Team:LMU-Munich/Bacillus Introduction
From 2012.igem.org
| Line 53: | Line 53: | ||
Visit our [[Team:LMU-Munich/Lab_Notebook/Protocols|Protocols page]] for details how to work with ''B. subtilis'' | Visit our [[Team:LMU-Munich/Lab_Notebook/Protocols|Protocols page]] for details how to work with ''B. subtilis'' | ||
| + | |||
| + | <p align="justify"> | ||
| + | There are two major differences between ''B. subtilis'' and ''E. coli'' that are of interest to us: | ||
| + | <br> | ||
| + | <br> | ||
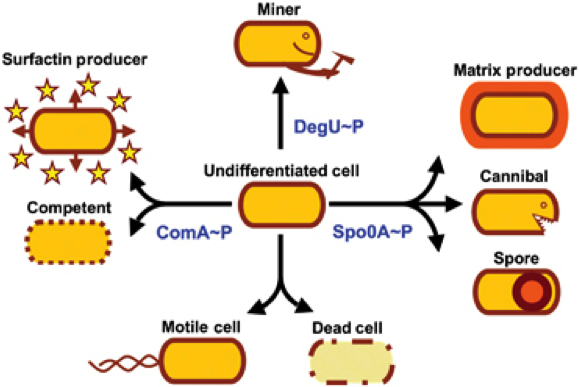
| + | '''1.''' ''B. subtilis'' is able to differentiate into cells with different morphology and function ('''Fig. 1'''), the most severe form being the endospore which is formed under stress conditions. But there are also phenomena like cannibalism which makes ''B. subtilis'' a lot more diverse to work with. We will exploit the production of endospores in our project '''Bead'''zillus. The life cycle of ''B. subtilis'' is depicted in '''Fig. 2'''. | ||
| + | <br> | ||
| + | </p> | ||
{| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | {| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | ||
| style="width: 70%;background-color: #EBFCE4;" | | | style="width: 70%;background-color: #EBFCE4;" | | ||
{| | {| | ||
| - | |[[File:LMU Bacillus life.png|300px||center| | + | |[[File:LMU Bacillus life.png|300px||center|300px]] |
|- | |- | ||
| style="width: 70%;background-color: #EBFCE4;" | | | style="width: 70%;background-color: #EBFCE4;" | | ||
{| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
| - | <font color="#000000"; size="2"></font> | + | <font color="#000000"; size="2">Fig. 1:</font> |
|} | |} | ||
|} | |} | ||
| Line 74: | Line 82: | ||
{| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
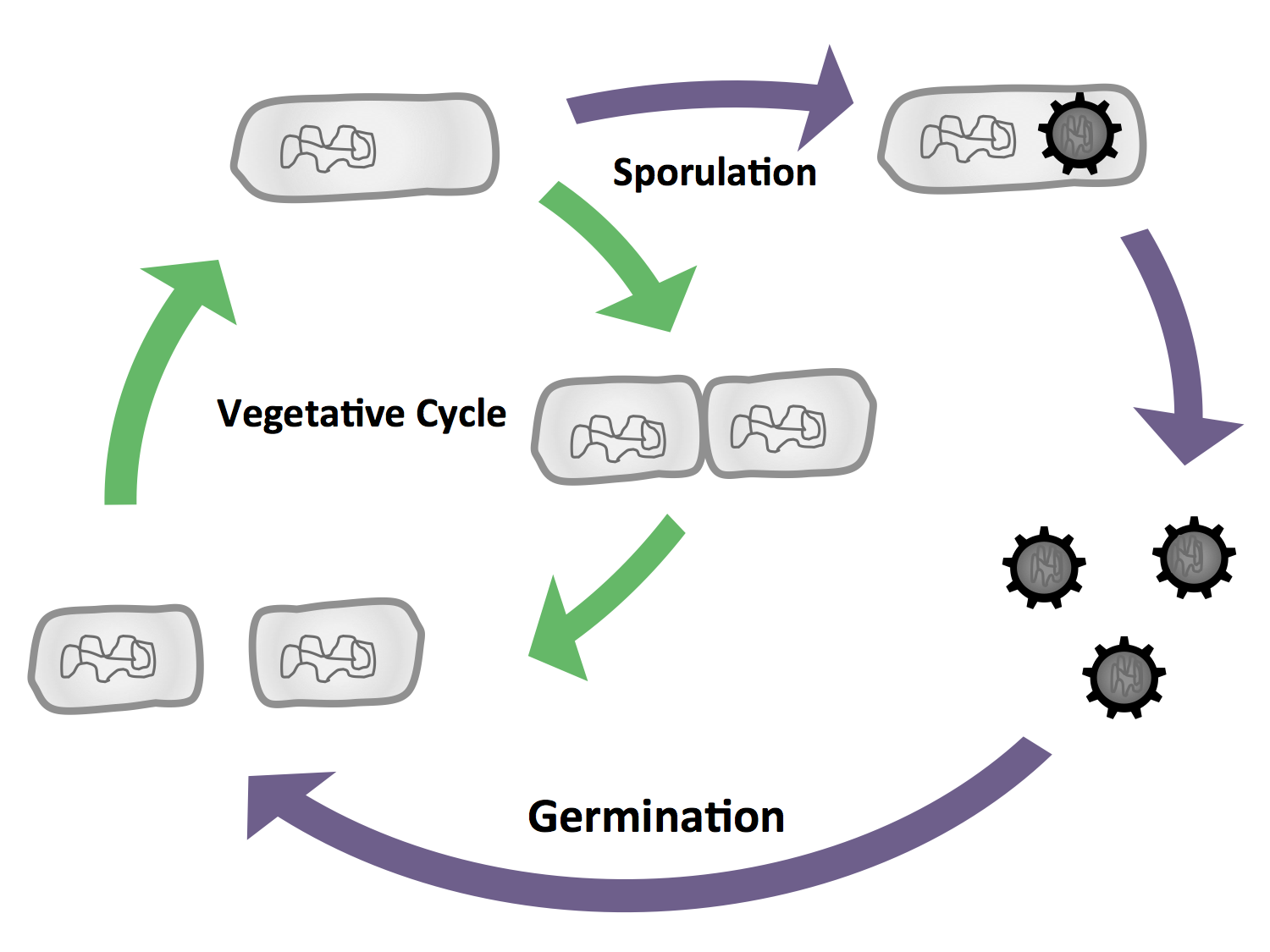
| - | <font color="#000000"; size="2"><p align="justify">''' | + | <font color="#000000"; size="2"><p align="justify">'''Fig. 2:''' The vegetative cycle is very similiar to the one of ''E. coli.'' But if there is a stress condition like starvation, the cells enter sporulation, where they first undergo a polar cell division, followed by the formation of the endospore. If the enviromental conditions are suitable again, the spore will then germinate and reenter the vegetative cycle.</p></font> |
|} | |} | ||
|} | |} | ||
|} | |} | ||
| - | + | <br> | |
| + | <p align="justify"> | ||
| + | '''2.''' ''B. subtilis'' can replicate exogenic DNA via an origin of replication on a plasmid as ''E. coli'' does, but there is a much more elegant way of bringing in exogenic DNA stretches. When flanked by homologous regions to the bacterial genome, it will integrate at high efficiency via homologous recombination at this locus and furthermore be replicated with the genome. This has the advantage that if comparing different things, not only the enviroment is always the same, but also the copy number is from cell to cell and from strain to strain the same, which is not always the case for replicative plasmids. This integrative way of bringing in exogenic DNA will be exploited by us when producing the BioBrick compatible ''Bacillus'' vectors. The comparision between these two ways of bringing in exogenic DNA is depicted in Figure 2. | ||
| + | <br> | ||
| + | For these reasons in some cases ''B. subtilis'' can be the chassis of choice. Sadly very few teams have worked with this model organism and there is at this time no established BioBrick system to use ''B. subtilis'' as a chassis. | ||
| + | </p> | ||
| + | <br> | ||
| + | <br> | ||
{| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | {| style="color:black;" cellpadding="3" width="70%" cellspacing="0" border="0" align="center" style="text-align:left;" | ||
| Line 89: | Line 104: | ||
{| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | {| style="color:black;" cellpadding="0" width="95%" cellspacing="0" border="0" align="center" style="text-align:center;" | ||
|style="width: 70%;background-color: #EBFCE4;" | | |style="width: 70%;background-color: #EBFCE4;" | | ||
| - | <font color="#000000"; size="2"><p align="justify">''' | + | <font color="#000000"; size="2"><p align="justify">'''Fig. 3:''' Exogenic DNA is shown in red while the bacterial genome is black a) The propagation of exogenic DNA if brought in as replicative plasmid. The number of plasmids per cell can vary. b) The propagation of exogenic DNA if it is able to integrate into the genome via homologous recombination</p></font> |
|} | |} | ||
|} | |} | ||
| Line 96: | Line 111: | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
Revision as of 14:30, 26 September 2012

The LMU-Munich team is exuberantly happy about the great success at the World Championship Jamboree in Boston. Our project Beadzillus finished 4th and won the prize for the "Best Wiki" (with Slovenia) and "Best New Application Project".
[ more news ]

Bacillus subtilis - a new chassis for iGEM
We chose to work with Bacillus subtilis to set new horizons and offer tools for this model organism to the Escherichia coli-dominated world of iGEM. In addition, this organism produces spores which we use for our Sporobeads. To introduce B. subtilis to the iGEM world we want to present you some important aspect of this organism in contrast to the commonly used Escherichia coli.
| Organism | B. subtilis | E. coli |
|---|---|---|
| type | gram positive rod | gram negative rod |
| Motility | pos | pos |
| Transformation | natural competence | artificial competence |
| Vectors | integrative and replicative | replicative |
| Sporulation | pos | neg |
| Differentiation | pos | neg |
| Safety | GRAS | GRAS |
In contrast to E. coli, the model organism B. subtilis is a gram-postivite rod. It is a facultative aerobe soil bacterium which can move with his peritrichous (?). Under very good living conditions it has a doubling time of 45 minutes. B. subtilis is, in contrast to E. coli, naturally competent, what means that e.g. under nutrient limitation 10% cells of a populaton get competent and take up DNA. Also unique for B. subtilis is that the vegetative cells can differentiate under nutrient limitations in spores which we use in our project Beadzillus. Concerning the work with B. subtilis it is important to know that there are integrative vectors which can integrate into the genome of the organism in contrast to E. coli where there are only replicative vectors.
Visit our Protocols page for details how to work with B. subtilis
There are two major differences between B. subtilis and E. coli that are of interest to us:
1. B. subtilis is able to differentiate into cells with different morphology and function (Fig. 1), the most severe form being the endospore which is formed under stress conditions. But there are also phenomena like cannibalism which makes B. subtilis a lot more diverse to work with. We will exploit the production of endospores in our project Beadzillus. The life cycle of B. subtilis is depicted in Fig. 2.
|
|
2. B. subtilis can replicate exogenic DNA via an origin of replication on a plasmid as E. coli does, but there is a much more elegant way of bringing in exogenic DNA stretches. When flanked by homologous regions to the bacterial genome, it will integrate at high efficiency via homologous recombination at this locus and furthermore be replicated with the genome. This has the advantage that if comparing different things, not only the enviroment is always the same, but also the copy number is from cell to cell and from strain to strain the same, which is not always the case for replicative plasmids. This integrative way of bringing in exogenic DNA will be exploited by us when producing the BioBrick compatible Bacillus vectors. The comparision between these two ways of bringing in exogenic DNA is depicted in Figure 2.
For these reasons in some cases B. subtilis can be the chassis of choice. Sadly very few teams have worked with this model organism and there is at this time no established BioBrick system to use B. subtilis as a chassis.
|

| 
| 
| 
|
| Bacillus Intro | Bacillus BioBrickBox | Sporobeads | Germination STOP |
 "
"