Team:HokkaidoU Japan/Notebook/aggregation Week 9
From 2012.igem.org
August 27th
Colony PCR
Colony PCR to confirm that whether the pT7-RBS-Ag43-dT on pSB1C3 was successfully ligated or not.
| DNA solution | 4 ul |
| Kapa-Taq(Taq polymerase) | 5 ul |
| Forward Primer(ag43-f4 primer) | 0.5 ul |
| Reverse Primer(200bp down primer) | 0.5 ul |
| Total | 10 ul |
| Number | Degree | Second |
| 1 | 95 | 120 |
| 2 | 95 | 30 |
| 3 | 53.0 | 30 |
| 4 | 72 | 60 |
| 5 | 72 | 60 |
| 6 | 4 | HOLD |
Cycle:2~4 x 35
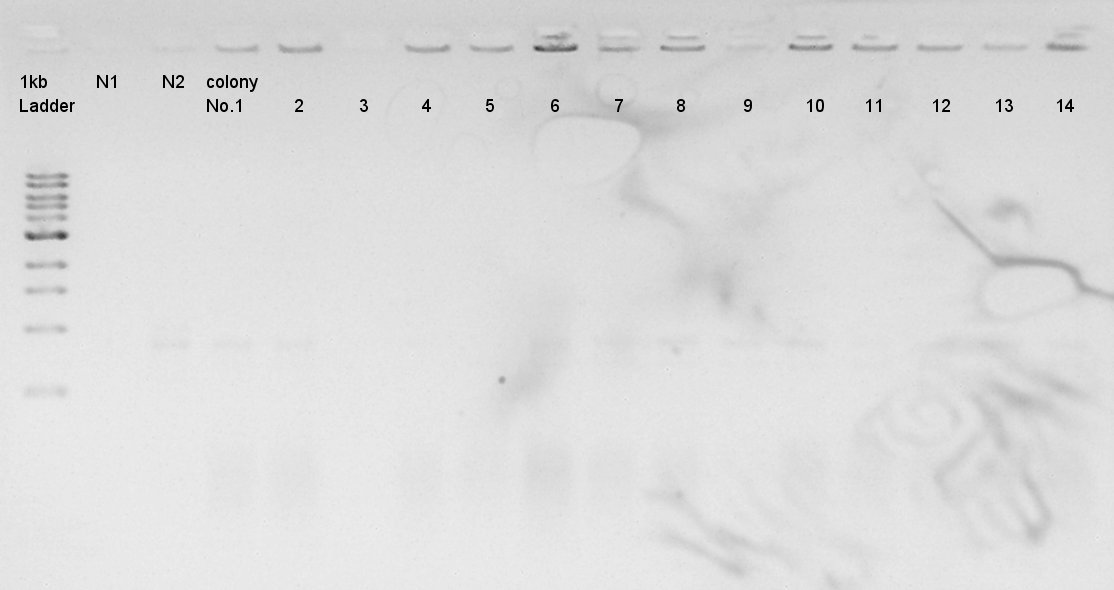
We used N1 (DW only) and N2 (Ag43-dT on pSB1AK3) as controls. Desired product is about 695bp.
The result did not show the band clearly. We selected No.1 and 2 colony for incubation.
Incubation for mini-prep of pT7-RBS-Ag43-dT on pSB1C3
Incubation of pT7-RBS-Ag43-dT on pSB1C3 in LBC liquid medium.
- Prepared 2 ml LBC into culture tubes.
- Re-suspended 2 colonies (No.1 and No.2 respectively).
- Incubated at 37C for 15 hrs.
Transformation of ptet-RBS(B0034)-CFP-dT on pSB1A2
- Mixed 1 ul DNA to 50 ul of thawed competent cells on ice.
- Incubated on ice for 30 min.
- Mixed 350 ul of LB.
- Prepared and Labeled two plastic plates with LBC.
- Plated 300 ul of the culture onto first dish and spread.
- Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread.
- Incubated the plates at 37C for 15 hours.
PCR of RBS-YFP-dT
Amplified the construct with 100bp-up-EX primer and 200bp-down-PS primer. Mixed PCR solutions.
| Solution | Volume(ul) |
| DNA | 1 |
| 100bp-up-EX | 1 |
| 200bp-down-PS | 1 |
| MgSO4 | 3 |
| dNTP | 5 |
| 10x KOD-Plus-Neo Buffer | 5 |
| KOD-Plus-Neo | 1 |
| DW | 33 |
| Total | 50 |
| Number | Degree | Second |
| 1 | 94 | 120 |
| 2 | 98 | 10 |
| 3 | 58.2 | 30 |
| 4 | 68 | 60 |
| 5 | 4 | HOLD |
Cycle:2~4 x 35
August 28th
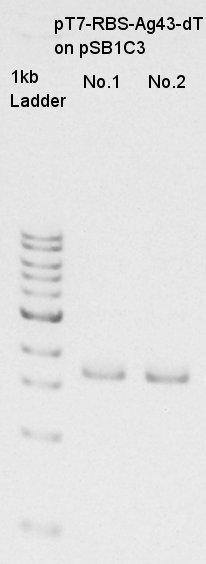
mini-prep of pT7-RBS-Ag43-dT on pSB1C3
Mini-prep of pT7-RBS-Ag43-dT on pSB1C3. We re-suspended No.1,2 colonies and incubated.
Estimation of Concentration of RBS-eYFP-dT and ptetR-pSB1A2
| Solution | Value(ul) |
| 1kb ladder | 1 |
| Insert | 1 |
| Vector | 1 |
From this result, We estimated that the concentration of Insert DNA solution is about 40ng/ul, and Vector DNA is also about 40ng/ul.
Digestion of ptet-pSB1A2 and RBS-Ag43-dT
ptet-pSB1A2(Vector) was digested with SpeI and PstI, and RBS-Ag43-dT(Insert) was cut with XbaI and PstI. Vector
| DNA solution ( 40ng/ul) | 5 ul |
| SpeI | 0.5 ul |
| PstI | 0.5 ul |
| 10xH buffer | 2 ul |
| DW | 12 ul |
| Total | 20 ul |
Inset
| DNA solution ( 40ng/ul) | 14 ul |
| XbaI | 1 ul |
| PstI | 1 ul |
| 10xM buffer | 2 ul |
| DW | 2 ul |
| Total | 20 ul |
| Number | Degree | Minute |
| 1 | 37 | 180 |
| 2 | 60 | 15 |
| 3 | 4 | HOLD |
From this result, we confirmed that Insert and Vector DNA were digested.
Ethanol Precipitation
Ethanol precipitation for ptet-pSB1A2 and RBS-eYFP-dT digestion products.
- Added 5 ul of NaoAc, 1.5 ul of glycogen and 125 ul of 100% ethanol.
- Centrifuged in 15000 rpm, 15 min at 4C.
- Remove supernatant and added 220 ul of 70% ethanol.
- Centrifuged in 15000 rpm, 10 min at 4C.
- Remove supernatant and air drying in room temperature then added 10 ul of DW.
Ligation
Ligation for ptet-pSB1A2 and RBS-eYFP-dT. We used Ligation Mighty Mix (TAKARA BIO INC.) which contains ligase and buffer.
| Vector DNA | 3 ul |
| Insert DNA | 3 ul |
| Ligation Mighty Mix | 6 ul |
| Total | 12 ul |
Ligation reaction time was in detail below.
| Degree | Minute |
| 16 | 30 |
| 65 | 10 |
| 4 | Hold |
Transformation
Transformation for ligation product (ptet-RBs-eYFP-dT on pSB1A2). We used E.coli strain DH5α.
- Mixed 1 ul of DNA to 50 ul of thawed competent cells on ice.
- Incubated on ice for 30 min.
- Mixed 350 ul of LB.
- Prepared and Labeled two plastic plates with LBA.
- Plated 300 ul of the culture onto first dish and spread.
- Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread.
- Incubated the plates at 37C for 15 hours.
Gel extraction product check of pT7-RBS-Ag43-dT on pSB1C3
We couldn't get desired plasmid DNA by transformation. We doubted contamination of non-digested vector DNA and decided to test the gel extraction product of pT7-RBS on pSB1C3 was successfully separated from non-digested product or not by transformation.
- Mixed 1 ul of each DNA of ligation product and digestion product to 50 ul of thawed competent cells on ice.
- Incubated on ice for 30 min.
- Mixed 350 ul of LB.
- Incubated for 2 hrs to get the resistance to Chloramphenicol.
- Prepared and Labeled two plastic plates with LBC.
- Plated 300 ul of the culture onto first dish and spread.
- Mixed 450 ul of LB to 50 ul of the culture and plated 300 ul of it onto second dish and spread.
- Incubated the plates at 37C for 16 hours.
August 29th
PCR of eCFP(E0020)
Amplified the part with 100bp-up-EX primer and 200bp-down-PS primer. Desired product is about 800~900bp. Mixed PCR solutions.
| Solution | Volume(ul) |
| DNA | 1 |
| 100bp-up-EX | 1 |
| 200bp-down-PS | 1 |
| MgSO4 | 3 |
| dNTP | 5 |
| 10x KOD-Plus-Neo Buffer | 5 |
| KOD-Plus-Neo | 1 |
| DW | 33 |
| Total | 50 |
| Number | Degree | Second |
| 1 | 94 | 120 |
| 2 | 98 | 10 |
| 3 | 58.2 | 30 |
| 4 | 68 | 60 |
| 5 | 4 | HOLD |
Cycle:2~4 x 35
Colony PCR
Colony PCR of pT7-RBS-Ag43-dT on pSB1C3 transformed in 28th.
| DNA solution | 4 ul |
| Kapa-Taq(Taq polymerase) | 5 ul |
| Forward Primer(ag43-f4 primer) | 0.5 ul |
| Reverse Primer(200bp down primer) | 0.5 ul |
| Total | 10 ul |
| Number | Degree | Second |
| 1 | 95 | 120 |
| 2 | 95 | 30 |
| 3 | 53.0 | 30 |
| 4 | 72 | 60 |
| 5 | 72 | 60 |
| 6 | 4 | HOLD |
Cycle:2~4 x 35
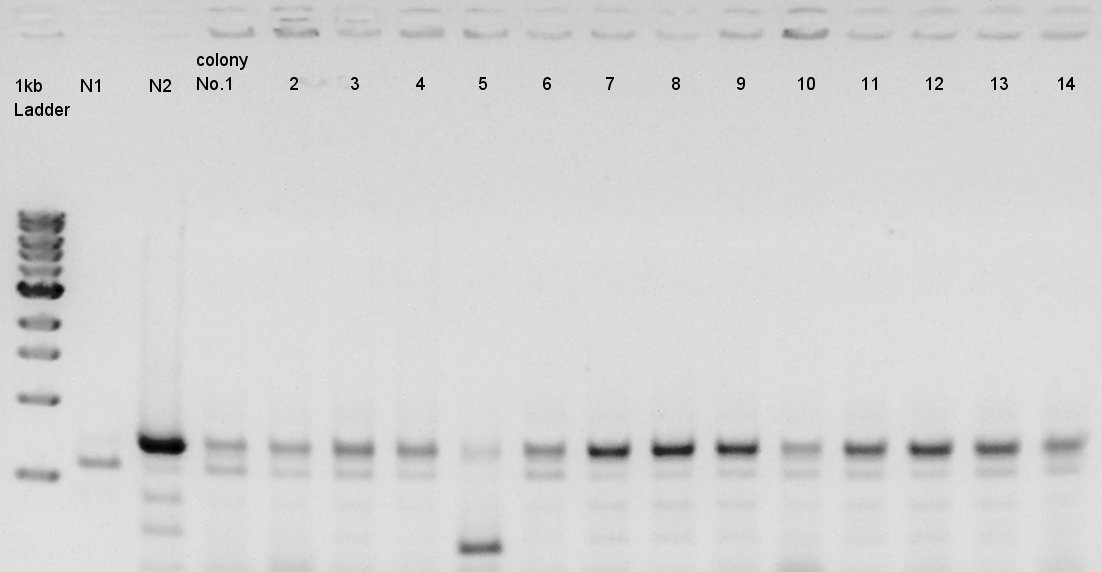
We used N1 (DW only) and N2 (Ag43-dT on pSB1AK3) as controls. Desired product is about 695bp. This length is almost same as N2.
We decided to incubate the No.7 and 8 colony.
Incubation for mini-prep of pT7-RBS-Ag43-dTonpSB1C3 and Ag43-dTonpSB1AK3
Incubation of pT7-RBS-Ag43-dTonpSB1C3 (and Ag43-dT on pSB1AK3) in LBC (LBA) liquid medium.
- Prepared 2 ml LBC (LBA) into culture tubes.
- Re-suspended 2 colonies No.7 and No.8 respectively. Ag43-dT on pSB1AK3 was re-suspended the N2 colony.
- Incubated at 37C for hrs.
Estimation of Concentration of RBS-eYFP-dT (PCR product) and ptetR-pSB1A2
| Solution | Value(ul) |
| 1kb ladder | 1 |
| Insert | 1 |
| Vector | 1 |
From this result, We estimated that the concentration of Insert DNA solution is about 36ng/ul, and Vector DNA is also about 29ng/ul.
Digestion of ptet-pSB1A2 and RBS-Ag43-dT
ptet-pSB1A2(Vector) was digested with EcoRI and SpeI, and RBS-Ag43-dT(Insert) was cut with EcoRI and XbaI. And we used construct of pT7-RBS on pSB1C3 as control for the function of XbaI. Insert (eCFP)
| DNA solution ( 36ng/ul) | 5 ul |
| EcoRI | 1 ul |
| SpeI | 1 ul |
| 10xH buffer | 2 ul |
| DW | 5 ul |
| Total | 20 ul |
Vector(RBS on pSB1A2)
| DNA solution ( 29ng/ul) | 6 ul |
| EcoRI | 1 ul |
| XbaI | 1 ul |
| 10xM buffer | 2 ul |
| DW | 10 ul |
| Total | 20 ul |
control (pT7-RBs on pSB1C3)
| DNA solution (30 ng/ul) | 6 ul |
| XbaI | 1 ul |
| 10xM buffer | 2 ul |
| 100x BSA | 0.2 |
| DW | 11 ul |
| Total | 20 ul |
| Number | Degree | Minute |
| 1 | 37 | 120 |
| 2 | 60 | 15 |
| 3 | 4 | HOLD |
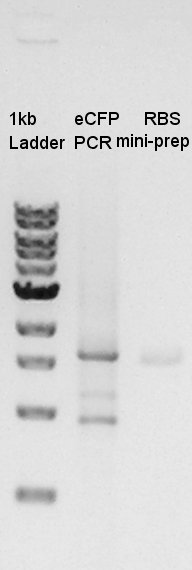
From this result, Vector DNA were digested with XbaI, but remains some amount of non-digested DNA. And about Insert DNA, the DNA solution were not existed in both d- and d+ lines. This means we failed to extract from gel after electrophoresis. We decided to retry PCR.
PCR of eCFP(E0020)
Amplified the part with 100bp-up-EX primer and 200bp-down-PS primer. Desired product is about 800~900bp. Mixed PCR solutions.
| Solution | Volume(ul) |
| DNA | 1 |
| 100bp-up-EX | 1 |
| 200bp-down-PS | 1 |
| MgSO4 | 3 |
| dNTP | 5 |
| 10x KOD-Plus-Neo Buffer | 5 |
| KOD-Plus-Neo | 1 |
| DW | 33 |
| Total | 50 |
| Number | Degree | Second |
| 1 | 94 | 120 |
| 2 | 98 | 10 |
| 3 | 58.0 | 30 |
| 4 | 68 | 60 |
| 5 | 4 | HOLD |
Cycle:2~4 x 35
 "
"