Team:NTU-Taida/Project/Stability
From 2012.igem.org
(→1. Multimer Resolution System:Tn1000(gamma delta) resolution system) |
(→2. Partition System:from Pseudomonas putida KT2440) |
||
| Line 28: | Line 28: | ||
====2. Partition System:from '''<em>Pseudomonas putida''' KT2440</em>==== | ====2. Partition System:from '''<em>Pseudomonas putida''' KT2440</em>==== | ||
| - | : | + | :Bacterium distributes chromosomes/plasmids evenly to its progenies by a dynamic system that includes SMC like proteins, type Ia partition system and so on[[#Ref6|[6] ]]]]. Type Ia partition system segregates chromosomes/plasmids in a process akin to Eukaryotic mitosis. It can be found on most eubacteria. However, E. coli is not the case. |
[[FIle:NTU-Taida-Par2.png|700px|center]] | [[FIle:NTU-Taida-Par2.png|700px|center]] | ||
Revision as of 02:54, 27 September 2012
Stability
Every GM system that functions outside the laboratories will face two major problems: system stability and safety! Without selective pressure, we have to deal with plasmid instability; to make our ''E. coli'' colonized bowel we have to use recA+ strains which may cause plasmid multimer. Our genetically modified ''E. coli'' will also contact with many kinds of bacteria and is under the risk of horizontal gene transfer.The following segment is our struggle against these obstacles.
Contents |
Stability of Delivery System
Briefing
As our PepdEx system consists of several plasmids and will function outside of the laboratory (human gut) which lacks of antibiotic selection pressure, plasmid segregation stability is critical that determines whether every single E. Coli cell contains the original system with the designated function. Inspired by natural plasmid & mobile gene element, we cope with segregation instability by incorporating three modules, i.e. partition system, Multimer resolution system and toxin antitoxin system, on top of our system. We think that this layer of regulation will potentially benefit the entire synthetic biology community, as synthetic systems are becoming increasingly complex with fine controls (e.g. on low copy plasmids).
Obstacles
Plasmid instability problems come majorly from the segregational instability,the burden effect and the dimer catastrophe.
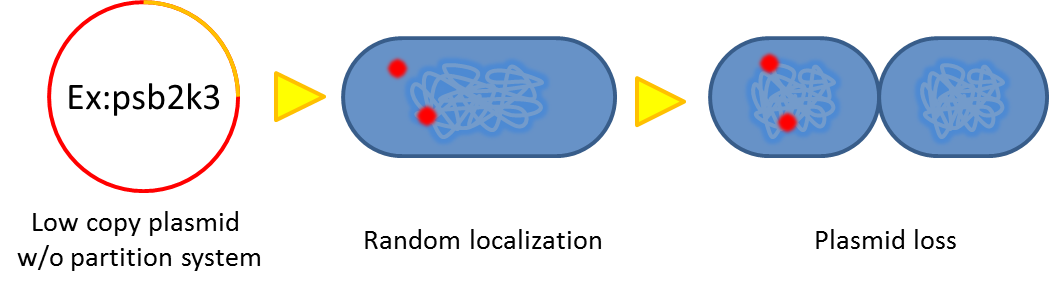
- Segregational instability
Plasmids are randomly (possibly uneven) distributed inside a bacterium. After cell division, one of the progenies may lose plasmids.
- Burden Effect
Cells bearing our PepdEx system have to spend extra energy/resource, so will not grow as good as plasmid-free cells. This metabolic burden will cause our coli to gradually lose pieces of circuits, and the growth rate difference will eventually eliminate plasmid-bearing bacteria in the population.
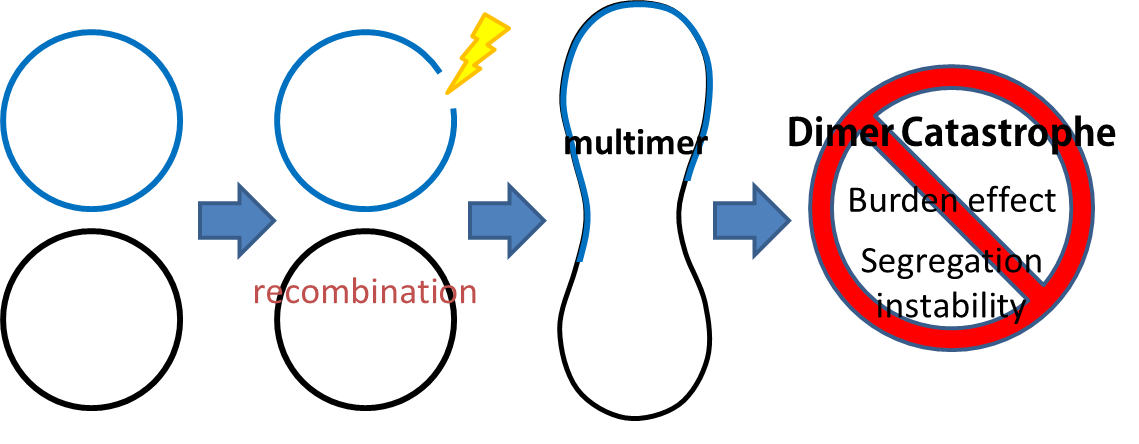
- Dimer Catastrophe
Homologues recombination may cause plasmid multimers which will increases segregational instability and burden[1] . This is the reason why most lab coli are recA1 mutants. But since our coli must be able to colonize in the gut, it should be recA+ wild type strain. That's the problem!
Stabilization Modules
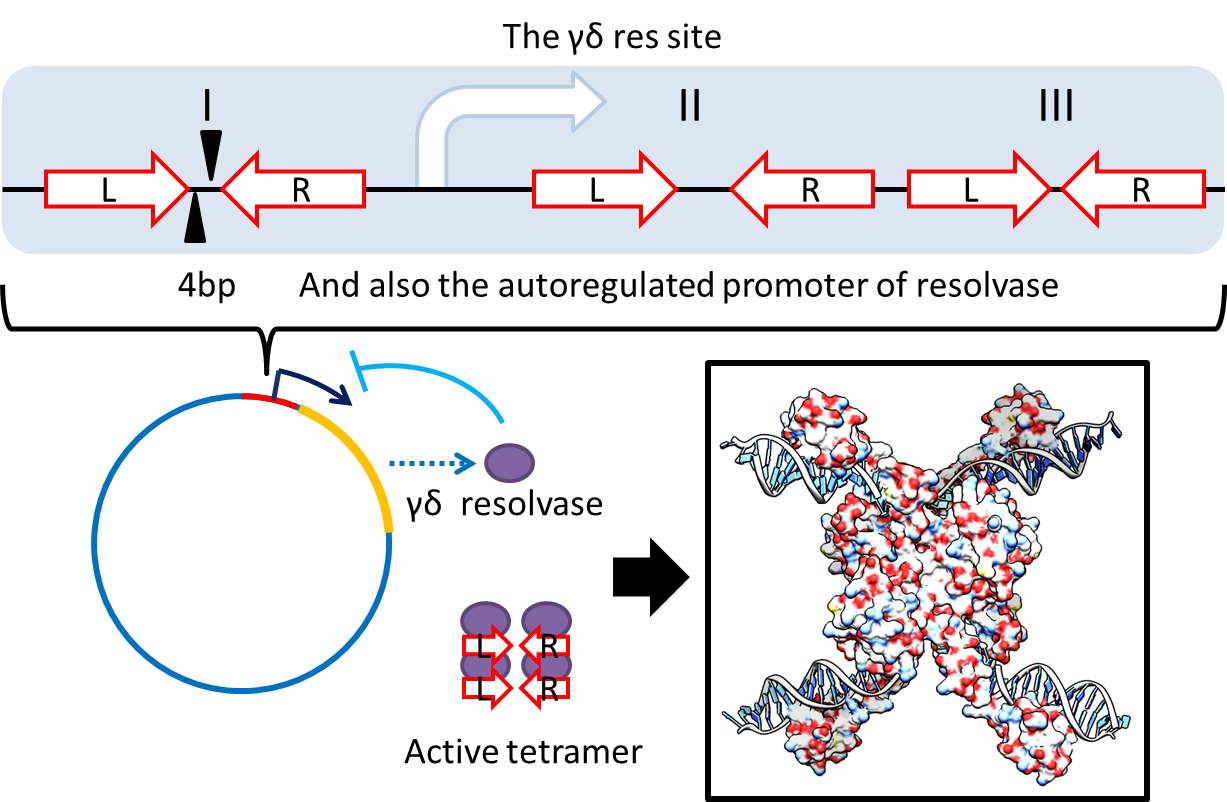
1. Multimer Resolution System:Tn1000(gamma delta) resolution system
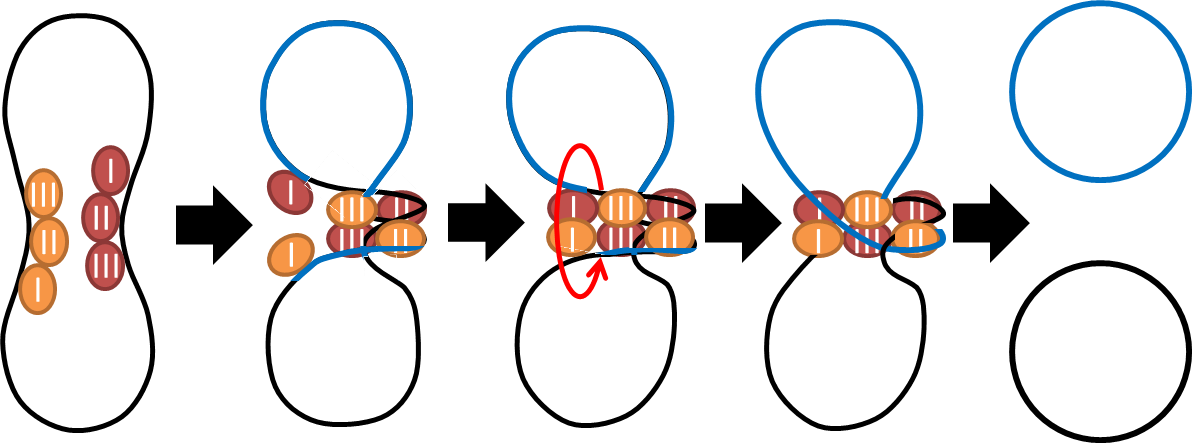
- To deal with multimerization, we cloned an cassette (BBa_K817018) which encodes an autoregulated resolves (serine type recombinase) from E.coli F plasmid transposon tn1000 (tn3 family)[2] . Its promoter region consists of 3 sub-sites(res site) and can process recombination[3] . This cassette can resolve multimer that formed during replicative transposition[4] . It can also resolve plasmid multimer to increase plasmid stability by avoiding dimer catastrophe. Multimer resolution system (MRS) provides analogous functions to E.coli chromosome XerCD/dif but acts independent of cell cycle, DNA localization and may have higher efficiency on plasmids compared with slow XerCD system[5] .
2. Partition System:from Pseudomonas putida KT2440
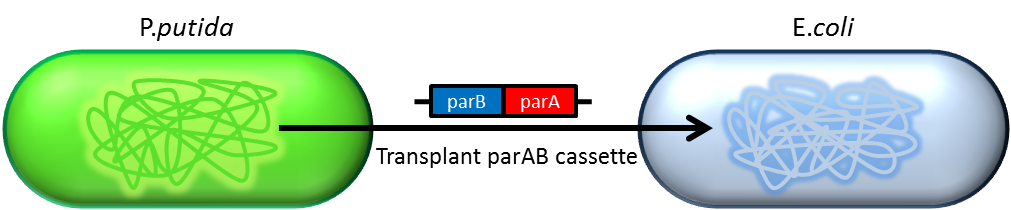
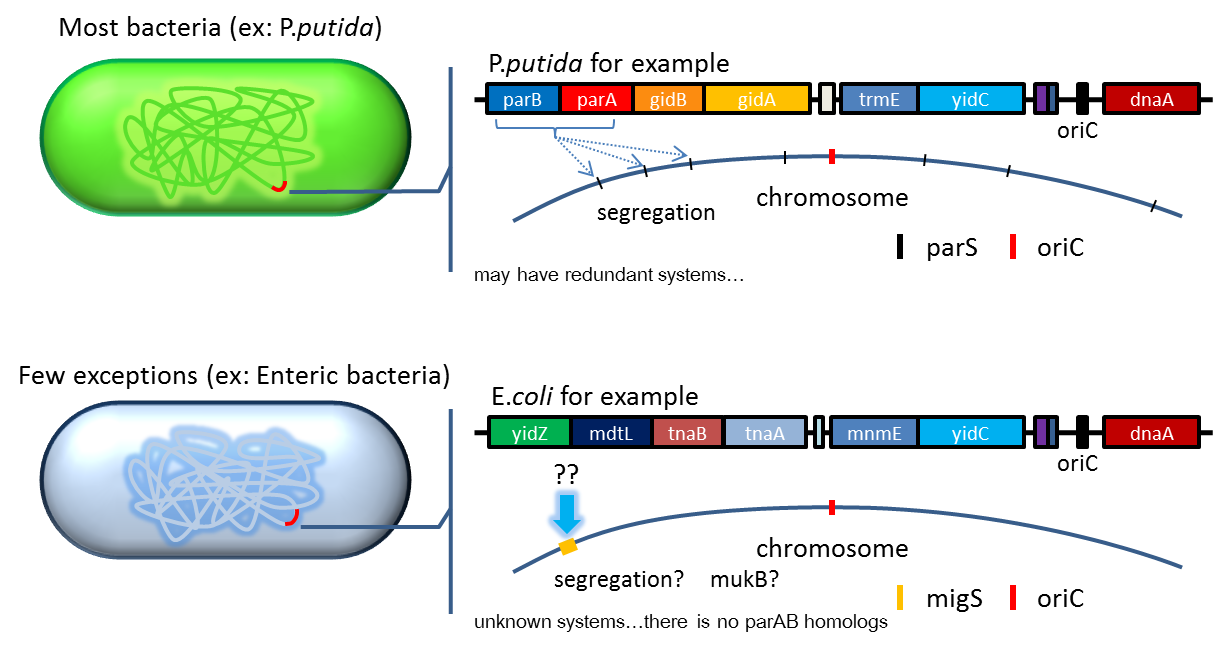
- Bacterium distributes chromosomes/plasmids evenly to its progenies by a dynamic system that includes SMC like proteins, type Ia partition system and so on[6] ]]. Type Ia partition system segregates chromosomes/plasmids in a process akin to Eukaryotic mitosis. It can be found on most eubacteria. However, E. coli is not the case.
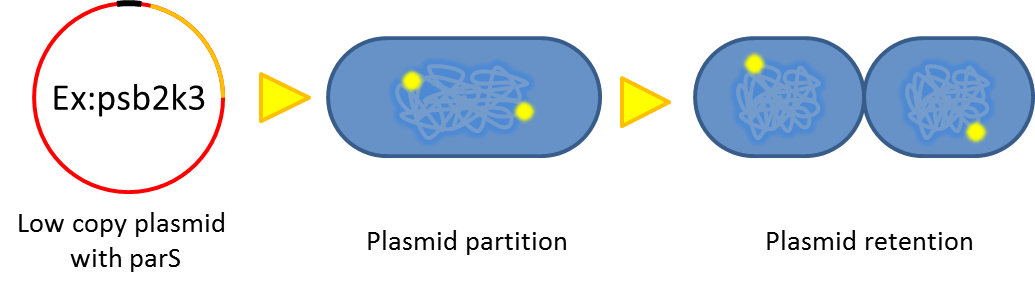
- Previous study have shown that when provide parAB of P.putida in trans, the low copy plasmid(mini F)carrying conserved parS site can be stabilized in E.coli[7] ]].(Anne-Marie etl. 2002)pSB2K3 is mini F plasmid thus can be stabilized by this way too, we make a new version of pSB2K3 with parS site. This plasmid that have lower gene dosage(copy number)and can be partitioned is an ideal vector to harbor our PepdEX system.
3. Post-Segregation Killing:srnBC toxin-antitoxin system
- No matter how well partition system and multimer resolution system work, inevitably there still will have some bacteria losing plasmid. We sentence them to death to solve the problem.

| 
|
- Type I toxin-antitoxin srnBC is an ideal executor which belongs to hok/sok homologues.[8] It expresses stable toxin encoded RNA and short lived antitoxin RNA that can neutralize toxin RNA by RNA interaction and RNase III cleavage[9] . It acts as post segregation killing system, which kills bacteria when it loses the DNA (genomic islands[10] , plasmids, mobile gene elements) that contains it. Therefore we use it to reduce plasmid loss rate to make applications without antibiotic selection more feasible. Following pictures show the mechanism (based on hok/sok).
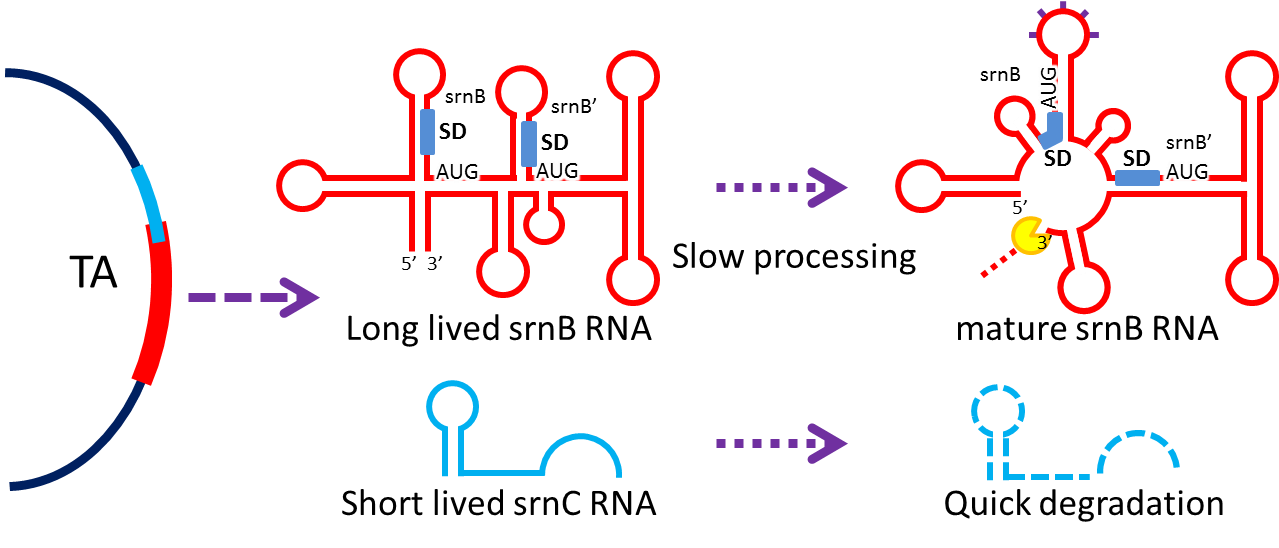
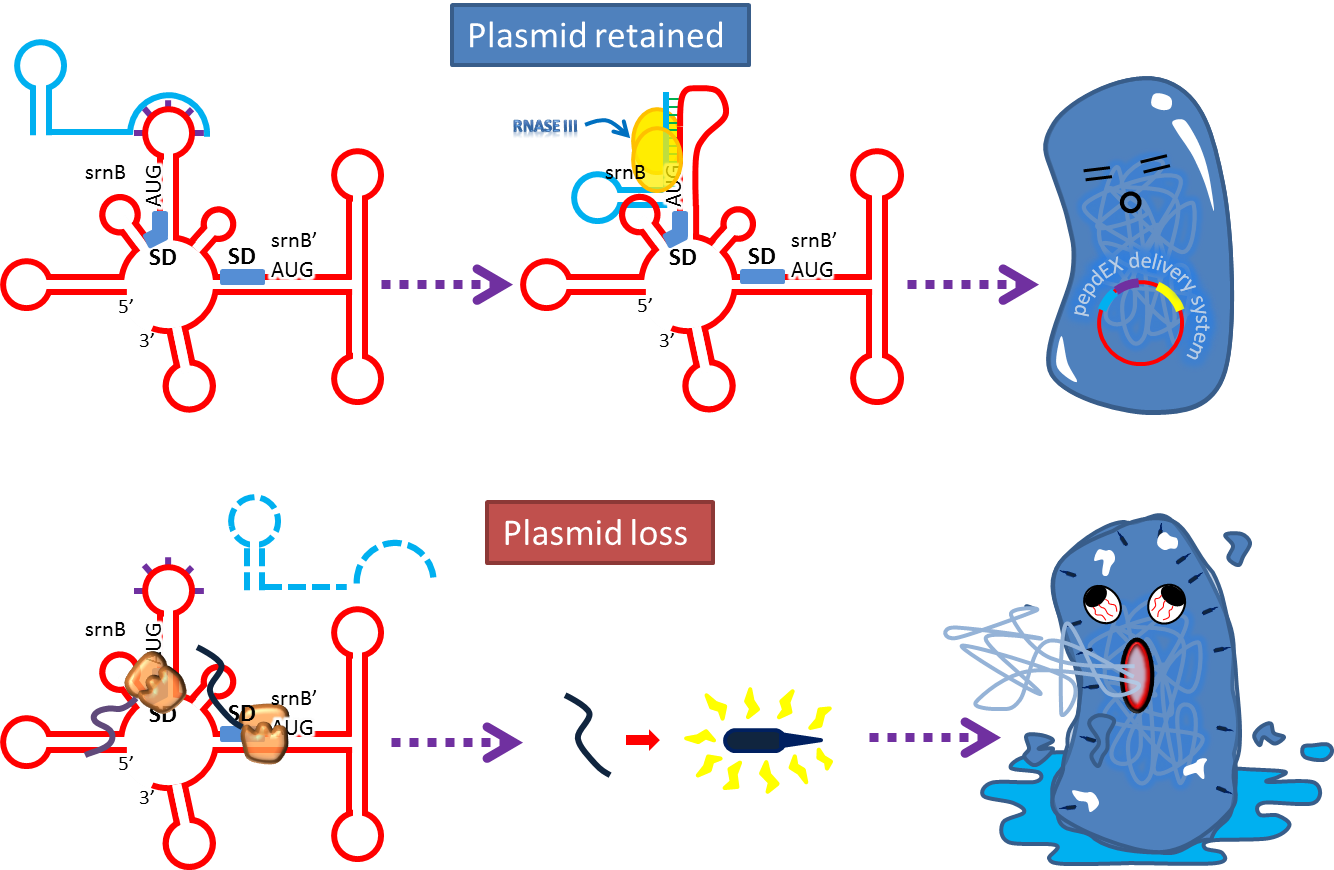
4. Reduce Burden Effect
- Besides these modules, reducing the burden effect is also important to the system stability. Well-designed system and lower gene dosage may help.
- High level gene expression and gene dosage cause burden effect. If having the same outcome, low copy plasmid is preferred than high copy ones. If having the same outcome and not for regulatory purpose, stabilizing mRNA is preferred than overexpression of it. Place yourselves in E. coli's position, reduce its metabolic burden as much as possible then it can work for you.
Reference
- Field, C.M. and D.K. Summers, Multicopy plasmid stability: revisiting the dimer catastrophe. Journal of Theoretical Biology, 2011. 291: p. 119-27.
- Falvey, E. and N.D.F. Grindley, Contacts between Gamma-Delta-Resolvase and the Gamma-Delta-Res Site. Embo Journal, 1987. 6(3): p. 815-821.
- Grindley, N.D.F., et al., Transposon-Mediated Site-Specific Recombination - Identification of 3 Binding-Sites for Resolvase at the Res-Sites of Gamma-Delta and Tn3. Cell, 1982. 30(1): p. 19-27.
- Wells, R.G. and N.D.F. Grindley, Analysis of the Gamma-Delta Res Site - Sites Required for Site-Specific Recombination and Gene-Expression. Journal of Molecular Biology, 1984. 179(4): p. 667-687.
- Guhathakurta, A., I. Viney, and D. Summers, Accessory proteins impose site selectivity during ColE1 dimer resolution. Molecular Microbiology, 1996. 20(3): p. 613-620.
- Mullins, R.D., Bacterial Chromosome Segregation. Annu Rev Cell Dev Biol, 2009.
- Godfrin-Estevenon, A.M., F. Pasta, and D. Lane, The parAB gene products of Pseudomonas putida exhibit partition activity in both P. putida and Escherichia coli. Molecular Microbiology, 2002. 43(1): p. 39-49.
- Gerdes, K., et al., Mechanism of killer gene activation. Antisense RNA-dependent RNase III cleavage ensures rapid turn-over of the stable hok, srnB and pndA effector messenger RNAs. Journal of Molecular Biology, 1992. 226(3): p. 637-49.
- Thisted, T. and K. Gerdes, Mechanism of post-segregational killing by the hok/sok system of plasmid R1. Sok antisense RNA regulates hok gene expression indirectly through the overlapping mok gene. Journal of Molecular Biology, 1992. 223(1): p. 41-54.
- Szekeres, S., et al., Chromosomal toxin-antitoxin loci can diminish large-scale genome reductions in the absence of selection. Molecular Microbiology, 2007. 63(6): p. 1588-1605.
- Faridani, O.R., et al., Competitive inhibition of natural antisense Sok-RNA interactions activates Hok-mediated cell killing in Escherichia coli. Nucleic Acids Research, 2006. 34(20): p. 5915-5922.
- Gerdes, K. and E.G.H. Wagner, RNA antitoxins. Current Opinion in Microbiology, 2007. 10(2): p. 117-124.
 "
"