Team:LMU-Munich/Project
From 2012.igem.org
| Line 31: | Line 31: | ||
=== Evaluate Promoters === | === Evaluate Promoters === | ||
| - | Another goal is to produce promoters of ''Bacillus subtilis'' in BioBrick mode and to evaluate them. These well-defined promoters | + | Another goal is to produce promoters of ''Bacillus subtilis'' in BioBrick mode and to evaluate them. These well-defined promoters will then be part of the ''Bacillus'' BioBrickBox which we will send to the registry but they can also be useful in our one project to express our fusion crust proteins on the outside of our spores. Therefore we will use different promoters which are the constitutive promoters from the Anderson collection from the partsregistry, the constitutive promoters P''liaG'', P''veg'' and P''lepA'' from ''B. subtilis'' as well as the inducible promoter P''liaI'' from ''B. subtilis''. For the characterization of the different promoters we will clone them upstream of reporter genes (lux operon, lacZ) to measure their activity in ''Bacillus subtilis''. Therefore we use e.g. the two reporter vectors from ''B. subtilis'' which are also part of the ''Bacillus'' BioBrickBox. One of the reporter vectors pSB<sub>Bs</sub>3C-<i>luxABCDE</i> contains the lux operon as a reporter. This is why the activity of the promoter can be measured as luminescence with the plate reader (BioTek) which is a result of the expression of the luceiferase. The second reporter vector used for the evaluation of the promoters is pSB<sub>Bs</sub>1C-''lacZ'' which contains the reporter gene ''lacZ''. The promoter activity leads to the production of the enzyme beta-galactosidase which cleaves the substrate ONPG. The product is detectable with the photometer and refers to the promoter activity. |
| + | |||
| + | There are three types of promoters which will be evaluated in this project as a part of the so-called BioBrickBox: | ||
| + | |||
| + | The first group of promoters are the promoters of the Anderson collection which we call ''Anderson promoters''. They have already been measured in ''Escherichia coli'' and they all showed a constitutive behaviour but with a different strength. In this project these Anderson promoters were measured in ''B. subtilis''. As they have already been measured in ''E. coli'', they are already in the partsregistry in the vector where they are fused to the so-called ''Red fluorescent protein'' (RFP). For evaluation they are cut out of the vector and then cloned in the expression vector pSB<sub>Bs</sub>3C-<i>luxABCDE</i> from the BioBrickBox containing the ''lux'' operon as a reporter for promoter activity. The gene expression which correlates to the promoter activity leads to the expression of the lux operon with the luciferase. The luminescence which is produced by the luciferase can be measured with the plate reader (BioTek). | ||
| + | |||
| + | The second group of promoters are constitutive promoters from B. subtilis. We will evaluate the promoters PliaG, Pveg and PlepA. Therefore we will use both reporter vectors pSB<sub>Bs</sub>3C-<i>luxABCDE</i> and pSB<sub>Bs</sub>1C-''lacZ'' to evaluate the promoters with different reporters, the ''lux'' operon and the ''lacZ'' gene. Therefore the promoters will be amplified from the genom of B. subtilis with primers that conatin the restriction sites of the BioBrick standard. Then these constitutive promoters will be cloned into the empty vector pSB1C3 to send them to the registry as well as the two reporter vectors to evaluate their strength in ''B. subtilis''. | ||
| + | |||
| + | The last group of promoters consist of inducible promoters of ''B. subtilis'' e.g. P''<sub>liaI</sub>''. They are useful if you want to turn the system on in a certain moment because these promoters need an inducer to start transcription. P''<sub>liaI</sub>' can be induced by antibiotics which interact with the lipidII cycle, e.g. bacitracin. In this project this promoter is also evaltuated with the reporter vector pSB<sub>Bs</sub>3C-<i>luxABCDE</i> which contains the lux operon as a reporter and with the vector pSB<sub>Bs</sub>1C-''lacZ'' which conatins the ''lacZ'' reporter gene. Therefore the promoters will be amplified from the genom of B. subtilis with primers that conatin the restriction sites of the BioBrick standard. Then these constitutive promoters will be cloned into the empty vector pSB1C3 to send them to the registry as well as the two reporter vectors to evaluate their strength in ''B. subtilis''. | ||
| - | |||
| - | |||
=== Fuse Spore Coat Proteins === | === Fuse Spore Coat Proteins === | ||
Revision as of 15:23, 18 August 2012
The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Overall project
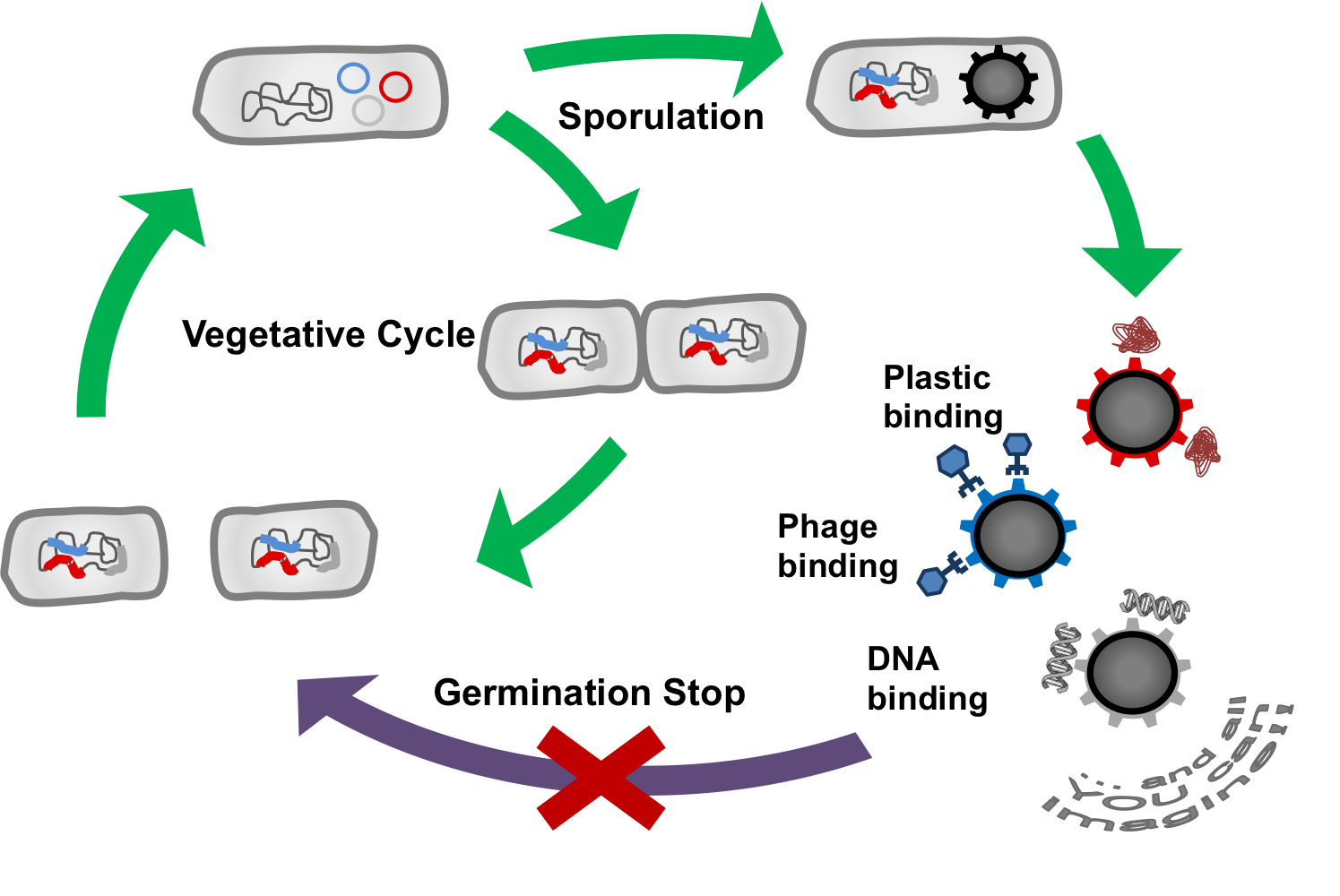
The goal of our project BEADzillus is the production of Bacillus subtilis spores which display different proteins with special features on their surface. For example, the spore could:
- bind harmful viruses
- bind toxic metals
- bind plastic molecules
- expose enzymes
These functions could be used to filter fluids or for use in laboratory.
As a safety consideration, we will stop spores from germinating. First, we want to knock out several genes which are important for germination from the spore phase; second, we want potentially germinating spores to be killed by a toxin which they produce themselves.
Project Details
Clone Vectors
Part of establishing Bacillus subtilis to the BioBrick mode is to modify standard vectors of Bacillus. We will mutate prohibited restriction sites (EcoRI, XbaI, SpeI, PstI) and excise redundant parts of the vectors.
Evaluate Promoters
Another goal is to produce promoters of Bacillus subtilis in BioBrick mode and to evaluate them. These well-defined promoters will then be part of the Bacillus BioBrickBox which we will send to the registry but they can also be useful in our one project to express our fusion crust proteins on the outside of our spores. Therefore we will use different promoters which are the constitutive promoters from the Anderson collection from the partsregistry, the constitutive promoters PliaG, Pveg and PlepA from B. subtilis as well as the inducible promoter PliaI from B. subtilis. For the characterization of the different promoters we will clone them upstream of reporter genes (lux operon, lacZ) to measure their activity in Bacillus subtilis. Therefore we use e.g. the two reporter vectors from B. subtilis which are also part of the Bacillus BioBrickBox. One of the reporter vectors pSBBs3C-luxABCDE contains the lux operon as a reporter. This is why the activity of the promoter can be measured as luminescence with the plate reader (BioTek) which is a result of the expression of the luceiferase. The second reporter vector used for the evaluation of the promoters is pSBBs1C-lacZ which contains the reporter gene lacZ. The promoter activity leads to the production of the enzyme beta-galactosidase which cleaves the substrate ONPG. The product is detectable with the photometer and refers to the promoter activity.
There are three types of promoters which will be evaluated in this project as a part of the so-called BioBrickBox:
The first group of promoters are the promoters of the Anderson collection which we call Anderson promoters. They have already been measured in Escherichia coli and they all showed a constitutive behaviour but with a different strength. In this project these Anderson promoters were measured in B. subtilis. As they have already been measured in E. coli, they are already in the partsregistry in the vector where they are fused to the so-called Red fluorescent protein (RFP). For evaluation they are cut out of the vector and then cloned in the expression vector pSBBs3C-luxABCDE from the BioBrickBox containing the lux operon as a reporter for promoter activity. The gene expression which correlates to the promoter activity leads to the expression of the lux operon with the luciferase. The luminescence which is produced by the luciferase can be measured with the plate reader (BioTek).
The second group of promoters are constitutive promoters from B. subtilis. We will evaluate the promoters PliaG, Pveg and PlepA. Therefore we will use both reporter vectors pSBBs3C-luxABCDE and pSBBs1C-lacZ to evaluate the promoters with different reporters, the lux operon and the lacZ gene. Therefore the promoters will be amplified from the genom of B. subtilis with primers that conatin the restriction sites of the BioBrick standard. Then these constitutive promoters will be cloned into the empty vector pSB1C3 to send them to the registry as well as the two reporter vectors to evaluate their strength in B. subtilis.
The last group of promoters consist of inducible promoters of B. subtilis e.g. PliaI. They are useful if you want to turn the system on in a certain moment because these promoters need an inducer to start transcription. PliaI' can be induced by antibiotics which interact with the lipidII cycle, e.g. bacitracin. In this project this promoter is also evaltuated with the reporter vector pSBBs3C-luxABCDE which contains the lux operon as a reporter and with the vector pSBBs1C-lacZ which conatins the lacZ reporter gene. Therefore the promoters will be amplified from the genom of B. subtilis with primers that conatin the restriction sites of the BioBrick standard. Then these constitutive promoters will be cloned into the empty vector pSB1C3 to send them to the registry as well as the two reporter vectors to evaluate their strength in B. subtilis.
Fuse Spore Coat Proteins
There are several different coat proteins in the spores of Bacillus subtilis. We chose CotZ and CgeA, which are located in the outermost layer of the spore coat. Our first step will be to fuse GFP to these proteins to see if they appear on the spore surface and if there is any effect on spore formation. The next step will be to fuse proteins with special features to CotZ and CgeA to produce functional "SporoBeads." SporoBeads could be used to filter fluids or in the laboratory, and could be capable of:
- binding harmful viruses
- binding toxic metals
- binding plastic molecules
- exposing enzymes
Stop Germination
To stop the SporoBeads from germinating, we have thought of two different manipulations. First, we will knock-out several genes which are necessary for all germination pathways, namely the cortex hydrolysis. Several gene deletion combinations will be tested for their outgrowth rates.
As a second mechanism to prevent germination, spores which begin to germinate despite the gene deletions shall be killed by an endotoxin system. Here, we will work with a sigma-factor from a different species which works in Bacillus subtilis but does not interfere with endogenous promoters. This sigma factor is placed downstream of a promoter that is strongly activated by sigmaG, the last active sigma factor in the endospore. Therefore, the alternative sigma factor is produced right before sporulation. The bacteria also have a second construct inserted into their genome: a promoter activated by the alternative sigma factor which leads to the transcription of an endotoxin. So, if a spore starts metabolism, the toxin gene will be activated and kill the germinating spore.
Results
 "
"