Team:NRP-UEA-Norwich/Parts
From 2012.igem.org
Contents |
<groupparts>iGEM012 NRP-UEA-Norwich</groupparts>
1) Part:BBa_K216005:Please see experience page
[1] (Group: iGEM09_Edinburgh (2009-09-25))
2) Part:BBa_K381001:Please see experience page [2] (Group: iGEM10_BCCS-Bristol (2010-10-05))
1)Bacterial-Mammalian Promoter, Part:BBa_K774000[3]
2)Mammalian-Bacterial Promoter, Part:BBa_K774001[4]
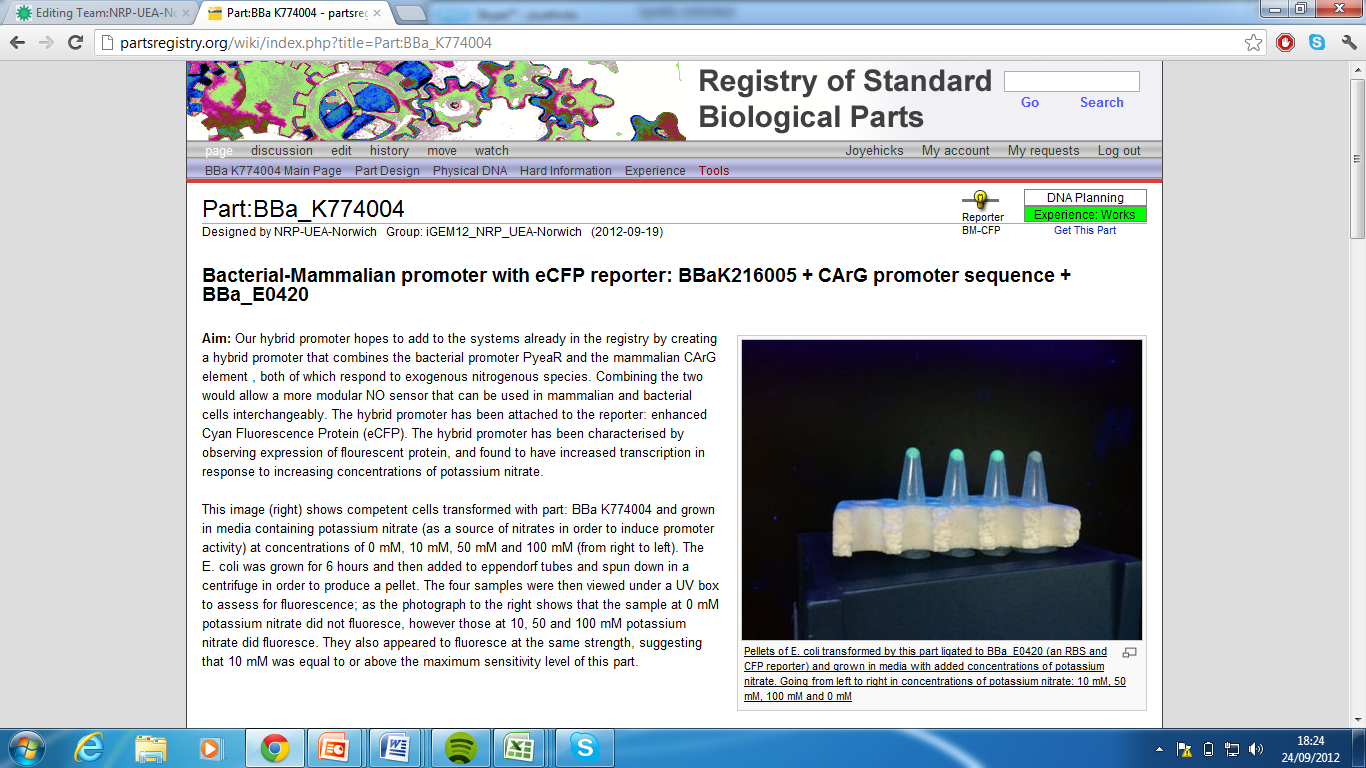
3)Bacterial-Mammalian promoter with eCFP reporter, Part:BBa_K774004[5]
4)Bacterial-Mammalian promoter with RFP reporter,Part:BBa_K774005[6]
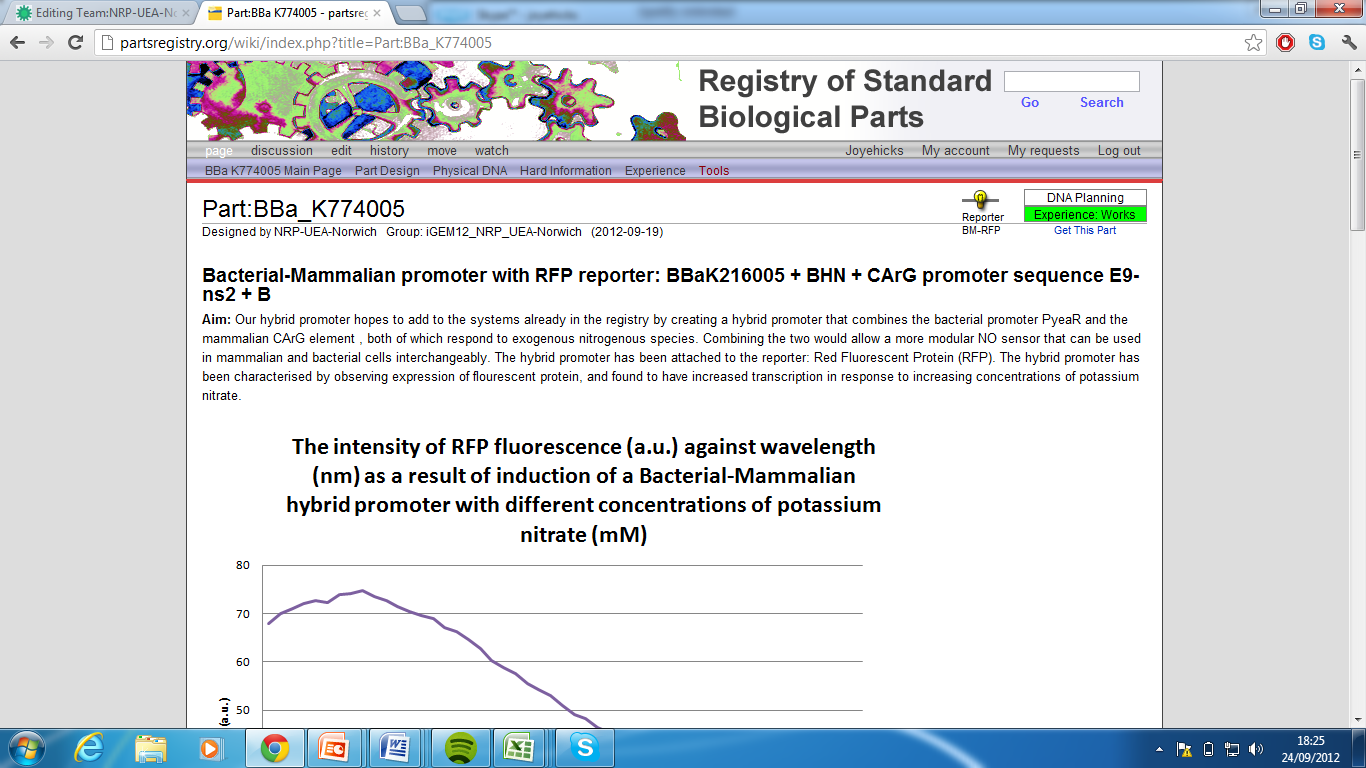

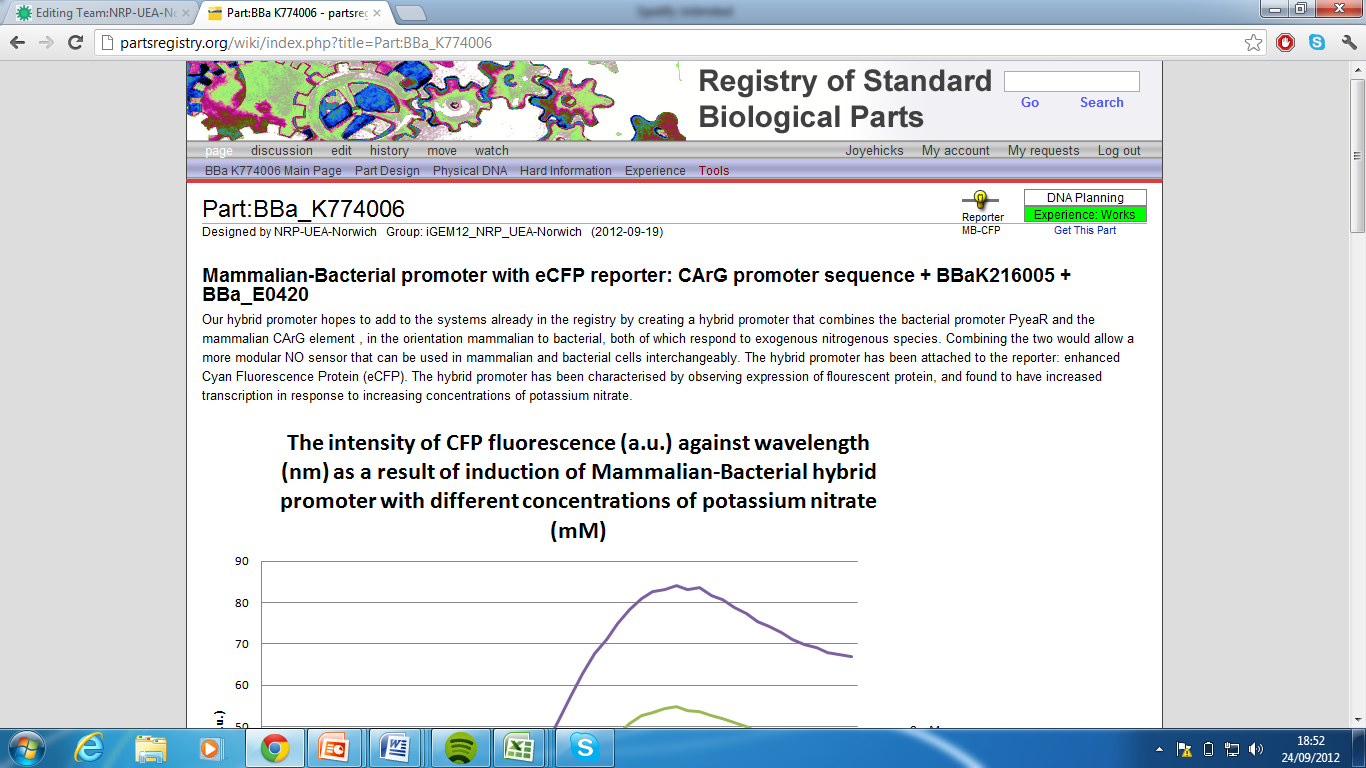
5)Mammalian-Bacterial promoter with eCFP reporter, Part:BBa_K774006[7]
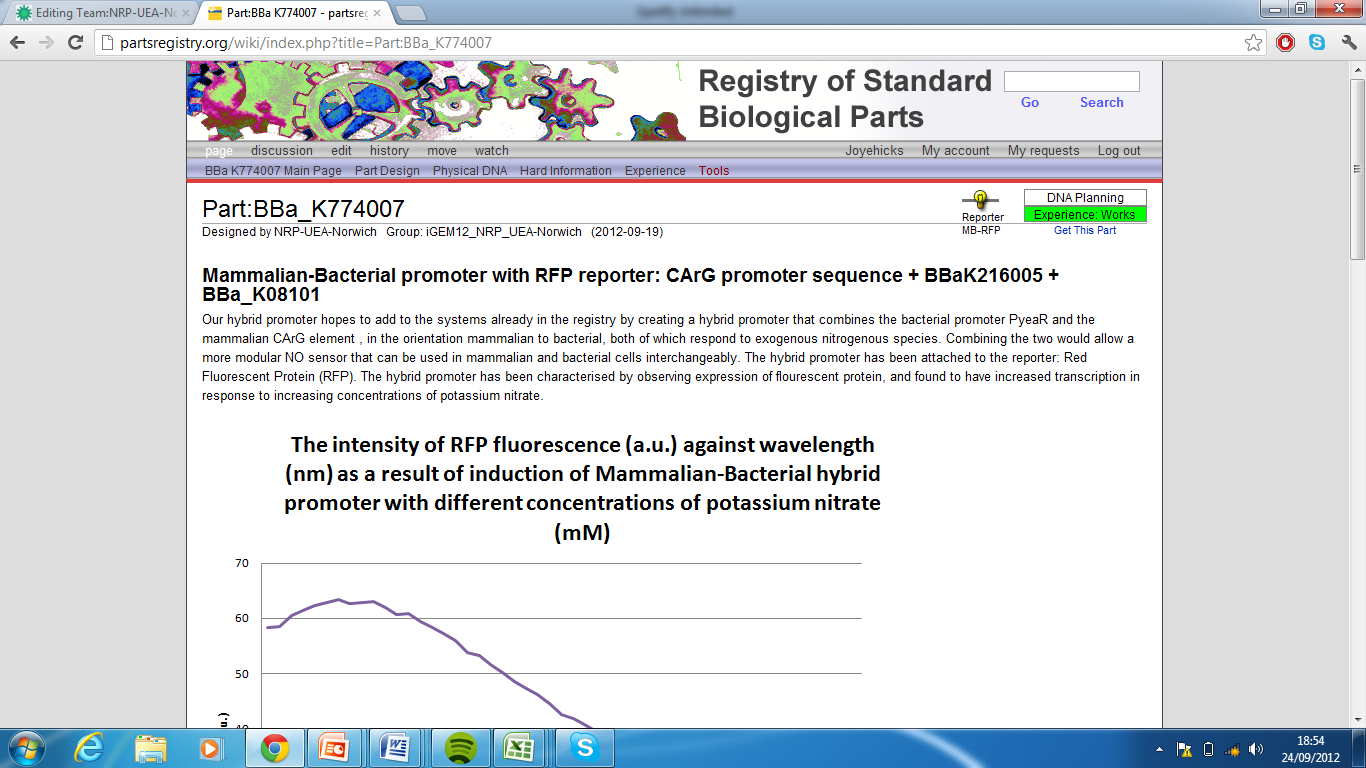
6)Mammalian-Bacterial promoter with RFP reporter, Part:BBa_K774007[8]
1)Comparator Circuit Part 1, Part:BBa_K774002
2)Comparator Circuit Part 2, Part:BBa_K774003
This system relies on two interacting mRNA transcripts, both of which would ordinary be translated into a reporter (a fluorescent protein in our case) in the presence of a particular substrate. The idea being that these transcripts will only be made in the presence of certain substrates due to differing promoter activity. Two promoters with overlapping specificity would be used and, crucially, if both promoters detect the same substrates but differ in that one extra substrate is detected by one of the promoters, it is this substrate and this substrate only that our system will be able to detect in a simple and quantitative way.
Our system relies on two constructs that interact via complimentary base pair sequences both before and after the ribosome binding site of the reporter protein. The idea being that, when both transcripts are present in the chassis, they would bind together, inhibiting the translation of the reporter proteins. Any imbalance of transcription due to the presence of the substrate of interest results in free mRNA of the gene system that detects that substrate.
Our team have constructed a countercurrent comparator circuit in which the reporter proteins are at the same end of the complimentary region, although a contracurrent system has been theorised. Both systems share a crucial subtractive nature comparable to an analogue computer. We envisage that, should the system be fine-tuned and expanded on, a variety of different business sectors from agriculture to spinoff pharmaceutical companies (such as the fictious QuantiCare) could capitalise on this novel genetic technology.