Team:NRP-UEA-Norwich/Project
From 2012.igem.org
| Line 1: | Line 1: | ||
{{UEANRP}} | {{UEANRP}} | ||
| + | |||
| + | |||
| + | |||
| + | Sensory BioBrick systems have been a large constituent of previous iGEM projects, in which teams have combined impressive amounts of logic with limitless creativity to produce synthetically engineered organisms with the potential of detecting the presence of a substrate within its environment via innovative combinations of various promoter and reporter BioBricks. | ||
| + | |||
| + | Our own project combines | ||
==The Hybrid Promoter== | ==The Hybrid Promoter== | ||
| + | |||
[[File:MB.png | 200px | right]] | [[File:MB.png | 200px | right]] | ||
[[File:BM.png | 200px | right]] | [[File:BM.png | 200px | right]] | ||
| - | |||
| - | |||
Our own hybrid promoter hopes to add to the systems already in the registry by creating a hybrid promoter that combines the bacterial promoter PyeaR and the mammalian CArG element , both of which respond to exogenous nitrogenous species. Combining the two would allow a more modular NO sensor that can be used in mammalian and bacterial cells interchangeably. | Our own hybrid promoter hopes to add to the systems already in the registry by creating a hybrid promoter that combines the bacterial promoter PyeaR and the mammalian CArG element , both of which respond to exogenous nitrogenous species. Combining the two would allow a more modular NO sensor that can be used in mammalian and bacterial cells interchangeably. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| Line 29: | Line 25: | ||
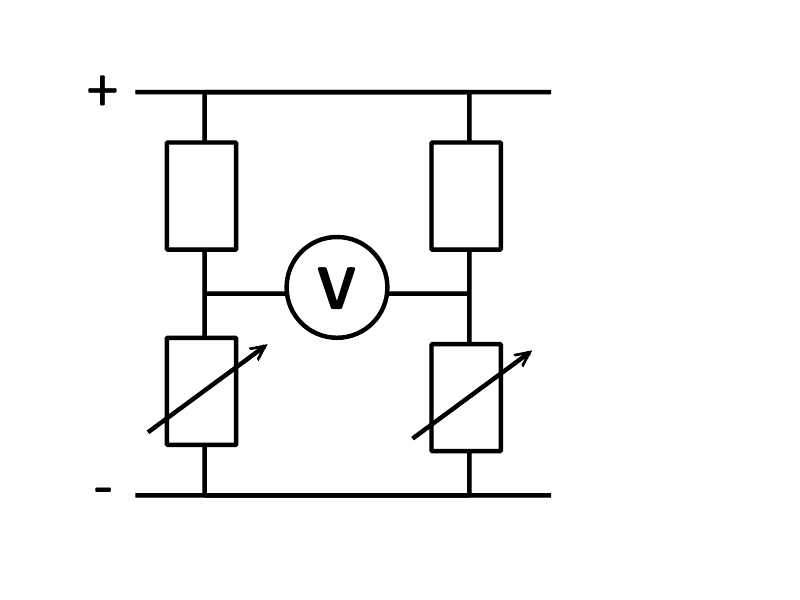
==The Comparator Circuit== | ==The Comparator Circuit== | ||
| - | [[File:ElectricalCircuit.png | 300px | | + | [[File:ElectricalCircuit.png | 300px |left]] |
This system relies on two interacting mRNA transcripts, both of which would ordinary be translated into a reporter (a fluorescent protein in our case) in the presence of a particular substrate. The idea being that these transcripts will only be made in the presence of certain substrates due to differing promoter activity. Two promoters with overlapping specificity would be used and, crucially, if both promoters detect the same substrates but differ in that one extra substrate is detected by one of the promoters, it is this substrate and this substrate only that our system will be able to detect in a simple and quantitative way. | This system relies on two interacting mRNA transcripts, both of which would ordinary be translated into a reporter (a fluorescent protein in our case) in the presence of a particular substrate. The idea being that these transcripts will only be made in the presence of certain substrates due to differing promoter activity. Two promoters with overlapping specificity would be used and, crucially, if both promoters detect the same substrates but differ in that one extra substrate is detected by one of the promoters, it is this substrate and this substrate only that our system will be able to detect in a simple and quantitative way. | ||
| Line 44: | Line 40: | ||
Due to the stop codon present in the scars of Assembly Standard 10 BioBricks, we decided that our constructs would have to be an Assembly Standard 23 BioBrick. Although the use of Bioscaffolds produced by previous iGEM teams was considered, time constraints meant that changing the Assembly Standard we put our BioBricks in would be the most convenient solution. | Due to the stop codon present in the scars of Assembly Standard 10 BioBricks, we decided that our constructs would have to be an Assembly Standard 23 BioBrick. Although the use of Bioscaffolds produced by previous iGEM teams was considered, time constraints meant that changing the Assembly Standard we put our BioBricks in would be the most convenient solution. | ||
| + | |||
| Line 58: | Line 55: | ||
The two gene constructs for the comparator circuit are currently being synthesised, and we look forward to receiving the synthetic gene soon and being able to characterise it! | The two gene constructs for the comparator circuit are currently being synthesised, and we look forward to receiving the synthetic gene soon and being able to characterise it! | ||
| + | |||
| + | |||
| + | ==Warrior Cell== | ||
| + | [[File:WarriorCell.png | 150px | right]] | ||
| + | |||
| + | |||
| + | |||
| + | A problem with current cancer therapeutic techniques is that they are not specific to cancer cells, leading to | ||
| + | patient discomfort. We hope to engineer E.coli cells that selectively identifies cancerous cells through their | ||
| + | anaerobic environment and secrete high concentrations of NO, mimicking macrophages, and thus killing | ||
| + | these cells. These cells are known to be hypoxic, and hypoxia is a hallmark for cancer cells already used in | ||
| + | tumour detection. Our Warrior Cell construct hopes to pave the way for a new kind of therapeutic strategy | ||
| + | that delivers a therapeutic agent only when and where it is needed. | ||
Revision as of 10:13, 31 August 2012
 "
"