Team:LMU-Munich/Bacillus Introduction
From 2012.igem.org
| Line 69: | Line 69: | ||
There are two major differences between ''B. subtilis'' and ''E. coli'' that are of interest to us: | There are two major differences between ''B. subtilis'' and ''E. coli'' that are of interest to us: | ||
| - | + | ||
'''1) Transformation of ''B. subtilis''''' | '''1) Transformation of ''B. subtilis''''' | ||
| - | + | <p align="justify">As ''B. subtilis'' and ''E. coli'' are model organisms, they have established genetics. The advantage of ''B. subtilis'' is that it is naturally competent. So it is very easy to conduct genetic manipulations. It can replicate plasmids as ''E. coli'' does, but there is a much more elegant way of bringing in exogenous DNA fragments. When flanked by regions homologous to the ''B. subtilis'' genome, it will be integrated at high efficiency via homologous recombination at this locus and subsequently be replicated with the chromosome (Fig. 2). This leads to stable, single-copy genomic alterations. Thereby avoiding, copy-number artifacts occuring with replicative plasmids. This different way of genetic manipulations requires the use of integrative vectors as provided by our [https://2012.igem.org/Team:LMU-Munich/Bacillus_BioBricks BacillusBioBrickBox]. For this reason, ''B. subtilis'' is an ideal genetic platform for Synthetic Bioloy. But so far, very few iGEM teams have worked with this model organism due to the lack of suitable BioBrick-compatible genetic tools.</p> | |
<br> | <br> | ||
| Line 95: | Line 95: | ||
<br> | <br> | ||
'''2) Differentiation''' | '''2) Differentiation''' | ||
| - | + | <p align="justify">''B. subtilis'' is able to differentiate into cells with different morphologies and functions (Fig. 1 and Fig. 3). The most characteristic form is the endospore, which is produced under nutrient starvation. In our project, we will exploit the production of endospores. Because they are extremely stable, they are perfect vehicles for the display of functional fusion proteins on their surface as illustrated by our [https://2012.igem.org/Team:LMU-Munich/Spore_Coat_Proteins '''Sporo'''bead] module. | |
<br> | <br> | ||
</p> | </p> | ||
Revision as of 01:33, 27 September 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

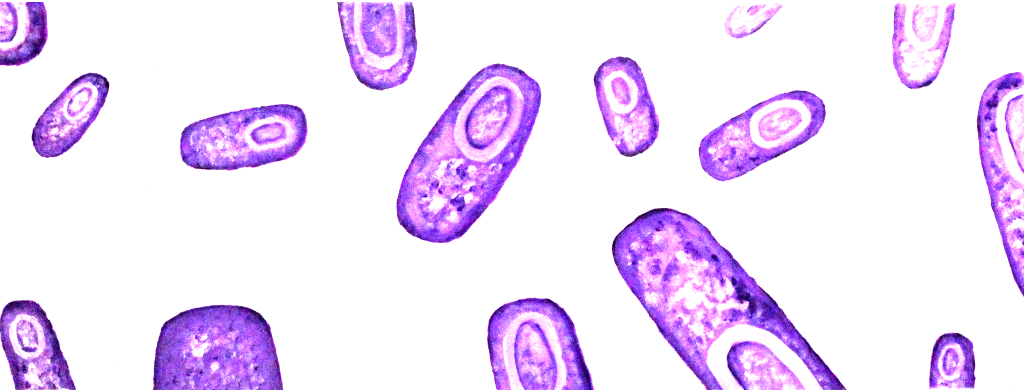
Bacillus subtilis - a new chassis for iGEM
We chose to work with Bacillus subtilis to set new horizons in the Escherichia coli-dominated world of iGEM by offering a set of tool for this Gram-positive model organism to the . To introduce B. subtilis, we first want to highlight some important aspects of this organism, which are listed in the table below.
| Organism | B. subtilis | E. coli |
|---|---|---|
| Type | Gram-positive rod | Gram-negative rod |
| Motility | + | + |
| Transformation | natural competence | artificial competence |
| Vectors | integrative & replicative | replicative |
| Sporulation | + | - |
| Differentiation | + | - |
| Safety | GRAS | GRAS |
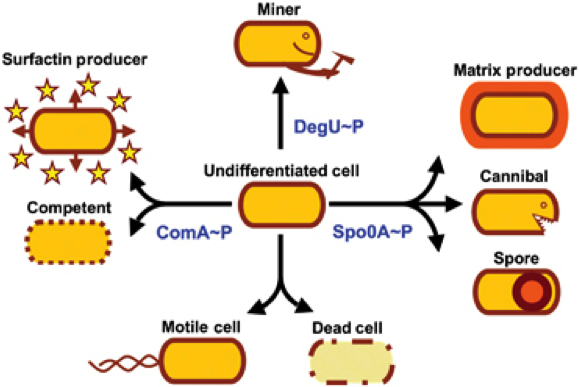
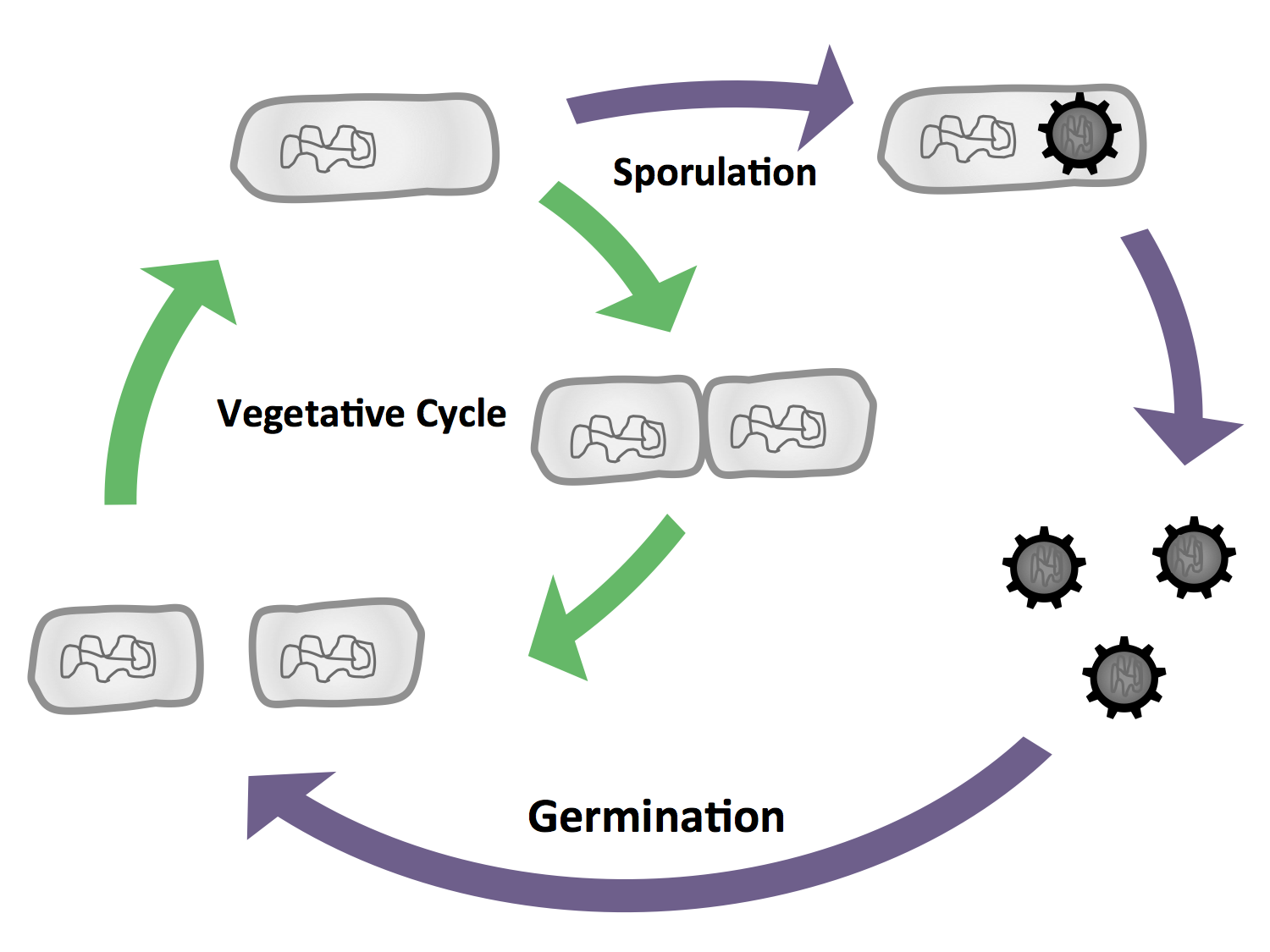
In general, bacteria can be divided into two major groups that differ primarily in their cell envelope: Gram-positive and Gram-negative. E. coli is the model organism for the Gram-negative bacteria. A model organism for the Gram-positive bacteria is B. subtilis, our favourite pet. The natural habitat of B. subtilis is the soil, so it is forced to adapt to numerous environmental changes. Accordingly, B. subtilis is characterized by quick and cunning reflexes and a complex lifestyle. There are many differentiations and survival strategies that B. subtilis can engage (Fig. 1): Due to its natural competence, it can take up exogenous DNA and integrate it into its genome. To be flexible to the environment and move towards nutrients or avoid toxic agents, it is motile with the aid of its peritrichous flagella. If starved some cells even become cannibals that feast on their siblings. If the conditions get too bad for survival, B. subtilis can form endospores. These are very dormant and highly stable stages that are resistant against e.g. desiccation, UV light, heat and pressure. Once these spores encounter better conditions they are able to germinate again. See this section for Details on germination.
|
There are two major differences between B. subtilis and E. coli that are of interest to us:
1) Transformation of B. subtilis
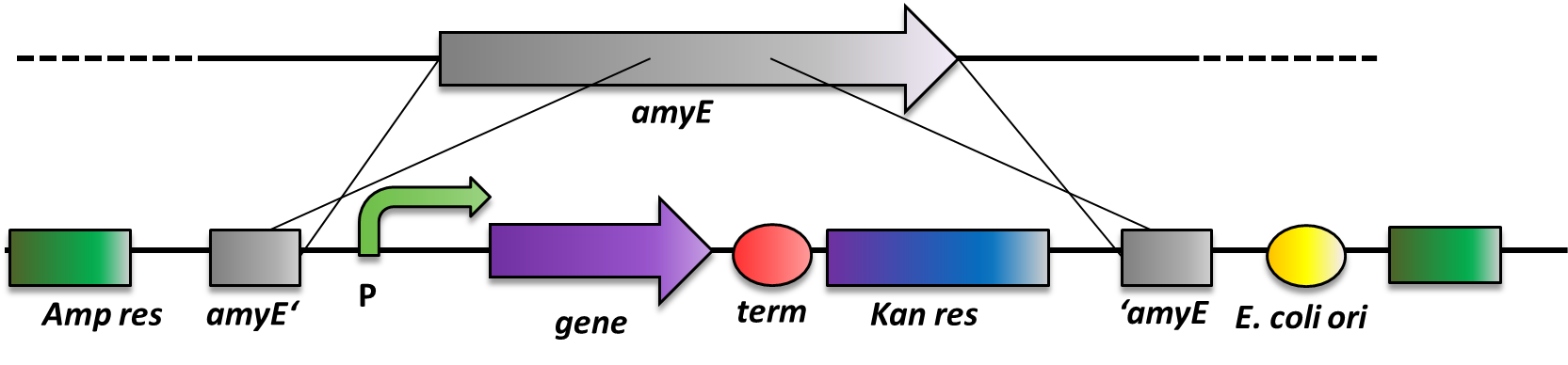
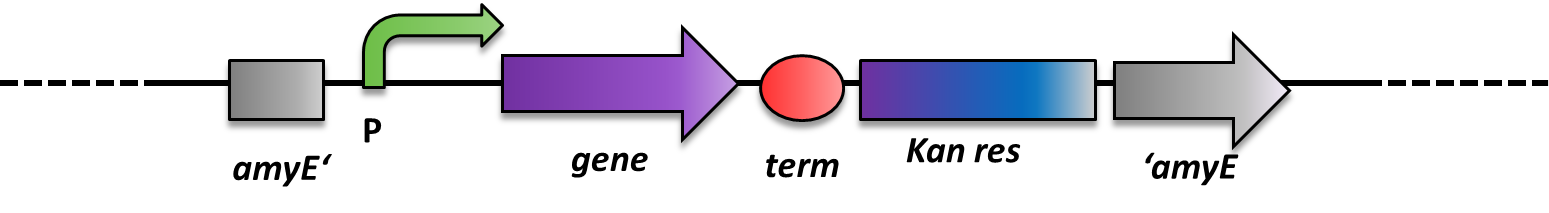
As B. subtilis and E. coli are model organisms, they have established genetics. The advantage of B. subtilis is that it is naturally competent. So it is very easy to conduct genetic manipulations. It can replicate plasmids as E. coli does, but there is a much more elegant way of bringing in exogenous DNA fragments. When flanked by regions homologous to the B. subtilis genome, it will be integrated at high efficiency via homologous recombination at this locus and subsequently be replicated with the chromosome (Fig. 2). This leads to stable, single-copy genomic alterations. Thereby avoiding, copy-number artifacts occuring with replicative plasmids. This different way of genetic manipulations requires the use of integrative vectors as provided by our BacillusBioBrickBox. For this reason, B. subtilis is an ideal genetic platform for Synthetic Bioloy. But so far, very few iGEM teams have worked with this model organism due to the lack of suitable BioBrick-compatible genetic tools.
B. subtilis is able to differentiate into cells with different morphologies and functions (Fig. 1 and Fig. 3). The most characteristic form is the endospore, which is produced under nutrient starvation. In our project, we will exploit the production of endospores. Because they are extremely stable, they are perfect vehicles for the display of functional fusion proteins on their surface as illustrated by our Sporobead module.
|
 "
"