Team:LMU-Munich/Spore Coat Proteins
From 2012.igem.org
Franzi.Duerr (Talk | contribs) |
Franzi.Duerr (Talk | contribs) |
||
| Line 31: | Line 31: | ||
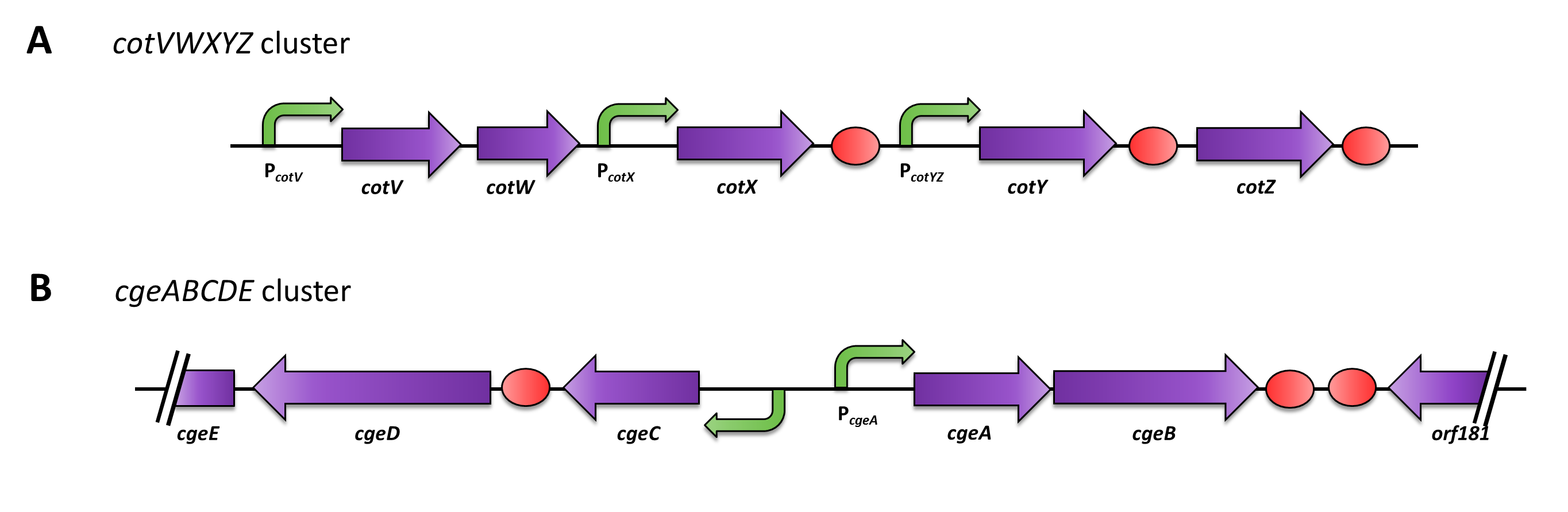
| - | <p align="justify">The gene ''cgeA'' is located in the ''cgeABCDE'' cluster and is expressed from its own promoter P<sub>''cgeA''</sub>. The cluster ''cotVWXYZ'' contains the gene ''cotZ'' which is cotranscribed with ''cotY'' and expressed from the promoter P<sub>''cotYZ''</sub>. Another promoter of this cluster, P<sub>''cotV''</sub>, is responsible for the transcription of the other three genes (Fig. 2). Those three promoters were | + | <p align="justify">The gene ''cgeA'' is located in the ''cgeABCDE'' cluster and is expressed from its own promoter P<sub>''cgeA''</sub>. The cluster ''cotVWXYZ'' contains the gene ''cotZ'' which is cotranscribed with ''cotY'' and expressed from the promoter P<sub>''cotYZ''</sub>. Another promoter of this cluster, P<sub>''cotV''</sub>, is responsible for the transcription of the other three genes (Fig. 2). Those three promoters were evaluated with the ''lux'' reporter genes ([http://partsregistry.org/Part:BBa_K823025 pSB<sub>''Bs''</sub>3C-''lux''ABCDE]) to analyze their strength and the time point of their activation (see [https://2012.igem.org/Team:LMU-Munich/Data/crustpromoters data]).</p> |
| Line 51: | Line 51: | ||
| - | [[File:Spore crust proteins cycle.jpg|600px|right]] | + | [[File:Spore crust proteins cycle.jpg|600px|right]] |
{| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="3" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
| style="width: 100%;background-color: #FFFFFF;" | | | style="width: 100%;background-color: #FFFFFF;" | | ||
| Line 77: | Line 77: | ||
{| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
| - | <font color="#000000"; size="2">Fig. | + | <font color="#000000"; size="2">Fig. 4: Section of the genome of ''B. subtilis'' with the various integrated constructs.</font> |
|} | |} | ||
|} | |} | ||
| Line 121: | Line 121: | ||
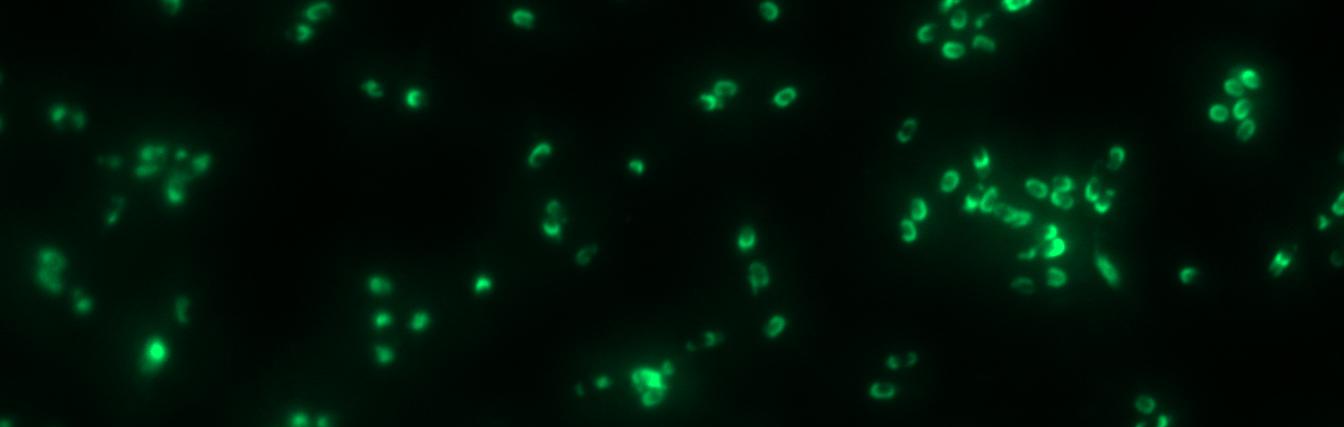
| - | <p align="justify">Finally, we started with the most important experiment for our GFP-'''Sporo'''beads, the fluorescence microscopy. We developed a sporulation protocol (for details see [https://static.igem.org/mediawiki/2012/e/e9/LMU-Munich_2012_Protocol_for_enhancement_of_mature_spore_numbers.pdf Protocol for enhancement of mature spore numbers]) that increases the rates of mature spores in our samples. The cells were fixed on agarose pads and investigated by phase contrast and fluorescence microscopy. While spores of the wild type only showed the known background fluorescence, all '''Sporo'''beads showed bright green fluorescence at the edge (on the surface) of the spores. '''Sporo'''bead from strain B 53 (containing the P<sub>''cotYZ''</sub>-''cotZ''-''gfp''-terminator construct)showed the highest fluorescence intensity (see | + | <p align="justify">Finally, we started with the most important experiment for our GFP-'''Sporo'''beads, the fluorescence microscopy. We developed a sporulation protocol (for details see [https://static.igem.org/mediawiki/2012/e/e9/LMU-Munich_2012_Protocol_for_enhancement_of_mature_spore_numbers.pdf Protocol for enhancement of mature spore numbers]) that increases the rates of mature spores in our samples. The cells were fixed on agarose pads and investigated by phase contrast and fluorescence microscopy. While spores of the wild type only showed the known background fluorescence, all '''Sporo'''beads showed bright green fluorescence at the edge (on the surface) of the spores. '''Sporo'''bead from strain B 53 (containing the P<sub>''cotYZ''</sub>-''cotZ''-2aa-''gfp''-terminator construct) showed the highest fluorescence intensity (see Fig. 5 and all [https://2012.igem.org/Team:LMU-Munich/Data/gfp_spore data]). Hence, this strain was chosen for further experiments.</p> |
| Line 138: | Line 138: | ||
| - | <p align="justify">Since there were still some vegetative cells left after 24 hours of growth in Difco Sporulation Medium, we wanted to purify the '''Sporo'''beads. We tried three different methods for this approach: treatment with French Press, sonification and lysozyme. By means of the microscopy results we were able to conclude that lysozyme treatment was the only successful method (see [https://2012.igem.org/Team:LMU-Munich/Data/Sporepurification data]). Additionally, this treatment did not harm the crust fusion proteins as green fluorescence was still detectable afterwards (see data for details).</p> | + | <p align="justify">Since there were still some vegetative cells left after 24 hours of growth in Difco Sporulation Medium, we wanted to purify the '''Sporo'''beads. We tried three different methods for this approach: treatment with French Press, sonification and lysozyme. By means of the microscopy results we were able to conclude that lysozyme treatment was the only successful method (see [https://2012.igem.org/Team:LMU-Munich/Data/Sporepurification data]). Additionally, this treatment did not harm the crust fusion proteins as green fluorescence was still detectable afterwards (see [https://2012.igem.org/Team:LMU-Munich/Data/Sporepurification data] for details).</p> |
<p align="justify">Moreover, we wondered if clean deletions of the native spore crust genes would show any difference in fusion protein expression in our '''Sporo'''beads. Thus, we deleted the native ''cotZ'' and ''cgeA'' using the pMAD based gene deletion strategy described by [http://www.ncbi.nlm.nih.gov/pubmedterm=New%20Vector%20for%20Efficient%20Allelic%20Replacement%20in%20Naturally%20Nontransformable%2C%20Low-GC-Content%2C%20Gram-Positive%20Bacteria Arnaud ''et al''., 2004].</p> | <p align="justify">Moreover, we wondered if clean deletions of the native spore crust genes would show any difference in fusion protein expression in our '''Sporo'''beads. Thus, we deleted the native ''cotZ'' and ''cgeA'' using the pMAD based gene deletion strategy described by [http://www.ncbi.nlm.nih.gov/pubmedterm=New%20Vector%20for%20Efficient%20Allelic%20Replacement%20in%20Naturally%20Nontransformable%2C%20Low-GC-Content%2C%20Gram-Positive%20Bacteria Arnaud ''et al''., 2004].</p> | ||
Revision as of 01:13, 27 September 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Sporobeads - what protein do you want to display?
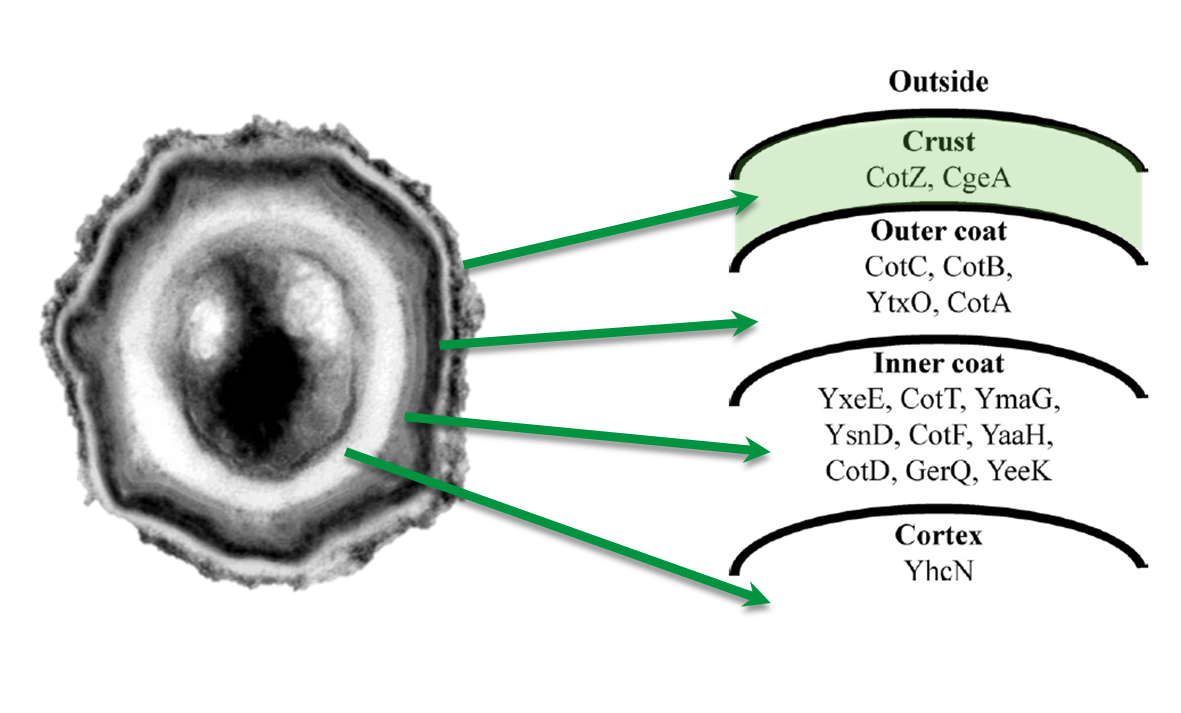
Sporobeads are Bacillus subtilis spores modulated in a way that they can be used as a platform for individual protein display. The aim of this module is to create these spores that display fusion proteins on their surface. There are several different proteins forming the spore coat layers of B. subtilis spores (Fig. 1). The outermost layer, the so-called spore crust, is composed of two proteins, CotZ and CgeA (Imamura et al., 2011). This is why we used them to create functional fusion proteins to be expressed on our Sporobeads.
|
The gene cgeA is located in the cgeABCDE cluster and is expressed from its own promoter PcgeA. The cluster cotVWXYZ contains the gene cotZ which is cotranscribed with cotY and expressed from the promoter PcotYZ. Another promoter of this cluster, PcotV, is responsible for the transcription of the other three genes (Fig. 2). Those three promoters were evaluated with the lux reporter genes (pSBBs3C-luxABCDE) to analyze their strength and the time point of their activation (see data).
|
Based on this data, two of the three promoters could be used for expression of spore crust fusion proteins (Fig. 3). First, we used gfp as a proof of principle and fused it to cgeA and cotZ. This way, we can determine if it is possible to display proteins on the spore crust and if their expression has any effect on spore formation.
|
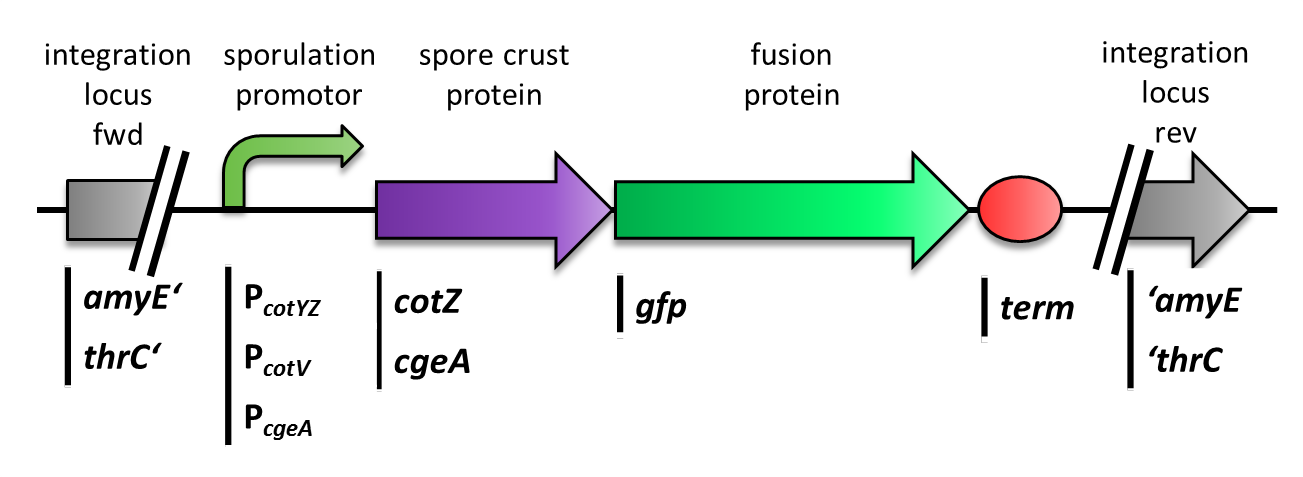
We constructed the BioBrick for cotZ, cgeA and gfp in Freiburg Standard. The cotZ gene was then fused to its two native promoters, PcotV and to PcotYZ, and PcgeA, which regulates the transcription of cgeA. For cgeA we only used its native promoter PcgeA and the stronger one of the two promoters of the cotVWXYZ cluster, PcotYZ (for more details see crust promotor evaluation. While gfp was ligated to the terminator B0014 (see Registry). When these constructs were finished and confirmed by sequencing, we fused them together by applying the Freiburg standard to create constructs, in which gfp is fused C-terminally to cotZ or cgeA flanked by one of the three promoters and the terminator. This way we created C-terminal fusion proteins (Fig. 4).
|
| |
|
But as we did not know if C- or N-terminal fusion would influence the fusion protein expression, our second aim was to construct N-terminal fusion proteins as well. For this purpose we wanted to fuse the genes for the crust proteins cotZ and cgeA to the terminator and gfp to the three chosen promoters. Unfortunately, there occured a mutation in the XbaI site during construction of gfp in Freiburg Standard. Therefore we were not able to finish these constructs. Finally we needed to clone our constructs into an empty Bacillus vector, so that they could get integrated into the genome of B. subtilis after transformation. For that purpose, we picked the empty vector pSBBS1C from our BacillusBioBrickBox, for the cotZ constructs. This vector integrates into the amyE locus, which allows to easily check the integration by starch test. In order to also express both crust protein constructs in one strain, the cgeA fusion proteins had to be cloned into another empty vector, pSBBS4S. Unfortunately, for unknown reasons, the cloning of the constructs with cgeA into this vector has so far not been successful.
But we were able to finish five constructs and integrated them into wild type W168 and the ΔcotZ mutant:
| recipient strain W168 | recipient strain B 49 (W168 ΔcotZ) | |
|---|---|---|
| pSBBs1C-PcotYZ-cotZ-2aa-gfp-terminator | B 53 | B 70 |
| pSBBs1C-PcotYZ-cotZ-gfp-terminator | B 54 | B 71 |
| pSBBs1C-PcotV-cotZ-2aa-terminator | B 55 | B 72 |
| pSBBs1C-PcotV-cotZ-terminator | B 56 | B 73 |
| pSBBs1C-PcgeA-cotZ-2aa-terminator | B 52 | B 69 |
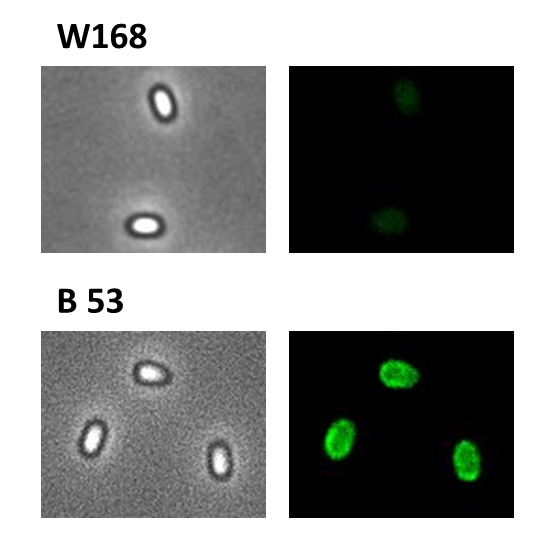
Finally, we started with the most important experiment for our GFP-Sporobeads, the fluorescence microscopy. We developed a sporulation protocol (for details see Protocol for enhancement of mature spore numbers) that increases the rates of mature spores in our samples. The cells were fixed on agarose pads and investigated by phase contrast and fluorescence microscopy. While spores of the wild type only showed the known background fluorescence, all Sporobeads showed bright green fluorescence at the edge (on the surface) of the spores. Sporobead from strain B 53 (containing the PcotYZ-cotZ-2aa-gfp-terminator construct) showed the highest fluorescence intensity (see Fig. 5 and all data). Hence, this strain was chosen for further experiments.
|
Since there were still some vegetative cells left after 24 hours of growth in Difco Sporulation Medium, we wanted to purify the Sporobeads. We tried three different methods for this approach: treatment with French Press, sonification and lysozyme. By means of the microscopy results we were able to conclude that lysozyme treatment was the only successful method (see data). Additionally, this treatment did not harm the crust fusion proteins as green fluorescence was still detectable afterwards (see data for details).
Moreover, we wondered if clean deletions of the native spore crust genes would show any difference in fusion protein expression in our Sporobeads. Thus, we deleted the native cotZ and cgeA using the pMAD based gene deletion strategy described by Arnaud et al., 2004.
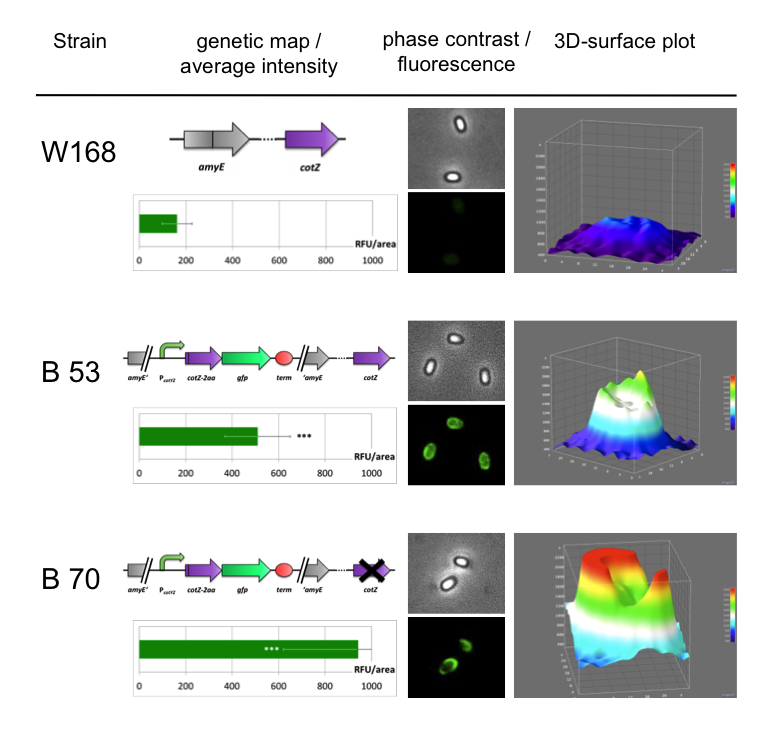
Because of the low but distinct fluorescence of wild type spores, we measured and compared the fluorescence intensity of 100 spores per construct (see data). We obtained significant differences between wild type spores and all of our Sporobeads (see Data). The intensity bar charts shown in Fig. 6 below the fluorescence difference between wild type (W168) and B53-/ B70-Sporobeads (B70 = B53 cotZ deletion). To demonstrate the distribution of the fusion proteins we created 3D graphs, which show the fluorescence intensity AUSTAUSCHEN VON DATENspread across the spore surface. For analysis we measured the fluorescence intensity of an area of 750px per spore by using ImageJ and evaluated it with the statistical software R. The following graph (Fig. 6) shows the results of microscopy and ImageJ analysis of the strongest construct integrated into wildtype W168 (B53) and the clean deletion mutant of cotZ (B 70).
|
| |
|
As shown in Fig. 6, the wild type spore has hardly any fluorescence, whereas both Sporobeads with the integrated construct pSBBs1C-PcotYZ-cotZ-2aa-gfp-terminator give a distinct fluorescence signal around the edge of the spore. Furthermore, it demonstrates that strain B 70 has the highest fluorescence intensity.
In summary we successfully developed functional sporobeads that are capable of displaying any protien of choice on the surface of modified B. subtilis endospores.

| 
| 
| 
|
| Bacillus Intro | Bacillus BioBrickBox | Sporobeads | Germination STOP |
 "
"