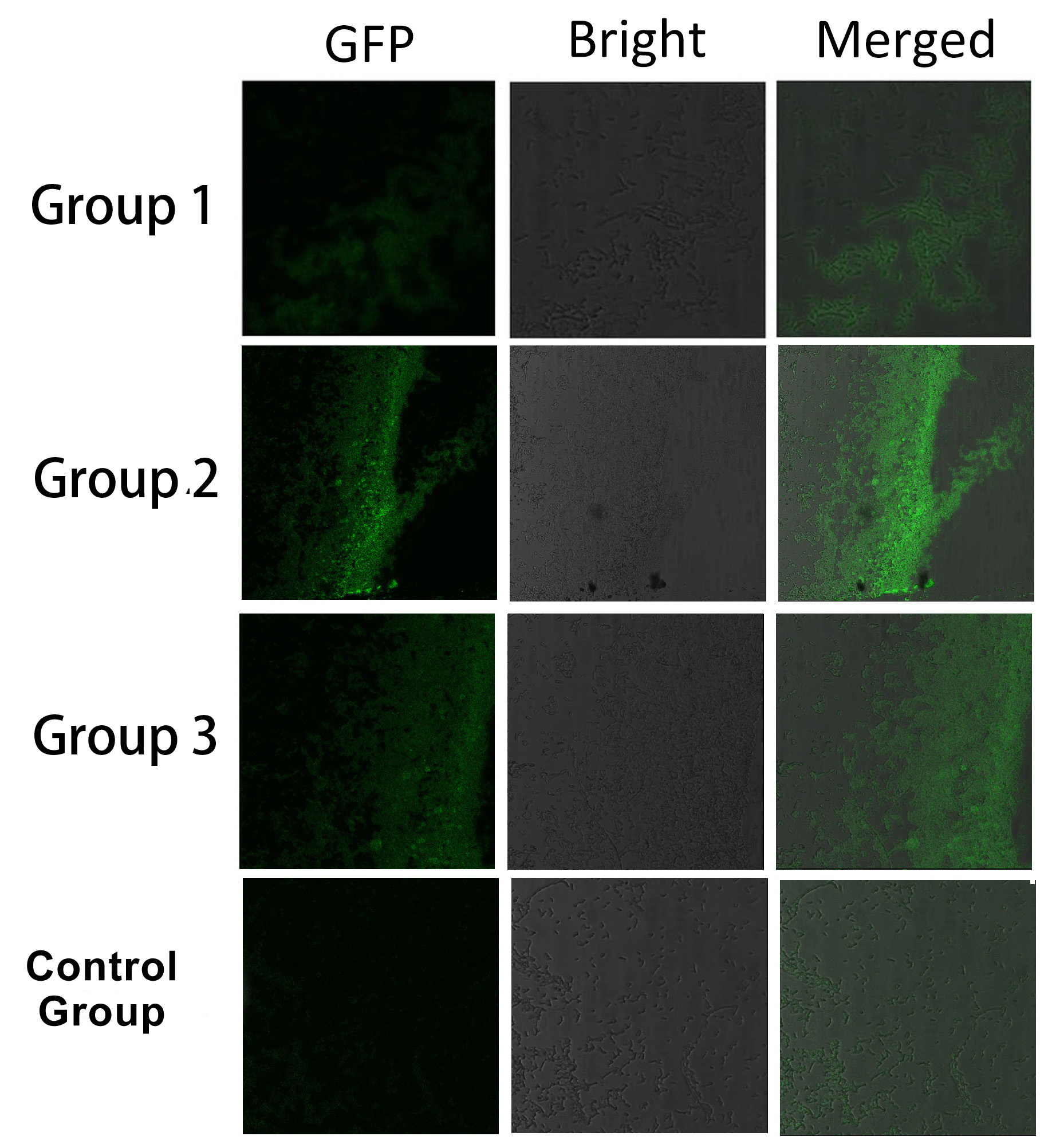
To rationally assemble and arrange enzymes onto inner membrane of ''E.coli'', we incorporate three protein domains that can interact with their cognate ligands. Protein-protein interaction domains and ligands from metazoan cells (mouse SH3 and PDZ domains and rat GBD domain) were recruited to assemble enzymes on membrane. Each domain could assemble with its cognate ligand. | To rationally assemble and arrange enzymes onto inner membrane of ''E.coli'', we incorporate three protein domains that can interact with their cognate ligands. Protein-protein interaction domains and ligands from metazoan cells (mouse SH3 and PDZ domains and rat GBD domain) were recruited to assemble enzymes on membrane. Each domain could assemble with its cognate ligand. | ||
Revision as of 01:28, 24 October 2012
| ||||||||||||||
|
 "
"