Team:LMU-Munich/Spore Coat Proteins
From 2012.igem.org
Franzi.Duerr (Talk | contribs) |
Franzi.Duerr (Talk | contribs) |
||
| Line 55: | Line 55: | ||
===Cloning Results=== | ===Cloning Results=== | ||
{| "width=100%" style="text-align:center;" style="align:right"| | {| "width=100%" style="text-align:center;" style="align:right"| | ||
| - | |<p align="justify">Main results of the various constructs | + | |<p align="justify">Main results of the various constructs that were created</p> |
|[[File:LMU Firstspore.jpg|150px|right|link=Team:LMU-Munich/Spore_Coat_Proteins/result]] | |[[File:LMU Firstspore.jpg|150px|right|link=Team:LMU-Munich/Spore_Coat_Proteins/result]] | ||
|- | |- | ||
| Line 62: | Line 62: | ||
</div> | </div> | ||
| - | + | <div class="box"> | |
| - | + | ===Purification methods=== | |
| - | < | + | {| "width=100%" style="text-align:center;" style="align:right"| |
| - | + | |<p align="justify">Description of the different purification methods of the spores</p> | |
| - | + | |[[File:Treatments.png|200px|right|link=Team:LMU-Munich/Spore_Coat_Proteins/purification]] | |
| - | + | ||
| - | {| | + | |
|- | |- | ||
| - | + | ! colspan="2" |[[File:LMU Arrow purple.png|40px|link=Team:LMU-Munich/Spore_Coat_Proteins/purification]] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | ! | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | |[[File: | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
|} | |} | ||
| + | </div> | ||
Revision as of 11:07, 24 October 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Sporobeads - What Protein Do You Want to Display?
Crust Promoter Evaluation
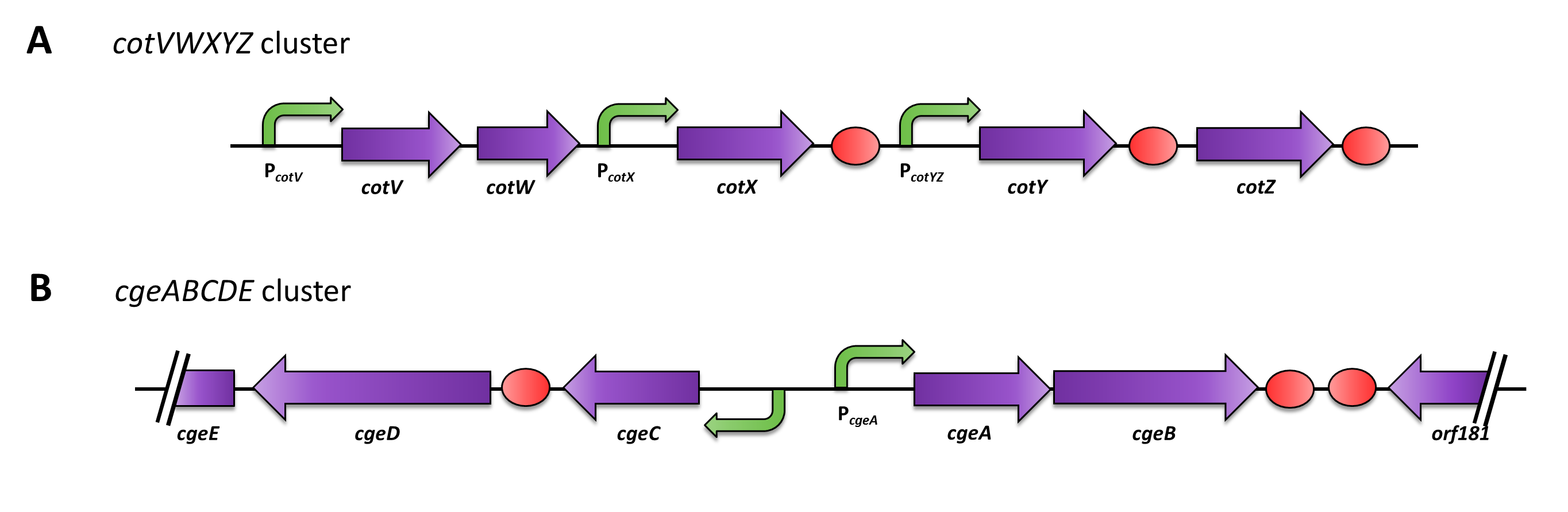
Three sporulation-dependent promoters (PcotYZ, PcotV, PcgeA) were constructed, two of which show the expected behavior. | 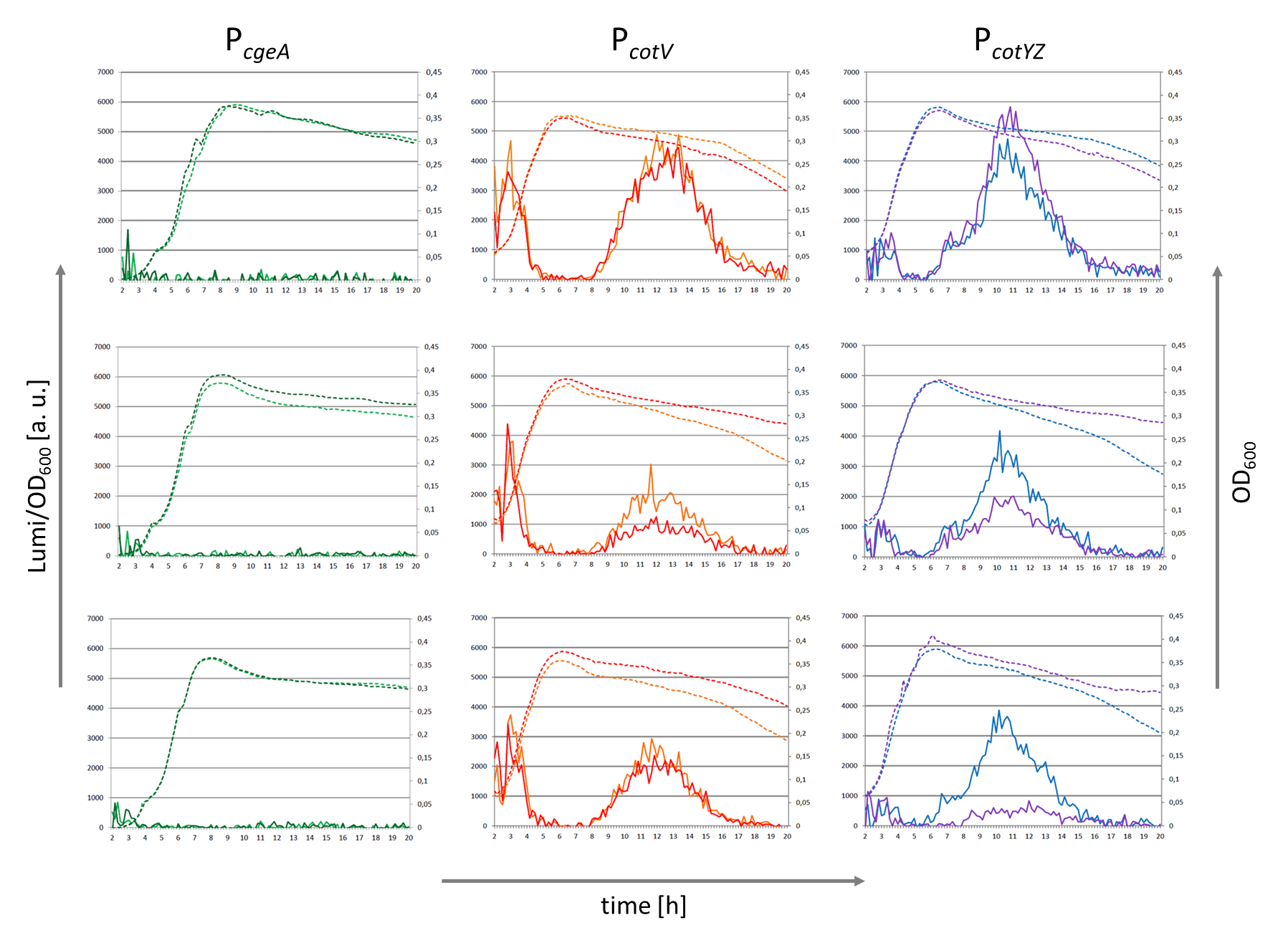
|
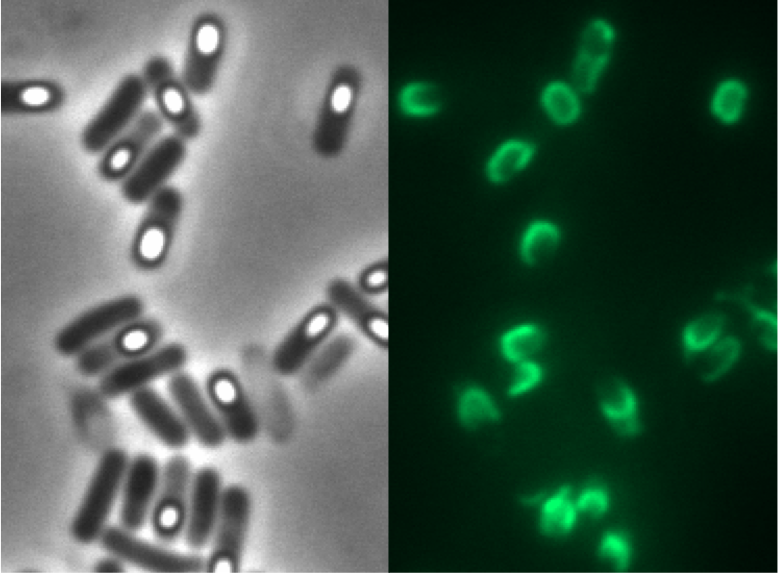
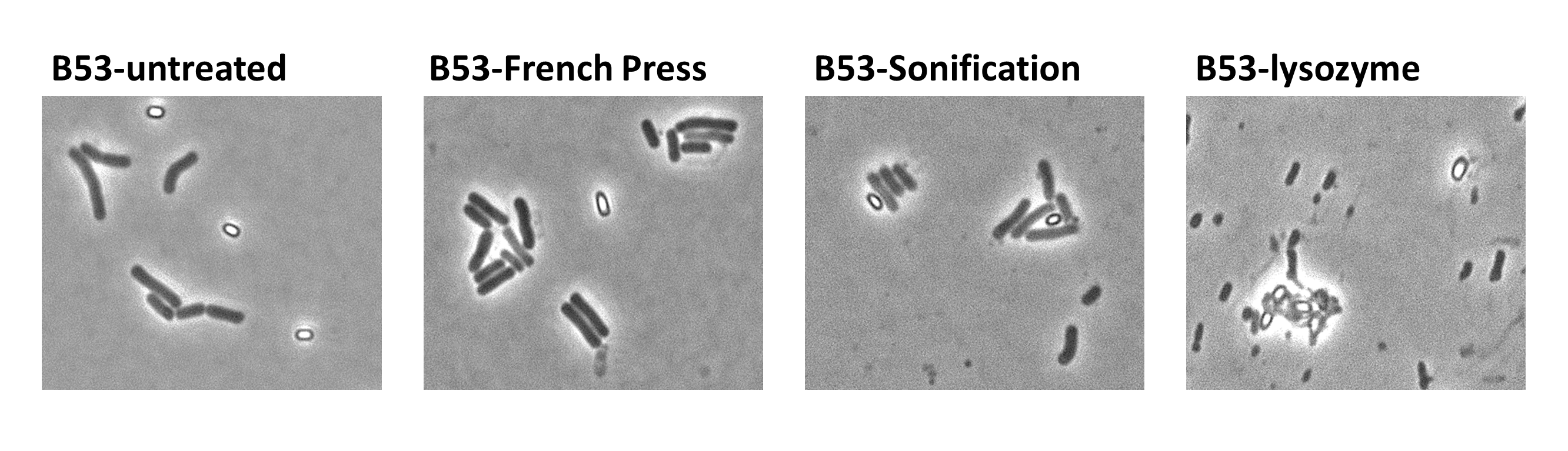
Since there were still some vegetative cells left after 24 hours of growth in Difco Sporulation Medium, we wanted to purify the Sporobeads. We tried three different methods for this approach: treatment with French Press, sonification and lysozyme. By means of the microscopy results we were able to conclude that lysozyme treatment was the only successful method (see data). Additionally, this treatment did not harm the crust fusion proteins as green fluorescence was still detectable afterwards (see data for details).
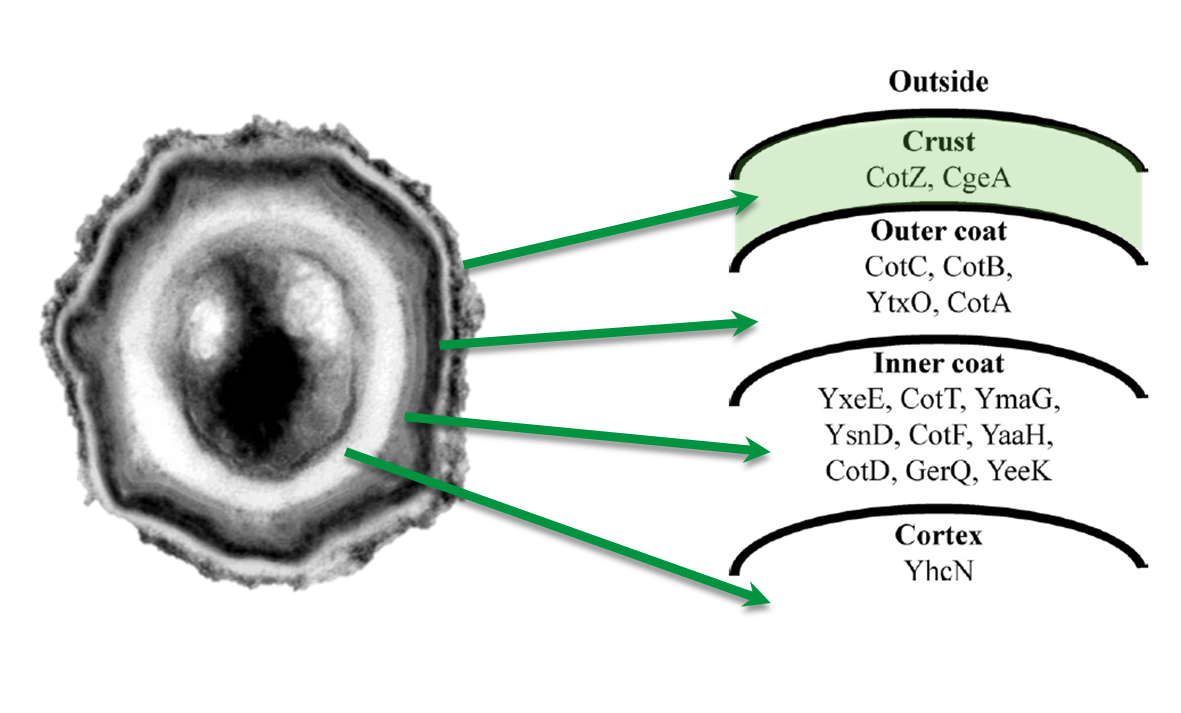
Moreover, we wondered if clean deletions of the native spore crust genes would show any difference in fusion protein expression in our Sporobeads. Thus, we deleted the native cotZ and cgeA using the pMAD based gene deletion strategy described by Arnaud et al., 2004.
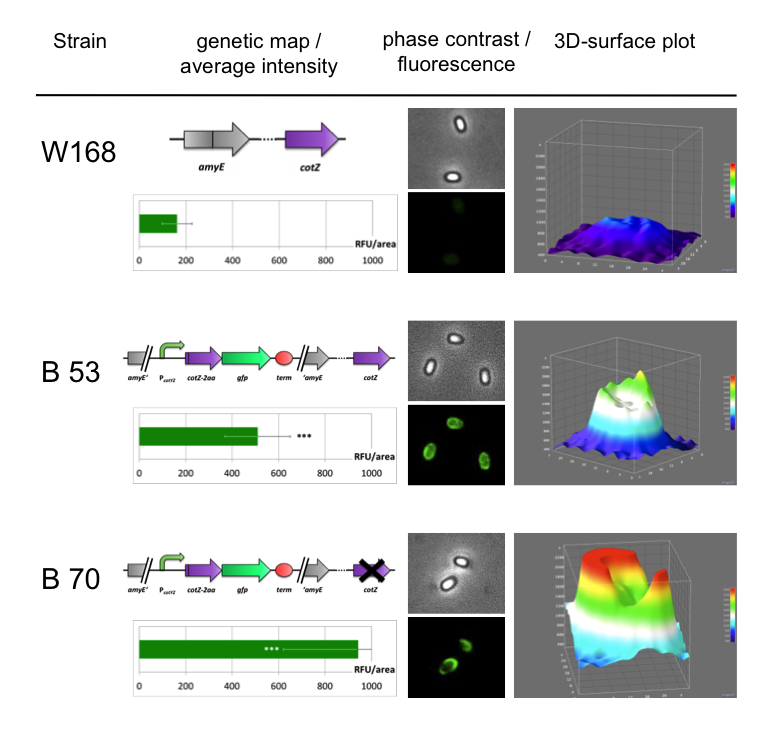
Because of the low but distinct fluorescence of wild type spores, we measured and compared the fluorescence intensity of 100 spores per construct (see data). We obtained significant differences between wild type spores and all of our Sporobeads (see data). The intensity bar charts in Fig. 6 show the fluorescence intensity, while the 3D graphs illustrate the distribution of fluorescence intensity across the spore surface. This correlates with the localization of our fusion proteins in the crust. For image analysis we measured the fluorescence intensity of an area of 750 pixel per spore by using ImageJ and evaluated the results with the statistical software R. The following graph (Fig. 6) shows the results of microscopy and ImageJ analysis of the strongest construct integrated into wildtype W168 (B53) and the deletion strain B 49 (B70).
|
| |
|
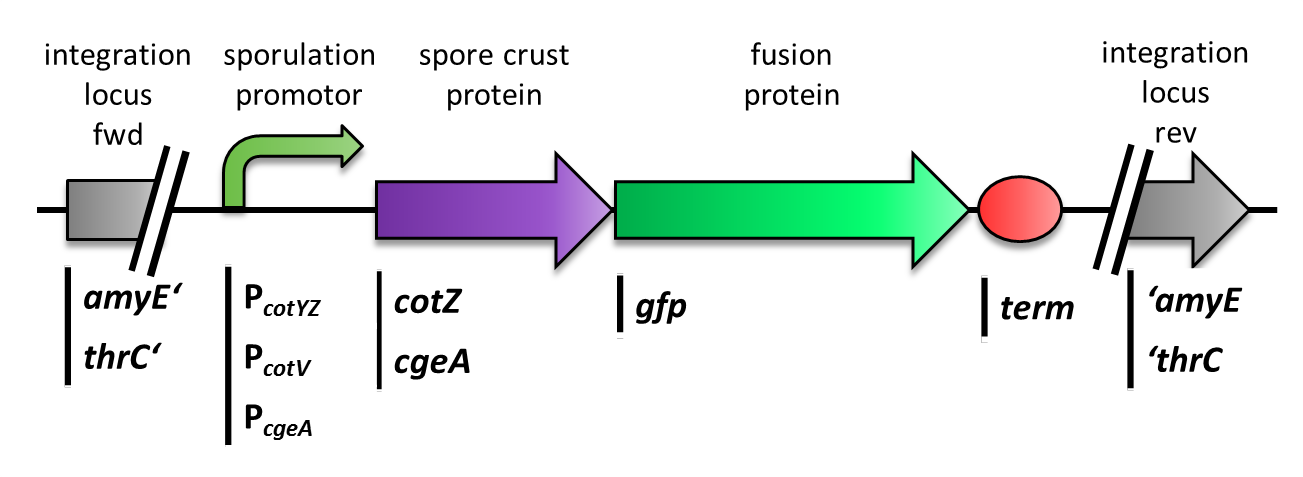
As shown in Fig. 6, the wild type spore has hardly any fluorescence, whereas both Sporobeads with the integrated construct pSBBs1C-PcotYZ-cotZ-2aa-gfp-terminator give a distinct fluorescence signal around the edge of the spore. Furthermore, it demonstrates that strain B 70 has the highest fluorescence intensity.
In summary we successfully developed functional sporobeads that are capable of displaying any protein of choice on the surface of modified B. subtilis endospores.

| 
| 
| 
|
| Bacillus Intro | Bacillus BioBrickBox | Sporobeads | Germination STOP |
|
 "
"