Team:NRP-UEA-Norwich/NOSensing
From 2012.igem.org
Welcome to the NRP UEA iGEM 2012 Wiki Projects Menu
Please choose the relevant link to view an overview of each project!
Nitric Oxide Sensing & The Hybrid Promoters | The Comparator Circuit | Theoretical Projects
Our main project has resulted in the production of a hybrid bacterial and mammalian promoter optimised for induction by nitric oxide, nitrates and nitrites. We have ligated PyeaR, a known bacterial promoter and Part BBa_K216005 in the parts registry, with its mammalian counterpart, CArG. The resulting hybrid promoter has been synthesised in two orientations; PyeaR (bacterial, B) upstream of CArG (mammalian, M), nicknamed (B-M); and CArG upstream of PyeaR (M-B). These orientations were submitted to the parts registry as our first two biobricks.
Each orientation of the promoter was ligated to enhanced cyan fluorescent protein (eCFP) and red fluorescent protein (RFP) to produce four new biobricks which have been submitted to the parts registry. These promoter + fluorescent protein biobricks have been characterised following transformation into Escherichia coli and induction by potassium nitrate using methods such as flow cytometry, fluorescence-activated cell sorting (FACS) and scanning with a fluorometer. The data from these experiments has proved that our promoter works as we expected it to. We have also transfected M-B + eCFP into a human breast cancer cell line, MCF7, and have proved the system is flexible and can be used in both eukaryotes and prokaryotes.
We believe the promoters we have produced have relevant uses in cancer therapeutics, soil fertilisation and detection of emissions from industries such as construction.
Parts produced from this project:
Bacterial-Mammalian/B-M (PyeaR-CArG) Hybrid Promoter -- Mammalian-Bacterial/M-B (CArG-PyeaR) Hybrid Promoter -- B-M + eCFP -- B-M + RFP -- M-B + eCFP -- M-B + RFP
The University of East Anglia hosts many research teams whose work focuses on studying nitrogenous species and the way in which bacteria use and modify those species. One major problem that is faced by these teams involves working with nitric oxide (NO), as NO is highly reactive with a low half life, therefore making it difficult to trace and quantitatively measure accurately. Many of the methods currently used to measure NO levels are unable to distinguish between homogenous species, such as nitrates (NO3) and nitrites (NO2), therefore the figure given for NO levels is often inaccurate as other nitrogenous species are taken into account.
The ability to be able to accurately detect NO levels is one with a great deal of potential for the future. Nitric oxide has been noted as a possible cancer therapy due to its physiological use as an apoptosis inducer by macrophages, however NO is also known to be used by cancerous cells to establish a baseline and use it to induce apoptosis and promote proliferation of a tumour; being able to accurately sense nitric oxide and go on to act on that information could be very useful to prevent the NO baseline being established by cancerous cells, but to also use NO for its apoptosis-inducing abilities. There are also other potential applications in the construction business, in 2008 NEED TO FIND REFERENCE OF PEOPLE released legislation encouraging construction companies to monitor their NO output as it is inversely proportional to carbon monoxide (a toxic substance which needs to be regulated); the ability to accurately detail levels of NO being released in these circumstances would be highly useful.
Nitric oxide is an extremely physiologically relevant molecule found within both eukaryotes and prokaryotes, where it is utilised by different enzymes and systems for various roles. The aim of the experiment was to devise a hybrid promoter that could be applied to eukaryotes and prokaryotes in order to begin to more accurately sense NO and report on its specific levels.
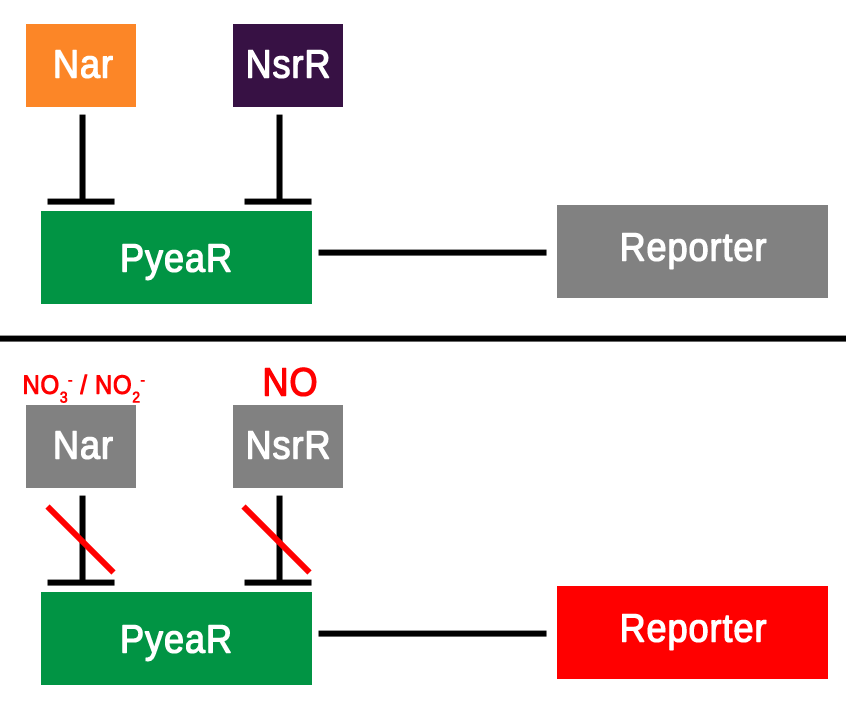
The hybrid promoter was designed to take on both bacterial and mammalian elements in order to be compatible with both bacterial and mammalian cells. After research around the subject and searching the parts registry a promoter known as PyeaR was decided upon as the bacterial element of the hybrid promoter; PyeaR is found in the yeaR/yoaG operon of Escherichia coli and is associated with induction by nitric oxide, nitrates and nitrites (Lin et al., 2007). PyeaR is repressed by two main repressors; Nar, which is regulated by nitrates and nitrites; and NsrR, which is regulated by nitric oxide (Figure 1.). One of the key elements of PyeaR is that it is not repressed in aerobic conditions, allowing for easier carrying out of experiments. The PyeaR aspect of the hybrid promoter has been known throughout the project as the bacterial promoter, or simply B.
The mammalian element of the hybrid promoter was produced by nine CArG elements (repeated elements of CC(A/T)(6)GG), a promoter previously used synthetically for nitric oxide synthase as a cancer therapy (Worthington et al., 2005) and developed from the EGR1 gene for early growth response protein 1 (Scott et al., 2002). The CArG aspect of the hybrid promoter has been known throughout the project as the mammalian promoter, or simply M.
Following identification of the two elements of the hybrid promoters the B (PyeaR) and M (CArG) aspects were ligated to one another in two orientations; B upstream of M (B-M) and M upstream of M (M-B) (Figure 2.). The hybrid promoters were synthesised in a pUC57 backbone with the standard iGEM restriction sites of EcoR1/Xba1 upstream of the promoter, and Spe1/Pst1 downstream of the promoter. A BamH1 restriction site was included in between the B and M sequences in order to allow for the B and M elements to be separated, as well as for easy verification of the promoter having been ligated into the iGEM backbone in future experiments (as BamH1 does not already exist in the pSB1C3 backbone).
The DNA for the synthesised genes of B-M and M-B had been supplied in the pUC57 backbone, therefore it was necessary for B-M and M-B to be digested from the pUC57 backbone and ligated into the pSB1C3 backbone. The synthesised gene was transformed into competent ’’E. coli’’ cells, which in turn were grown on agar plates containing 100 µg/ml ampicillin (due to pUC57 containing ampicillin resistance); colonies that had grown were then inoculated into liquid culture, and the liquid culture was subsequently mini-prepped using either the Bioline ISOLATE Plasmid DNA Mini Kit or the Promega Wizard® Plus SV Minipreps DNA Purification System. The DNA that had been extracted through mini-preps and the pSB1C3 backbone, as provided by the iGEM registry, were then digested using EcoR1 and Pst1 and a ligation was carried out using standard assembly protocol. The product of ligation was then transformed into competent E. coli cells, which were grown on agar plates containing 2.5 µg/ml chloramphenicol (due to pSB1C3 containing chloramphenicol resistance); this was done to eliminate any bacteria that had been transformed with undesirable ligation products.
The colonies that had grown were then grown in liquid culture and mini-prepped in order to extract the DNA; the extracted DNA was then sent for sequencing, and the returned sequenced matched the expected sequence. The DNA was then sent to the parts registry as the Bacterial-Mammalian Hybrid Promoter and the Mammalian-Bacterial Hybrid Promoter. The biobricks for B-M and M-B were then used for further experiments, including ligation with a fluorescent protein reporter and growth studies.
From weeks one through to five the team worked on producing the first biobricks of the hybrid promoter. Despite this proving difficult due to various reasons such as low amounts of DNA being produced from early mini-preps and ligation strategies not working, E. coli transformed by the biobrick DNA was successfully grown and proven to have the relevant resistance by the beginning of week six.
In order to test the activity of the hybrid promoters a reporter needed to be ligated. As the hybrid promoter did not already contain a ribosome binding site (RBS) both the RBS and the reporter were needed to be ligated to the promoter; in order to help improve experimental efficiency the parts registry was searched for relevant reporters that also contained an RBS. In week three two reporters were identified as BBa_E0420, a biobrick for enhanced CFP (eCFP) + RBS + terminators, and BBa_K081014, a biobrick for RFP + RBS + terminators.
Once the B-M and M-B biobricks had been created in week six work began in earnest on the fluorescent proteins and ligating the promoters to them in order to begin characterisation. Due to many set-backs with low levels of DNA and having to order more biobricks from the registry, a successful ligation of the promoter to a fluorescent protein reporter was finally achieved in week ten. In order to carry out the ligation the promoter was first digested using Spe1 and Pst1 in order to linearise the backbone downstream of the promoter; the fluorescent proteins were digested using Xba1 and Pst1 in order to remove the insert. A ligation was then carried out using standard assembly protocol and the ligation products were transformed into E. coli competent cells, which in turn were grown on agar plates 2.5 µg/ml chloramphenicol (due to pSB1C3 containing chloramphenicol resistance).
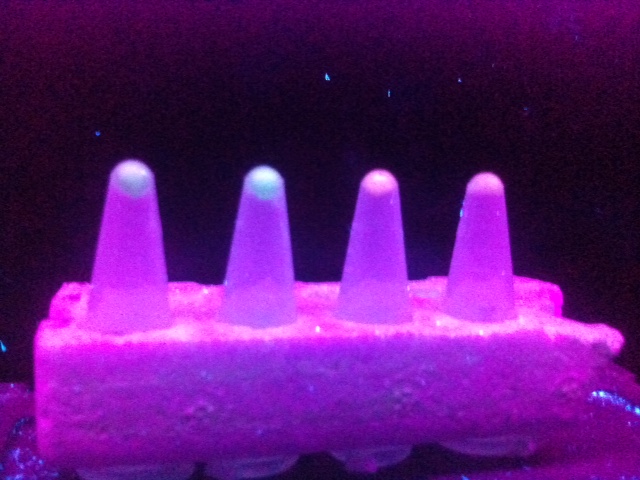
In order to quickly identify colonies of bacteria containing the promoter, RBS and reporter in a likely correct sequence a range of colonies were inoculated into media also containing potassium nitrate (KNO3) solution; this was done in order to inhibit the Nar repressor in PyeaR and result in activation of the promoter/induction of transcription/expression of the fluorescent protein reporter. Samples of the inoculated media containing KNO3 were then added to an eppendorf and spun down to form a pellet, which was viewed under a UV box and observed for fluorescence. After a week of various ligation and transformation experiments both promoters were successfully ligated to both fluorescent proteins and fluorescence of eCFP and RFP was observed under a UV box (Figure 3.).
From these experiments four more biobricks were produced and submitted to the registry: B-M + eCFP, B-M + RFP, M-B + eCFP, and M-B + RFP. These four new biobricks were then used to further characterise the activity of the M-B and B-M hybrid promoters.
In order to characterise the hybrid promoters ligated to fluorescent proteins a number of experiments were carried out to measure the level of fluorescent output at different concentrations of KNO3 (used to induce the promoter's activity). All of these experiments were carried out in week eleven and were as follows:
Three tubes of media were inoculated with bacteria transformed by the B-M + RFP biobrick. Each tube then had potassium nitrate added to it at different concentrations; 0 mM, 1 mM and 10 mM. The samples were grown over night and then fixed using PFA, before being run through a flow cytometer in order to count the number of cells fluorescing at the different concentrations of KNO3.
Note: This was the first time the flow cytometers at the University of East Anglia had been used with E. coli
Three tubes of media were inoculated with bacteria transformed by the B-M + RFP biobrick, and three tubes of media were inoculated with 200 µL bacteria transformed by the M-B + eCFP biobrick. Potassium nitrate was then added to some of the tubes in order to create samples as follows:
. B-M + RFP with 0 mM, 1 mM and 10 mM potassium nitrate
. M-B + eCFP with 0 mM, 1 mM and 10 mM potassium nitrate
These samples were then grown over night before being fixed with PFA and run through a flow cytometer optimised for FACS in order to count the number of cells fluorescing at different concentrations of KNO3.
Note: This was the first time the flow cytometers at the University of East Anglia had been used with E. coli
The main characterisation of the biobricks was carried out using a fluorometer. Five tubes of media containing 200 µL transformed bacteria and potassium nitrate were grown for each biobrick in concentrations as follows:
. B-M + RFP with 0 mM, 5 mM, 10 mM, 15 mM and 20 mM potassium nitrate
. B-M + eCFP with 0 mM, 5 mM, 10 mM, 15 mM and 20 mM potassium nitrate
. M-B + RFP with 0 mM, 5 mM, 10 mM, 15 mM and 20 mM potassium nitrate
. M-B + eCFP with 0 mM, 5 mM, 10 mM, 15 mM and 20 mM potassium nitrate
The samples were then spun down in a centrifuge and resuspended in Tris buffer before being subjected to sonication in order to lyse the cells. The resulting solution was then spun down in a centrifuge again before the supernatant was extracted (in order to separate the proteins from the rest of the cell). The supernatant was then run through a fluorometer in order to measure the intensity of fluorescence as a method to gauge the activity of the hybrid promoters. RFP-containing samples were excited with a wavelength of 560 nm and the emission measured from 600 – 650 nm; eCFP-containing samples were excited with a wavelength of 410 nm and emission measured from 440-490 nm.
These characterisation experiments gave rise to the characterisation which is now available on the main page for all hybrid promoter + fluorescent protein parts on the registry, and the experience section of the hybrid promoters alone on the registry.
In order to ascertain the flexibility of the hybrid promoter and help prove our original aim of producing a promoter that could be used in both eukaryotes and prokaryotes correct, the DNA for M-B + eCFP was transfected into the MCF7 human breast cancer cell line. An experiment was set up using S-Nitroso-N-acetyl-DL-penicillamine (SNAP), a nitric oxide donor, in order to induce expression of the fluorescent protein. A six-well transfection slide was produced containing:
. MCF7 cells, untransfected without SNAP
. MCF7 cells, untransfected with SNAP
. MCF7 cells, transfected without SNAP
. MCF7 cells, transfected with SNAP
The cells were then left for a day before they were imaged with a fluorescence microscope in order to observe expression of eCFP.
. Studied the growth rate
. Photos
. Graphs from flow and fluorometer
. Pictures etc. from mammalian transfection
. Growth study on MB/BM
BM/MB better or the same, RFP/CFP better or the same etc.
A problem with current cancer therapeutic techniques is that they are not specific to cancer cells, leading to patient discomfort. We hope to engineer E.coli cells that selectively identifies cancerous cells through their anaerobic environment and secrete high concentrations of NO, mimicking macrophages, and thus killing these cells. These cells are known to be hypoxic, and hypoxia is a hallmark for cancer cells already used in tumour detection. Our Warrior Cell construct hopes to pave the way for a new kind of therapeutic strategy that delivers a therapeutic agent only when and where it is needed.
. Full quantitative analysis to see where the values lie; combine with tuners for different sensitivity levels
. With different substrates (e.g. nitrite salt, NO donor)
. With different reporter/effector enzyme
Lin H.Y., Bledsoe P.J., Stewart V., (2007), Activation of yeaR-yoaG Operon Transcription by the Nitrate-Responsive Regulator NarL Is Independent of Oxygen- Responsive Regulator Fnr in Escherichia coli K-12▿, Journal of Bacteriology, 189: 7539 - 7548
Scott S.D., Joiner M.C., Marples B., (2002), Optimizing radiation-responsive gene promoters for radiogenetic cancer therapy., Gene Therapy, 9: 1396-1402
Worthington J., Robson T., Scott S., Hirst, D., (2005), Evaluation of a synthetic CArG promoter for nitric oxide synthase gene therapy of cancer, Gene Therapy, 12: 1417–1423