Team:NRP-UEA-Norwich/Week3
From 2012.igem.org
| Line 13: | Line 13: | ||
===Meetings=== | ===Meetings=== | ||
| - | Amy (our artist) came to visit us today in order to discuss our video. She, Joy and Rachel began to storyboard the film and work out how to incorporate the decision made over the weekend into the video. They came up with some fantastic ideas and the whole team sat down to discuss them before Amy left to organise making props. Joy also got in touch with the UEA Drama School in order to see whether we could hire a studio in order to film our video. | + | . Amy (our artist) came to visit us today in order to discuss our video. She, Joy and Rachel began to storyboard the film and work out how to incorporate the decision made over the weekend into the video. They came up with some fantastic ideas and the whole team sat down to discuss them before Amy left to organise making props. Joy also got in touch with the UEA Drama School in order to see whether we could hire a studio in order to film our video. |
===Labs=== | ===Labs=== | ||
| - | From the transformation of the two promoters carried out on Friday, we inoculated some colonies into culture. This will be used later for characterisation of the promoter. From the two plates, we noticed that one plate had bigger colonies and one had very small colonies. The larger colonies were from the agar plate inoculated with transformed "E.coli" with promoter M-B. | + | . From the transformation of the two promoters carried out on Friday, we inoculated some colonies into culture. This will be used later for characterisation of the promoter. From the two plates, we noticed that one plate had bigger colonies and one had very small colonies. The larger colonies were from the agar plate inoculated with transformed "E.coli" with promoter M-B. |
| - | To characterise the promoters and other Bio-Bricks we may be interested in, an input of NO will be needed. Our solution to this is to use a nitrate salt like potassium nitrate. A stock solution (1M) was made up today by Lukas. This was taken to be autoclaved. | + | . To characterise the promoters and other Bio-Bricks we may be interested in, an input of NO will be needed. Our solution to this is to use a nitrate salt like potassium nitrate. A stock solution (1M) was made up today by Lukas. This was taken to be autoclaved. |
==Day 2 (24/07/12)== | ==Day 2 (24/07/12)== | ||
===Labs=== | ===Labs=== | ||
| - | Joy prepared the pSB1C3 backbone by restriction. This gave them sticky ends which allows us to ligate DNA with complimentary ends into them. Lukas and Joy also prepared the synthesised hybrid promoters of both orientations by isolating the plasmids. | + | . Joy prepared the pSB1C3 backbone by restriction. This gave them sticky ends which allows us to ligate DNA with complimentary ends into them. Lukas and Joy also prepared the synthesised hybrid promoters of both orientations by isolating the plasmids. |
==Day 3 (25/07/12)== | ==Day 3 (25/07/12)== | ||
| Line 30: | Line 30: | ||
===Research=== | ===Research=== | ||
| - | After our hunt for the fluorometers and speaking to Dr. Yeoman the previous day we decided that we needed to find whether we could use fluorescent proteins today in order to go on to design the DNA for the control region and reporter/enzyme. After a social media call-out for fluorometers in our faculty we were pointed in the direction of Dr. Tom Clarke who very kindly offered us the use of his fluorometer, as well as training on it and potentially even the use of a fluorescence microscope. He also gave us advice on the project and suggested that pigments and the use of a spectrophotometer would perhaps give the better quantitative results and informed us that fluorescence may be particularly labour-intensive and time consuming. We decided to spend the rest of the day researching other ways of reporting on the substrate levels in the environment. | + | . After our hunt for the fluorometers and speaking to Dr. Yeoman the previous day we decided that we needed to find whether we could use fluorescent proteins today in order to go on to design the DNA for the control region and reporter/enzyme. After a social media call-out for fluorometers in our faculty we were pointed in the direction of Dr. Tom Clarke who very kindly offered us the use of his fluorometer, as well as training on it and potentially even the use of a fluorescence microscope. He also gave us advice on the project and suggested that pigments and the use of a spectrophotometer would perhaps give the better quantitative results and informed us that fluorescence may be particularly labour-intensive and time consuming. We decided to spend the rest of the day researching other ways of reporting on the substrate levels in the environment. |
| - | We began by investigating various pigments and looked at projects such as E. chromi to find biobricks of the pigments we would require. One problem we fell into was the absorbance of the pigments overlapping, which would ultimately cause our results to be inaccurate if they were produced in an environment. We then decided to look at proteins with unique absorbancies as our reporters, as well as enzymes that would produce colourful products. Our main problems involved using proteins that were not already incorporated into biobricks, using proteins whose absorbance and emission spectra did not overlap, and using proteins that would not interfere with the ribosome binding site and | + | . We began by investigating various pigments and looked at projects such as E. chromi to find biobricks of the pigments we would require. One problem we fell into was the absorbance of the pigments overlapping, which would ultimately cause our results to be inaccurate if they were produced in an environment. We then decided to look at proteins with unique absorbancies as our reporters, as well as enzymes that would produce colourful products. Our main problems involved using proteins that were not already incorporated into biobricks, using proteins whose absorbance and emission spectra did not overlap, and using proteins that would not interfere with the ribosome binding site and |
===Labs=== | ===Labs=== | ||
| - | Hybrid promoter M-B, was miniprepped. This involved isolation of the plasmid and running on a 1.4% agarose gel. The desired fragment was excised from the gel. | + | . Hybrid promoter M-B, was miniprepped. This involved isolation of the plasmid and running on a 1.4% agarose gel. The desired fragment was excised from the gel. |
| - | To characterise the PyeaR | + | . To characterise the PyeaR Bio-Brick, a culture mixture was prepared containing potassium nitrate, PyeaR-GFP composite, chloramphenicol and LB media. This was left overnight to grow. The isolated hybrid promoter M-B was digested and an 1.4% agarose gel was prepared to separate the fragments the following morning. |
| Line 45: | Line 45: | ||
===Research=== | ===Research=== | ||
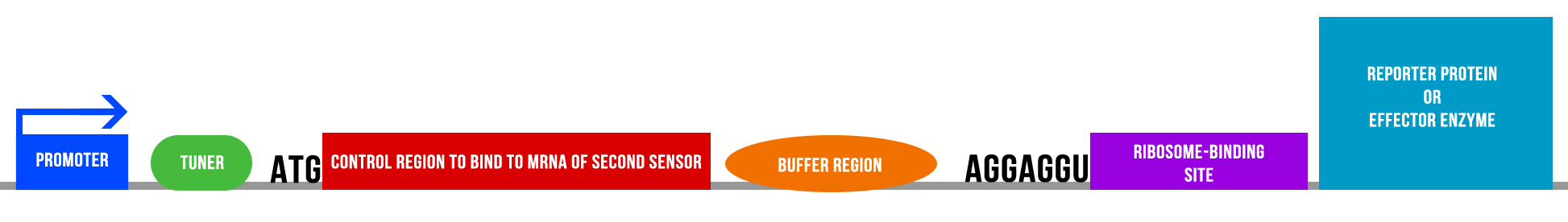
| - | The team have been going down all avenues to search for which reporter would be the best for our project, including fluorescence, absorbing proteins, pigments and enzymes with coloured produce. A decision was made to test fluorescent | + | . The team have been going down all avenues to search for which reporter would be the best for our project, including fluorescence, absorbing proteins, pigments and enzymes with coloured produce. A decision was made to test fluorescent BioBricks we already had to determine how easy they would be to use. Using IDTs OligoAnalyser, we also began to look at how to construct the synthetic gene needed for our comparator circuit idea. It soon became clear that the previous designs for the gene would be too complicated for the timescale of the project, so we decided that our efforts should be focused on designing 'zips' that engulf the ribosome binding site of the reporter gene of interest. These 'zips' on the two constructs will complimentary bind together when both present in the cell and form a duplex that will sequester the ribosome binding site and inhibit translation. |
However, these 'zips' need to consist of non-rare codons on both constructs and still display complimentary base pairing to one another. This is a challenge we hope to overcome over the coming weeks and have the synthetic gene sent of in good time so that, when it arrives, we have enough time to characterise it fully and convert it into a biobrick. | However, these 'zips' need to consist of non-rare codons on both constructs and still display complimentary base pairing to one another. This is a challenge we hope to overcome over the coming weeks and have the synthetic gene sent of in good time so that, when it arrives, we have enough time to characterise it fully and convert it into a biobrick. | ||
| Line 55: | Line 55: | ||
. We tried to purify M-B using the Bioline kit but both ourselves and an independent lab person retrieved no DNA. We suspect that either we are misinterpreting the instructions or the kit may be faulty at low concentrations of DNA. | . We tried to purify M-B using the Bioline kit but both ourselves and an independent lab person retrieved no DNA. We suspect that either we are misinterpreting the instructions or the kit may be faulty at low concentrations of DNA. | ||
| - | Using the nitrate cultures made the previous day, Russell, spun down a proportion of the cells and viewed them under UV light. He found a graduated intensity of green fluorescence. Rebecca prepared chloramphenicol plates at a concentration of 25µg/ml and also made a stock solution of ampicillin. Lukas and Joy ran the gel and found that there was no DNA. We suspect that during the process of isolation, the DNA was lost and the remaining was insufficient to show up on the gel. In the afternoon, Lukas purified the hybrid promoter B-M using gel electrophoresis and DNA was found. This was digested and again Joy prepared an agarose gel for following morning use. | + | . Using the nitrate cultures made the previous day, Russell, spun down a proportion of the cells and viewed them under UV light. He found a graduated intensity of green fluorescence. Rebecca prepared chloramphenicol plates at a concentration of 25µg/ml and also made a stock solution of ampicillin. Lukas and Joy ran the gel and found that there was no DNA. We suspect that during the process of isolation, the DNA was lost and the remaining was insufficient to show up on the gel. In the afternoon, Lukas purified the hybrid promoter B-M using gel electrophoresis and DNA was found. This was digested and again Joy prepared an agarose gel for following morning use. |
==Day 5 (27/07/12)== | ==Day 5 (27/07/12)== | ||
| Line 61: | Line 61: | ||
===Research=== | ===Research=== | ||
| - | Pascoe and Khadija continued their work on planning the comparator circuit construct. They had a limited number of codons to work with, as they needed to be both common codons in ''E. coli'' and have complimentary codons that are also common, and both codons needed to code for small, non polar amino acids that would not affect the reporter proteins tertiary structure enough to affect its reporter properties. The list of codons were collated into a list affectionately called 'The Good List' and from this Pascoe and Khadija began piecing them together and, using OligoAnalyser, analysing the folding of the constructs on their own. If the ribosome binding site or start codon were obstructed in the single construct strands, translation could be inhibit, which would hinder the comparator circuits functionality. | + | . Pascoe and Khadija continued their work on planning the comparator circuit construct. They had a limited number of codons to work with, as they needed to be both common codons in ''E. coli'' and have complimentary codons that are also common, and both codons needed to code for small, non polar amino acids that would not affect the reporter proteins tertiary structure enough to affect its reporter properties. The list of codons were collated into a list affectionately called 'The Good List' and from this Pascoe and Khadija began piecing them together and, using OligoAnalyser, analysing the folding of the constructs on their own. If the ribosome binding site or start codon were obstructed in the single construct strands, translation could be inhibit, which would hinder the comparator circuits functionality. |
===Labs=== | ===Labs=== | ||
| - | Lukas and Joy ran a gel of the double digest of the bacterial and mammalian hybrid promoter which was used to separate the fragments. We found DNA and hence the 250bp fragment was cut from the gel and purified. The ligation between the promoter (250bp fragment) and iGEM vector (pSB1C3) on Monday in preparation, the linearised backbone was digested. A different method of creating the bio-brick was attempted. As part of this method, the unpurified double digest sample was ligated into the pSB1C3 backbone. Three expected plasmid constructs might be observed and the desired one selected for next week. Rebecca made extra LB media which was poured into 5ml sample bottles and LB plates containing ampicillin at a concentration of 100 µg/ml. Lukas and Rebecca also isolated the plasmids containing the mammalian and bacterial hybrid promoters in both orientations. This was so that more DNA was ready to be used when needed. | + | . Lukas and Joy ran a gel of the double digest of the bacterial and mammalian hybrid promoter which was used to separate the fragments. We found DNA and hence the 250bp fragment was cut from the gel and purified. The ligation between the promoter (250bp fragment) and iGEM vector (pSB1C3) on Monday in preparation, the linearised backbone was digested. A different method of creating the bio-brick was attempted. As part of this method, the unpurified double digest sample was ligated into the pSB1C3 backbone. Three expected plasmid constructs might be observed and the desired one selected for next week. Rebecca made extra LB media which was poured into 5ml sample bottles and LB plates containing ampicillin at a concentration of 100 µg/ml. Lukas and Rebecca also isolated the plasmids containing the mammalian and bacterial hybrid promoters in both orientations. This was so that more DNA was ready to be used when needed. |
| - | After the previous days' investigation into the various reporter proteins for the project we decided to look at fluorescent proteins with vastly differing peaks. Ultimately we decided to investigate Red Fluorescence Protein (RFP) and Blue Fluorescence Protein (BFP) as there was very little overlap in their peaks, and it was concluded that we should characterise some biobricks that we already had to assess the fluorescent protein levels and their usefulness in our project. After looking at our kit plates we decided on using: | + | . After the previous days' investigation into the various reporter proteins for the project we decided to look at fluorescent proteins with vastly differing peaks. Ultimately we decided to investigate Red Fluorescence Protein (RFP) and Blue Fluorescence Protein (BFP) as there was very little overlap in their peaks, and it was concluded that we should characterise some biobricks that we already had to assess the fluorescent protein levels and their usefulness in our project. After looking at our kit plates we decided on using: |
| - | + | - [http://partsregistry.org/Part:BBa_R0080 BBa_R0080] - A promoter that should activate transcription in the presence of arabinose, in order to allow simple and easy activation of the fluorescence in transformed cells. | |
| - | + | - [http://partsregistry.org/Part:BBa_E0420 BBa_E0420] - A reporter expressing enhanced Cyan Fluorescence Protein (eCFP), a fluorescent protein with a similar peak to BFP that shouldn't overlap with RFP too much. This biobrick also contains the RBS and terminators meaning we can simply add the promoter of choice to complete the system. | |
| - | + | - [http://partsregistry.org/Part:BBa_K081014 BBa_K081014] - A reporter expression RFP. This biobrick also contains the RBS and terminators meaning we can simply add the promoter of choice to complete the system. | |
| - | Russell and Rachel transformed ''E. coli'' with the three biobricks of choice using the standard protocol described in previous experiments and plated onto agar plates containing 5µl ampicillin due to the plasmids for each biobrick including ampicillin resistance. The bacteria was left to grow overnight in the 37 °C incubator and placed in the fridge the next day following growth. | + | . Russell and Rachel transformed ''E. coli'' with the three biobricks of choice using the standard protocol described in previous experiments and plated onto agar plates containing 5µl ampicillin due to the plasmids for each biobrick including ampicillin resistance. The bacteria was left to grow overnight in the 37 °C incubator and placed in the fridge the next day following growth. |
The transformation protocol can be found [https://2012.igem.org/Team:NRP-UEA-Norwich/Protocol#Transformation Here] | The transformation protocol can be found [https://2012.igem.org/Team:NRP-UEA-Norwich/Protocol#Transformation Here] | ||
Revision as of 21:30, 21 September 2012
 "
"