Team:LMU-Munich/Data/Constitutive
From 2012.igem.org
| Line 8: | Line 8: | ||
==Constitutive ''Bacillus'' Promoters== | ==Constitutive ''Bacillus'' Promoters== | ||
<br> | <br> | ||
| - | + | ===Luminescence measurements=== | |
| + | |||
<p align="justify"> | <p align="justify"> | ||
| - | The constitutive promoters '''P<sub>''liaG''</sub>''' and '''P<sub>''lepA''</sub>''' were evaluated in the reporter vector pSB<sub>Bs</sub>3C-<i>luxABCDE</i> which contains the ''lux'' operon [[File:Lux operon.png|100px]]. The promoter activity leads to gene expression and to the production of the protein luciferase. The luminescence produced by this protein can be measured with the plate reader ''Synergy2'' (Biotek) ('''Fig.1'''). All clones show a normal growth behaviour. The activity of both promoters increases during transition from log to stationary phase. P<sub>''liaG''</sub> has an activity maximum of about 100.000 Lumi/OD<sub>600</sub>. P<sub>''lepA''</sub> shows a maximum of about 400.000 Lumi/OD<sub>600</sub>. Comparing these two constitutive promoters the activity of P<sub>''lepA''</sub> is about four times higher than the activity of P<sub>''liaG''</sub>. In the late stationary phase the activity completely disappears. The second clone of the promoters P<sub>''lepA''</sub> and P<sub>''liaG''</sub> did not show any luminescence activity. Therefore additional clones should be measured.</p> | + | The constitutive promoters '''P<sub>''liaG''</sub>''' and '''P<sub>''lepA''</sub>''' were evaluated in the reporter vector pSB<sub>Bs</sub>3C-<i>luxABCDE</i> which contains the ''lux'' operon [[File:Lux operon.png|100px]]. The promoter activity leads to gene expression and to the production of the protein luciferase. The luminescence produced by this protein can be measured with the plate reader ''Synergy2'' (Biotek) ('''Fig.1'''). </p> |
| + | |||
| + | {| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | ||
| + | | style="width: 70%;background-color: #EBFCE4;" | | ||
| + | {| | ||
| + | |[[File:Auswertung_plate_reader_andere_promotoren.png|400px]] | ||
| + | |- | ||
| + | | style="width: 70%;background-color: #EBFCE4;" | | ||
| + | {| style="color:black;" cellpadding="0" width="100%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
| + | |style="width: 70%;background-color: #EBFCE4;" | | ||
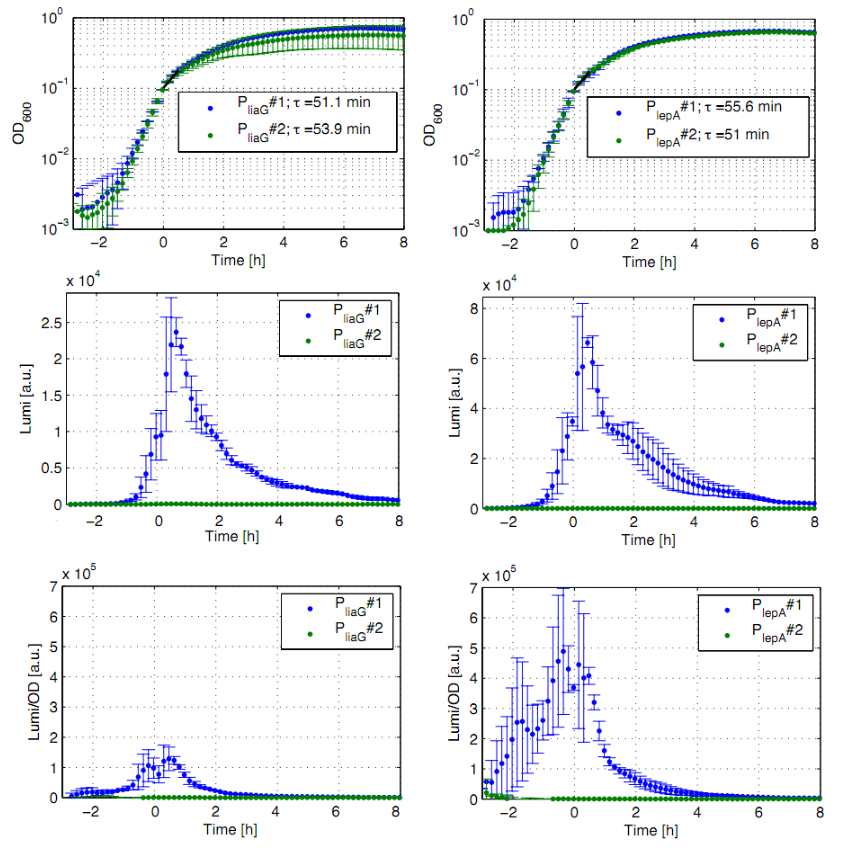
| + | <font color="#000000"; size="2"><p align="justify">'''Fig. 1: Luminescence measurement of the constitutive ''Bacillus'' promoters P<sub>''liaG''</sub> and P<sub>''lepA''</sub> in the reporter vector pSB<sub>''Bs''</sub>3C-''luxABCDE'''''. OD<sub>600</sub> (up), LUMI (middle) and LUMI per OD<sub>''600''</sub> (down) depending on the time (h) are shown for two different clones (green/blue). Data derive from three independent experiments, the graphs show the mean fo the three experiments and the standard deviation. Curves were fitted over each other (t=0, OD<sub>''600''</sub>=0,3) and smoothed by taking the average of three neighboring values.</p></font> | ||
| + | |} | ||
| + | |} | ||
| + | |} | ||
| + | |||
| + | <p align="justify">All clones show a normal growth behaviour. The activity of both promoters increases during transition from log to stationary phase. P<sub>''liaG''</sub> has an activity maximum of about 100.000 Lumi/OD<sub>600</sub>. P<sub>''lepA''</sub> shows a maximum of about 400.000 Lumi/OD<sub>600</sub>. Comparing these two constitutive promoters the activity of P<sub>''lepA''</sub> is about four times higher than the activity of P<sub>''liaG''</sub>. In the late stationary phase the activity completely disappears. The second clone of the promoters P<sub>''lepA''</sub> and P<sub>''liaG''</sub> did not show any luminescence activity. Therefore additional clones should be measured.</p> | ||
| + | ===β-galactosidase assay=== | ||
[[File:Englisch_Auswertung_PliaG_Pveg.png|thumb|right|400px| <p align="justify"> '''Fig. 2: β-galactosidase assay and growth curve of strains carrying the promoters P<sub>''liaG''</sub> (black) and P<sub>''veg''</sub> (grey) fused to ''lacZ'''''. β-galactosidase activity (Miller Units) and the growth curve values are the average of two independent clones with their standard deviation. Experiment shows representative data from three independent experiments.</p>]] | [[File:Englisch_Auswertung_PliaG_Pveg.png|thumb|right|400px| <p align="justify"> '''Fig. 2: β-galactosidase assay and growth curve of strains carrying the promoters P<sub>''liaG''</sub> (black) and P<sub>''veg''</sub> (grey) fused to ''lacZ'''''. β-galactosidase activity (Miller Units) and the growth curve values are the average of two independent clones with their standard deviation. Experiment shows representative data from three independent experiments.</p>]] | ||
Revision as of 17:24, 25 September 2012
The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Constitutive Bacillus Promoters
Luminescence measurements
The constitutive promoters PliaG and PlepA were evaluated in the reporter vector pSBBs3C-luxABCDE which contains the lux operon ![]() . The promoter activity leads to gene expression and to the production of the protein luciferase. The luminescence produced by this protein can be measured with the plate reader Synergy2 (Biotek) (Fig.1).
. The promoter activity leads to gene expression and to the production of the protein luciferase. The luminescence produced by this protein can be measured with the plate reader Synergy2 (Biotek) (Fig.1).
All clones show a normal growth behaviour. The activity of both promoters increases during transition from log to stationary phase. PliaG has an activity maximum of about 100.000 Lumi/OD600. PlepA shows a maximum of about 400.000 Lumi/OD600. Comparing these two constitutive promoters the activity of PlepA is about four times higher than the activity of PliaG. In the late stationary phase the activity completely disappears. The second clone of the promoters PlepA and PliaG did not show any luminescence activity. Therefore additional clones should be measured.
β-galactosidase assay
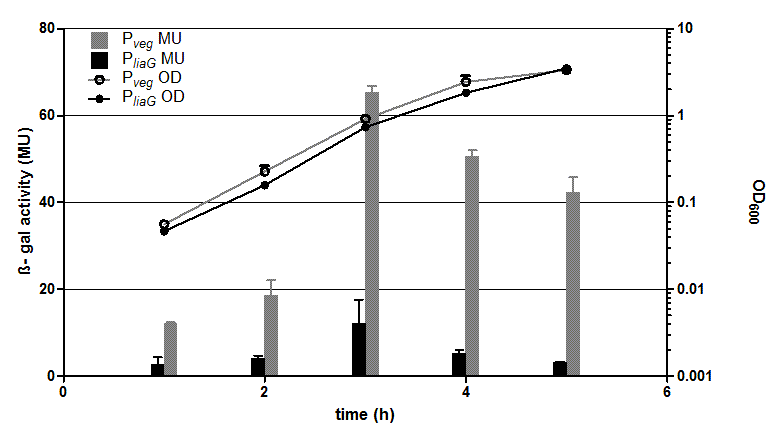
Fig. 2: β-galactosidase assay and growth curve of strains carrying the promoters PliaG (black) and Pveg (grey) fused to lacZ. β-galactosidase activity (Miller Units) and the growth curve values are the average of two independent clones with their standard deviation. Experiment shows representative data from three independent experiments.
The two constitutive promoters PliaG and Pveg were evaluated with the reporter vector pSBBs1C-lacZ which contains the lacZ reporter gene ![]() (Fig.2). Promoter activity leads to the expression of the β-galactosidase. The β-galactosidase assay of the constitutive Bacillus promoters Pveg and PliaG was repeated three times. The graph shows data of one representative experiment. In the beginning of the growth curve both promoters show only low activity before it increases to a maximum before it decreases to the begininng level after about seven hours (Data not shown). Summing up, the course of activity of both promoters Pveg and PliaG is very similar based on the growth curve. The highest β-galactosidase activity and therefore the highest activity of the promoter Pveg with a maximum of 65 Miller units can be found during the transition from the logarithmic to the stationary phase. This is about five times higher than the acitivity of the promoter PliaG with a maximum activity of about 12 Miller Units.
(Fig.2). Promoter activity leads to the expression of the β-galactosidase. The β-galactosidase assay of the constitutive Bacillus promoters Pveg and PliaG was repeated three times. The graph shows data of one representative experiment. In the beginning of the growth curve both promoters show only low activity before it increases to a maximum before it decreases to the begininng level after about seven hours (Data not shown). Summing up, the course of activity of both promoters Pveg and PliaG is very similar based on the growth curve. The highest β-galactosidase activity and therefore the highest activity of the promoter Pveg with a maximum of 65 Miller units can be found during the transition from the logarithmic to the stationary phase. This is about five times higher than the acitivity of the promoter PliaG with a maximum activity of about 12 Miller Units.
 "
"