Team:Frankfurt/Notebook
From 2012.igem.org
(Difference between revisions)
(→August 2012) |
(→July 2012) |
||
| Line 10: | Line 10: | ||
==July 2012== | ==July 2012== | ||
| - | + | 1. Plasmid isolation of p426, p423, pUD8e, pUD22e from ''E.coli'' | |
| - | + | 2. Isolation of chromosomal DNA of CEN.PK2-1C | |
| - | + | 3. Trials to get the genes, promoters and terminators via PCR | |
==August 2012== | ==August 2012== | ||
Revision as of 22:05, 26 September 2012

Contents |
Labwork
May and June 2012
- Arrangements for labwork
- preparation of competent cells (E.coli, S.cerevisiae), agarose plates (LB, YEPD, SCD-ura,…), medium for E.coli and S.cerevisiae
- Purchasing of the equipment (reaction tubes, glass bottles, pipette tips,..)
- Primer design
July 2012
1. Plasmid isolation of p426, p423, pUD8e, pUD22e from E.coli 2. Isolation of chromosomal DNA of CEN.PK2-1C 3. Trials to get the genes, promoters and terminators via PCR
August 2012
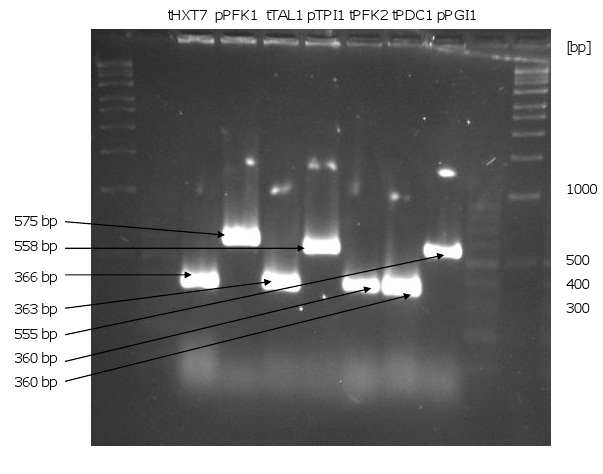
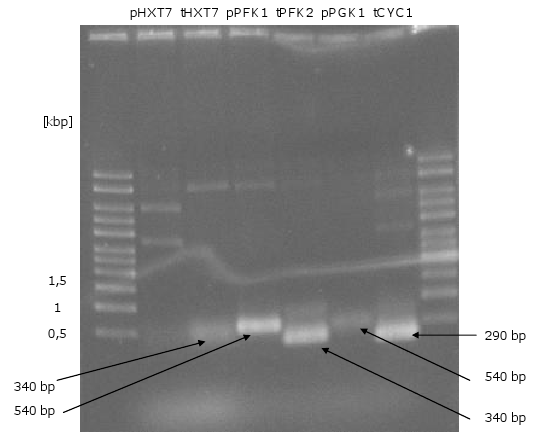
1. PCR of the genes, promoters and terminators
- all genes (without KO and KAH), promoters and terminators could be amplified
2. Linearization of p426 and p423 with SpeI and XhoI
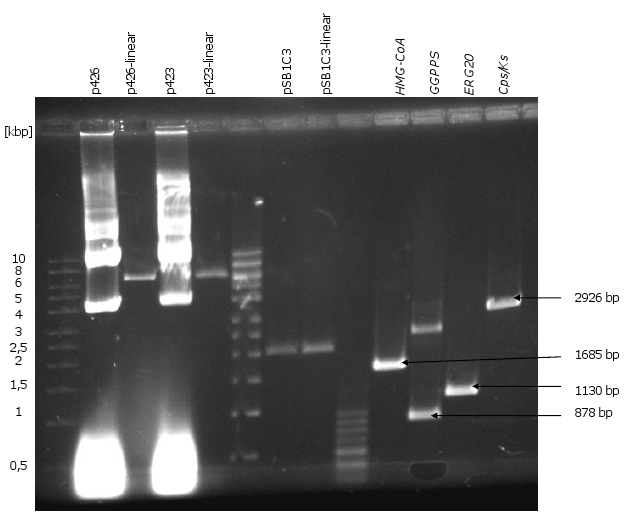
| Templates | Amplified DNA Fragments |
|---|---|
| synthesized sequence of HMG-CoA | HMG-CoA |
| synthesized sequence of GGPPS | GGPPS |
| synthesized sequence of Cps/Ks | CPS/KS |
| chromosomal DNA of CEN.PK2-1C | ERG20 |
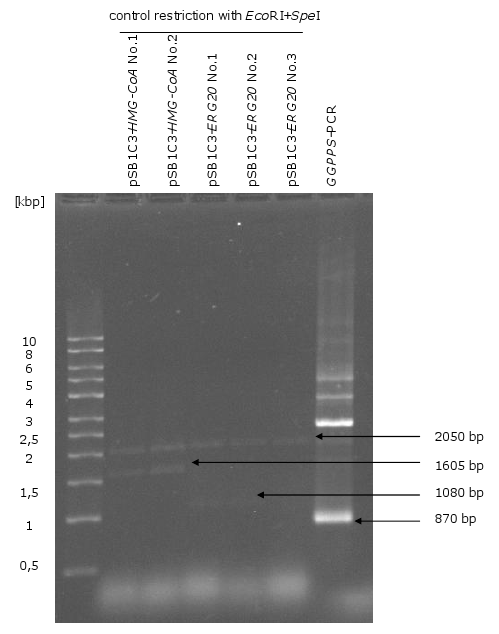
3. Biobrick production of the genes HMG-CoA, ERG20, CPS/KS
- restriction of 3 µg of the genes with EcoRI and PstI
- ligation of biobrick genes with linear pSB1C3
- transformation of the ligation in E.coli
- plasmid isolation of E.coli clones
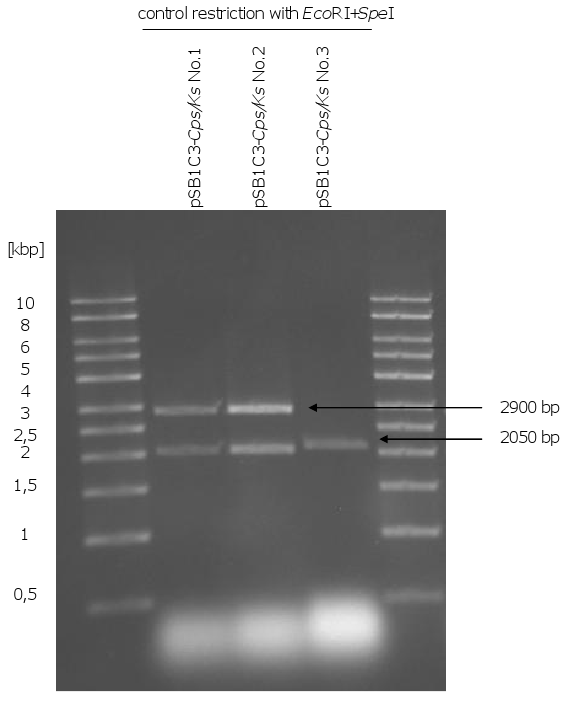
- control restriction of biobrick plasmids with EcoRI and SpeI
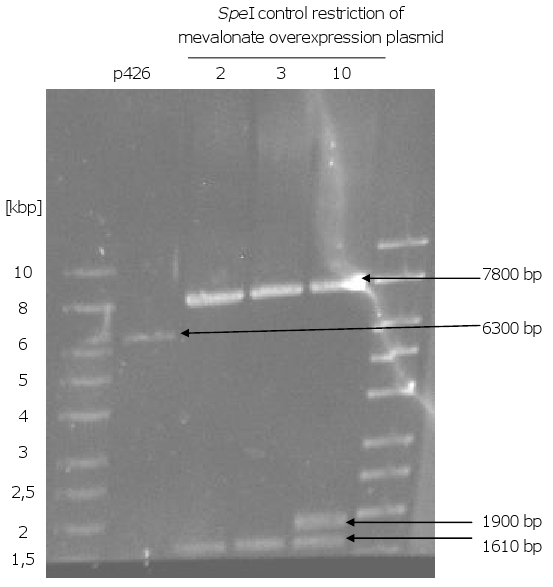
4. Formation of the mevalonate overexpression plasmid via gap repair
- first and second yeast transformation with equimolar quantities of DNA fragments for mevalonate overexpression (p426 with 7 inserts): only very small colonies could grow after the first and second transformation
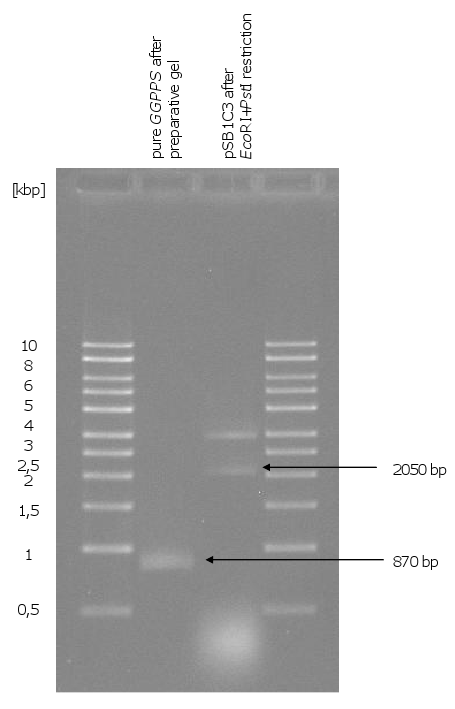
- using pure GGPPS (purification of a preparative gel) for the third yeast transformation: normal size of the colonies
- inoculation of several clones of the third yeast transformation
- plasmid preparation of the clones
- transformation of the plasmids in E.coli
5. Amplifying pSB1C3 for biobrick production
- trials to amplify pSB1C3, whose blunt ends were ligated and transformed in E.coli
- pSB1C3 should be linearized by EcoRI and PstI : did not work (two fragments instead of one)
- preparative gel of the linear fragment: very low concentration of linear pSB1C3 (was not sufficient for ligation)
6. GC analysis
- GC analysis of the wild type CEN.PK2-1C (standard GGOH): as expected no GGOH could be observed
7. Assembly of the KO and the KAH fragments
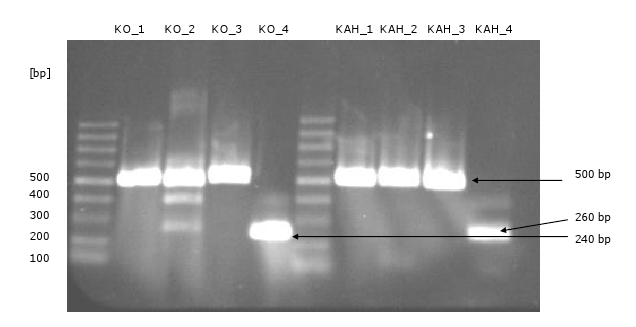
- amplification of the fragments via PCR (there are four fragments of each gene with an overhang to the fragment beside of 30 bp)
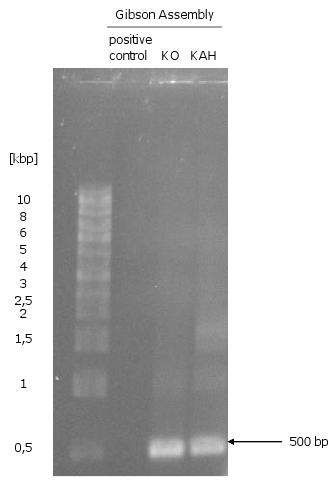
- Gibson assembly of the fragments of KO and KAH: did not work
September 2012
- Formation of the mevalonate overexpression plasmid via gap repair
- plasmid isolation (p426 with 7 inserts) from E.coli
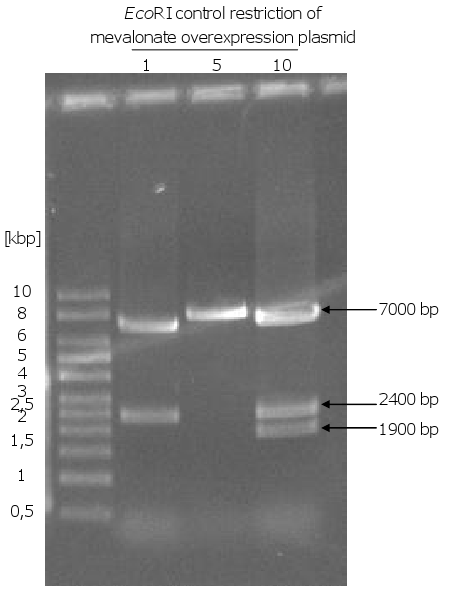
- control restriction of the plasmids with EcoRI and SpeI: one clone out of 10 got the right sizes
- Ergosterol experiment
- idea: maybe the clones of the first and the second yeast transformation grow better after ergosterol addition (0,02 g/l)): wildtype with and without ergosterol and one of the clones with and without ergosterol (no significant difference in growth could be observed)
- GGPPS-PCR
- PCR of GGPPS with shorter synthesis time in order to get only the correct fragment
- Amplification of pSB1C3 for biobrick production
- transformation of pSB1C3-RFP in E.coli
- isolation of the plasmid from E.coli
- linearization with EcoRI and PstI: it worked (only the correct fragment was observed)
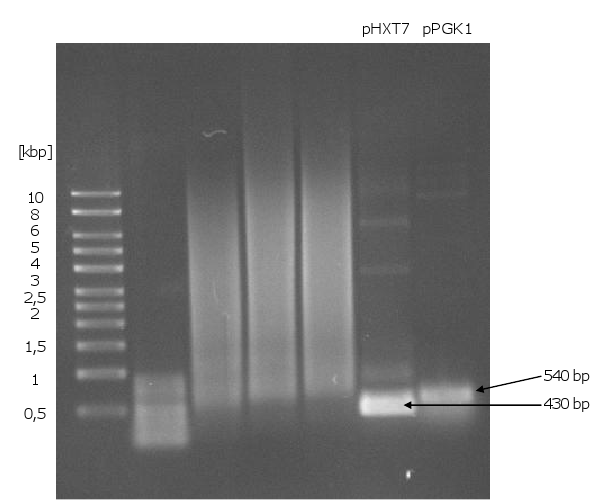
- PCR of biobrick-promoters and -terminators
- PCR of biobrick promoters and terminators that were used to build the mevalonate overexpression plasmid
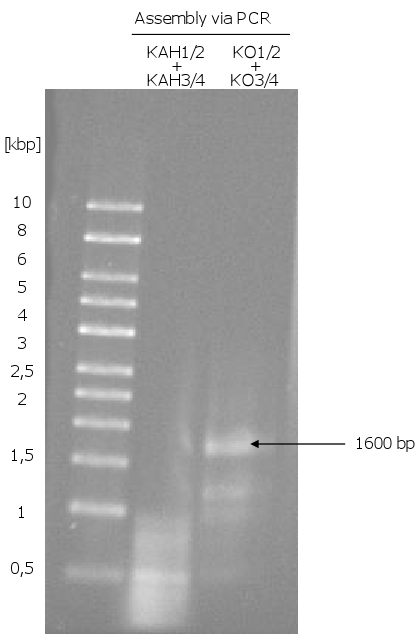
- Assembly of the KO and the KAH fragments
- trials to assembly the KO and the KAH fragments via PCR (the overhangs were used as primer): KO worked, KAH not
- amplification of KO
- Biobrick production of the genes GGPPS, KO and the promoters and terminators
- restriction of 3 µg of the DNA fragments with EcoRI and PstI
- ligation of the promoters, terminators, KO and the GGPPS with pSB1C3
- transformation of the ligation in E.coli
- plasmid isolation of E.coli clones
- control restriction of biobrick plasmids with EcoRI and SpeI
- Midi-preparation of plasmids for sequencing
- biobrick plasmids
- mevalonate overexpression plasmid
- GC
- GC analysis of the wild type CEN.PK2-1C with p426 and the wild type with the mevalonate overexpression plasmid in SCD-ura
- standard GGOH
- Formation of the plasmid for steviol synthesis
- yeast transformation with equimolar quantities of DNA fragments for steviol synthesis (p423 with 7 inserts)
- problem: could not assemble KAH, assembled KAH1/2 and KAH3/4 were used (KAH1/2 and KAH3/4 only got a 30 bp overhang instead of a 45 bp overhang): gap repair did not work
 "
"