Team:Frankfurt/Notebook
From 2012.igem.org
(→September 2012) |
(→September 2012) |
||
| (11 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | { | + | {{Template:Template_Frankfurt2012}} |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
=Labwork= | =Labwork= | ||
==May and June 2012== | ==May and June 2012== | ||
| - | + | 1. Arrangements for labwork <br> | |
| - | + | * preparation of competent cells (''E.coli'', ''S.cerevisiae''), agarose plates (LB, YEPD, SCD-ura,…), medium for ''E.coli'' and ''S.cerevisiae'' | |
| - | + | 2. Purchasing of the equipment (reaction tubes, glass bottles, pipette tips,..) | |
| - | + | 3. Primer design | |
==July 2012== | ==July 2012== | ||
| - | + | 1. Plasmid isolation of p426, p423, pUD8e, pUD22e from ''E.coli''<br> | |
| - | + | 2. Isolation of chromosomal DNA of CEN.PK2-1C<br> | |
| - | + | 3. Trials to get the genes, promoters and terminators via PCR<br> | |
==August 2012== | ==August 2012== | ||
| - | + | 1. PCR of the genes, promoters and terminators<br> | |
| - | + | * all genes (without ''KO'' and ''KAH''), promoters and terminators could be amplified | |
| - | + | 2. Linearization of p426 and p423 with ''SpeI'' and ''XhoI'' | |
{|class="wikitable zebra" | {|class="wikitable zebra" | ||
|- | |- | ||
| Line 44: | Line 31: | ||
|} | |} | ||
{|width="100%" align="center" | {|width="100%" align="center" | ||
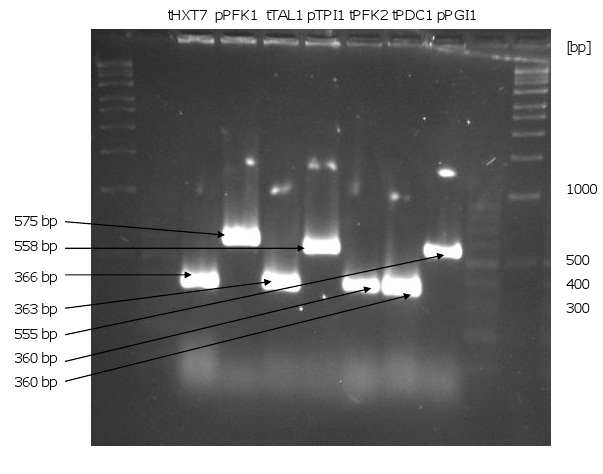
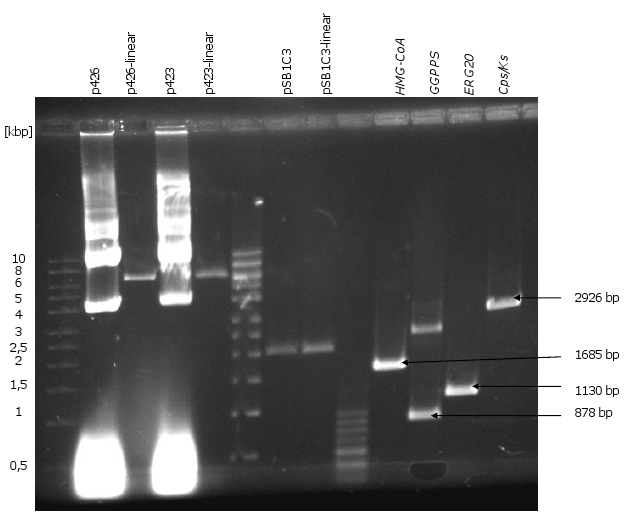
| - | [[Image:Gel1_1.png|450px|thumb|left|'''Gel 1: PCR of the promoters and terminators.'''<br>There are shown the DNA fragments of the PCR with appropriate annealing temperatures. All promoters and terminators could be amplified. As templates were used the plasmids pUD8e(''tHXT7'', ''pPFK1'', ''tTAL1'', ''pTPI1'') and pUD22e (''tPFK2'', ''tPDC1'', ''pPGI1'').]][[Image:Gel2_1.png|415px|thumb| | + | [[Image:Gel1_1.png|450px|thumb|left|'''Gel 1: PCR of the promoters and terminators.'''<br>There are shown the DNA fragments of the PCR with appropriate annealing temperatures. All promoters and terminators could be amplified. As templates were used the plasmids pUD8e(''tHXT7'', ''pPFK1'', ''tTAL1'', ''pTPI1'') and pUD22e (''tPFK2'', ''tPDC1'', ''pPGI1'').]][[Image:Gel2_1.png|415px|thumb|right|'''Gel 2: Linearization of p426, p423, pSB1C3 and PCR of the genes ''HMG-CoA'', ''GGPPS'', ''ERG20'', ''CPS/KS''.'''<br> As a control it is also the circular p426 and p423 shown. The linear p426 and p423 are about 6300 bp long. pSB1C3-linear means the restriction of the plasmid with ''EcoRI'' and ''PstI'' in this case (it is 2050 bp long). As PCR templates for ''HMG-CoA'', ''GGPP''S and ''CPS/KS'' were used the synthesized DNA fragments and for ''ERG20'' chromosomal DNA of CEN.PK2-1C.]] |
|} | |} | ||
| - | + | 3. Biobrick production of the genes ''HMG-CoA'', ''ERG20'', ''CPS/KS''<br> | |
| - | + | * restriction of 3 µg of the genes with ''EcoRI'' and ''PstI''<br> | |
| - | + | * ligation of biobrick genes with linear pSB1C3<br> | |
| - | + | * transformation of the ligation in ''E.coli''<br> | |
| - | + | * plasmid isolation of ''E.coli'' clones<br> | |
| - | + | * control restriction of biobrick plasmids with ''EcoRI'' and ''SpeI''<br> | |
{|width="100%" align="center" | {|width="100%" align="center" | ||
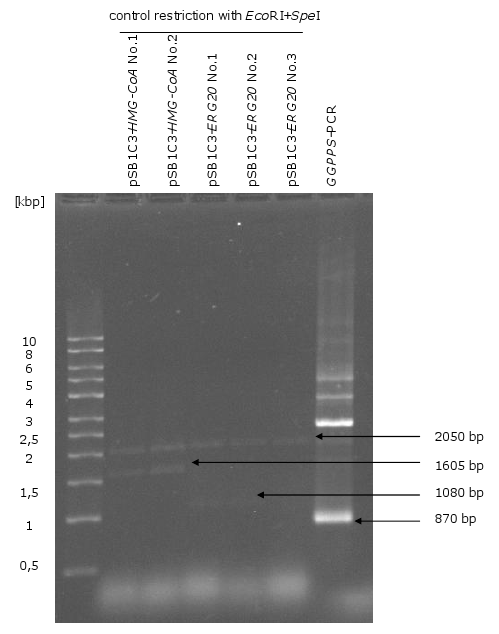
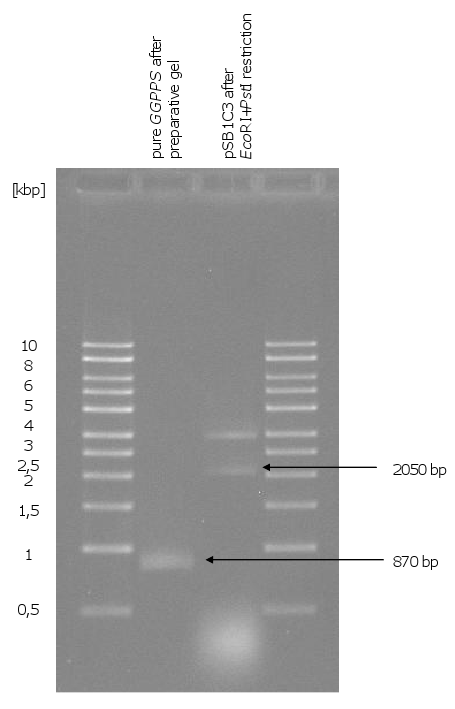
[[Image:Gel3_1.png|370px|thumb|left|'''Gel 3: Control restriction of pSB1C3-''HMG-CoA'', pSB1C3-''ERG20'' and PCR of ''GGPPS''.'''<br> | [[Image:Gel3_1.png|370px|thumb|left|'''Gel 3: Control restriction of pSB1C3-''HMG-CoA'', pSB1C3-''ERG20'' and PCR of ''GGPPS''.'''<br> | ||
| Line 59: | Line 46: | ||
on this picture. With the PCR of ''GGPPS'' could not only amplified the correct fragment (870 | on this picture. With the PCR of ''GGPPS'' could not only amplified the correct fragment (870 | ||
bp) but also a fragment of 3000 bp.]] | bp) but also a fragment of 3000 bp.]] | ||
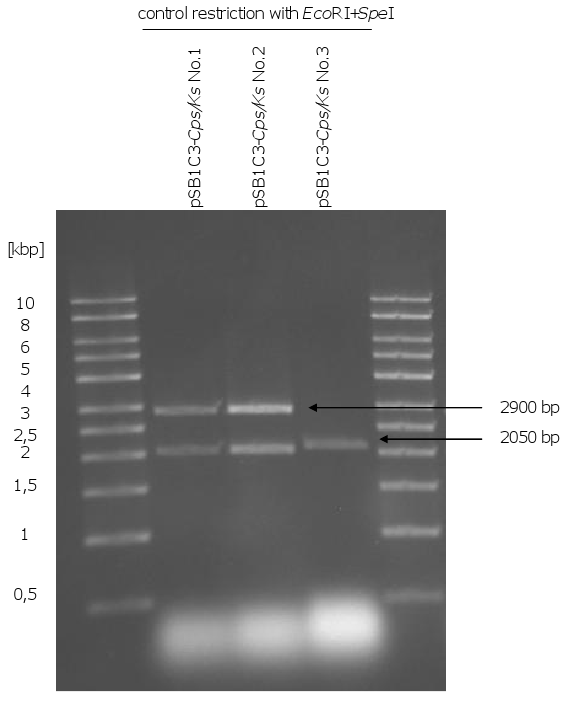
| - | [[Image:Gel5_1.png|370px|thumb| | + | [[Image:Gel5_1.png|370px|thumb|right|'''Gel 5: Control restriction of pSB1C3-''CPS/KS''.'''<br> |
The biobrick plasmids were cut with ''EcoRI'' and ''SpeI''. The correct sizes of pSB1C3-''CPS/ | The biobrick plasmids were cut with ''EcoRI'' and ''SpeI''. The correct sizes of pSB1C3-''CPS/ | ||
KS'' from two clones could be observed (2050 bp + 2900 bp). The plasmid of clone 3 is the | KS'' from two clones could be observed (2050 bp + 2900 bp). The plasmid of clone 3 is the | ||
pSB1C3 without the insert.]] | pSB1C3 without the insert.]] | ||
|} | |} | ||
| - | + | 4. Formation of the mevalonate overexpression plasmid via gap repair | |
| - | + | * first and second yeast transformation with equimolar quantities of DNA fragments for mevalonate overexpression (p426 with 7 inserts): only very small colonies could grow after the first and second transformation | |
| - | + | * using pure ''GGPPS'' (purification of a preparative gel) for the third yeast transformation: normal size of the colonies | |
| - | + | * inoculation of several clones of the third yeast transformation | |
| - | + | * plasmid preparation of the clones | |
| - | + | * transformation of the plasmids in ''E.coli''<br> | |
| - | + | 5. Amplifying pSB1C3 for biobrick production | |
| - | + | * trials to amplify pSB1C3, whose blunt ends were ligated and transformed in ''E.coli'' | |
| - | + | * pSB1C3 should be linearized by ''EcoRI'' and ''PstI'' : did not work (two fragments instead of one)<br> | |
{|width="100%" align="center" | {|width="100%" align="center" | ||
[[Image:Gel4_1.png|300px|thumb|left|'''Gel 4: Purified ''GGPPS'' and linearization of pSB1C3.'''<br>The 870 bp fragment of the ''GGPPS'' PCR (see gel 3) were cut out from a preparative gel and then purified with a gel extraction kit (The purified ''GGPPS'' could be used for transformation, but not for constructing a biobrick. Therefore another PCR was made with a very short synthesis time. So only the 870 bp fragment was obtained.)<br>Because of lack of pSB1C3 the remained linear pSB1C3 was ligated and transformed in ''E.coli''. After isolation of the plasmid it was restricted with ''EcoRI'' and ''PstI''. But there was not | [[Image:Gel4_1.png|300px|thumb|left|'''Gel 4: Purified ''GGPPS'' and linearization of pSB1C3.'''<br>The 870 bp fragment of the ''GGPPS'' PCR (see gel 3) were cut out from a preparative gel and then purified with a gel extraction kit (The purified ''GGPPS'' could be used for transformation, but not for constructing a biobrick. Therefore another PCR was made with a very short synthesis time. So only the 870 bp fragment was obtained.)<br>Because of lack of pSB1C3 the remained linear pSB1C3 was ligated and transformed in ''E.coli''. After isolation of the plasmid it was restricted with ''EcoRI'' and ''PstI''. But there was not | ||
only the linear fragment of 2050 bp but also another fragment about 4000 bp.]] | only the linear fragment of 2050 bp but also another fragment about 4000 bp.]] | ||
|} | |} | ||
| - | + | * preparative gel of the linear fragment: very low concentration of linear pSB1C3 (was not sufficient for ligation) | |
| - | + | 6. GC analysis | |
| - | + | * GC analysis of the wild type CEN.PK2-1C (standard GGOH): as expected no GGOH could be observed | |
| - | + | 7. Assembly of the ''KO'' and the ''KAH'' fragments | |
| - | + | * amplification of the fragments via PCR (there are four fragments of each gene with an overhang to the fragment beside of 30 bp) | |
| - | + | * Gibson assembly of the fragments of ''KO'' and ''KAH'': did not work<br> | |
{|width="100%" align="center" | {|width="100%" align="center" | ||
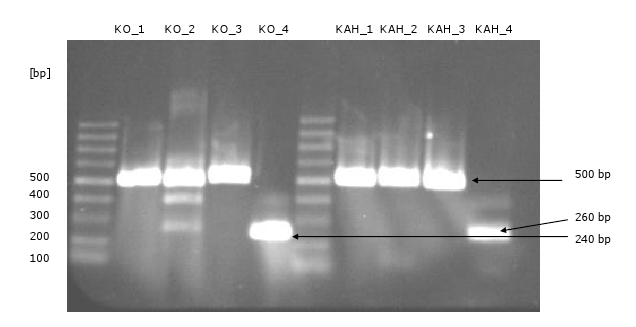
[[Image:Gel6_1.png|500px|thumb|left|'''Gel 6: PCR of ''KO'' and ''KAH'' fragments'''<br> | [[Image:Gel6_1.png|500px|thumb|left|'''Gel 6: PCR of ''KO'' and ''KAH'' fragments'''<br> | ||
Each gene consists of four fragments, which have a 30 bp overhang to the fragments beside. | Each gene consists of four fragments, which have a 30 bp overhang to the fragments beside. | ||
In order to do the Gibson Assembly, the fragments were amplified via PCR.]] | In order to do the Gibson Assembly, the fragments were amplified via PCR.]] | ||
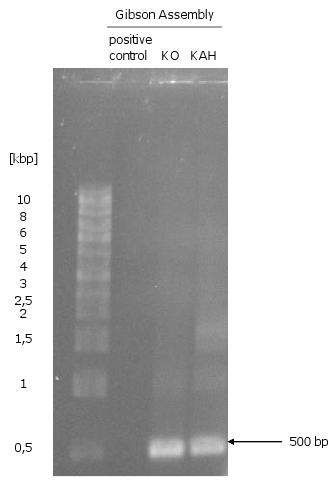
| - | [[Image:Gel7_1.png|370px|thumb| | + | [[Image:Gel7_1.png|370px|thumb|right|'''Gel 7: Gibson Assembly of ''KO'' and ''KAH'''''<br> |
The assembly did not work, because there are only the 500 bp of the single fragments.]] | The assembly did not work, because there are only the 500 bp of the single fragments.]] | ||
|} | |} | ||
==September 2012== | ==September 2012== | ||
| - | + | 1. Formation of the mevalonate overexpression plasmid via gap repair | |
| - | + | * plasmid isolation (p426 with 7 inserts) from ''E.coli'' | |
| - | + | * control restriction of the plasmids with ''EcoRI'' and ''SpeI'': one clone out of 10 got the right sizes | |
| - | + | ||
{|width="100%" align="center" | {|width="100%" align="center" | ||
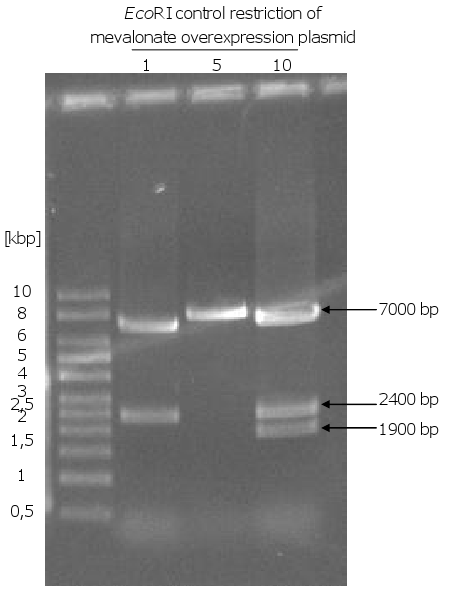
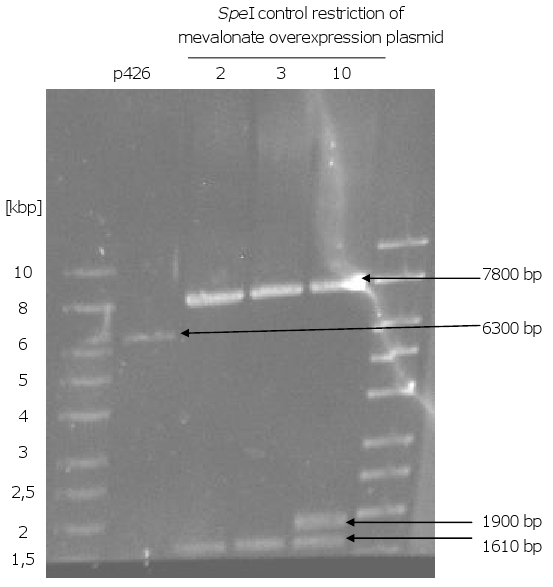
[[Image:Gel11_1.png|270px|thumb|left|'''Gel 11: Possible mevalonate overexpression plasmids from ''E.coli''.'''<br>After yeast transformation with p426 and 7 inserts, isolation of the plasmids, transformation | [[Image:Gel11_1.png|270px|thumb|left|'''Gel 11: Possible mevalonate overexpression plasmids from ''E.coli''.'''<br>After yeast transformation with p426 and 7 inserts, isolation of the plasmids, transformation | ||
| Line 105: | Line 91: | ||
be the correct one. To have a positive control p426 was also cut with ''SpeI'' (6300 bp).]] | be the correct one. To have a positive control p426 was also cut with ''SpeI'' (6300 bp).]] | ||
|} | |} | ||
| - | + | 2. Ergosterol experiment | |
| - | + | * idea: maybe the clones of the first and the second yeast transformation grow better after ergosterol addition (0,02 g/l)): wildtype with and without ergosterol and one of the clones with and without ergosterol (no significant difference in growth could be observed) | |
| - | + | 3. ''GGPPS''-PCR | |
| - | + | * PCR of ''GGPPS'' with shorter synthesis time in order to get only the correct fragment | |
| - | + | 4. Amplification of pSB1C3 for biobrick production | |
| - | + | * transformation of pSB1C3-RFP in ''E.coli'' | |
| - | + | * isolation of the plasmid from ''E.coli'' | |
| - | + | * linearization with ''EcoRI'' and ''PstI'': it worked (only the correct fragment was observed) | |
| - | + | 5. PCR of biobrick-promoters and -terminators | |
| - | + | * PCR of biobrick promoters and terminators that were used to build the mevalonate overexpression plasmid | |
{|width="100%" align="center" | {|width="100%" align="center" | ||
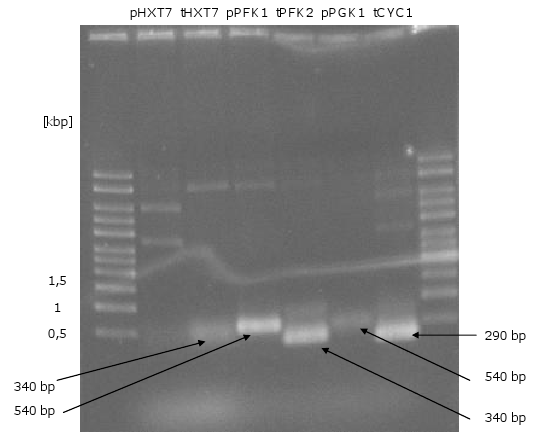
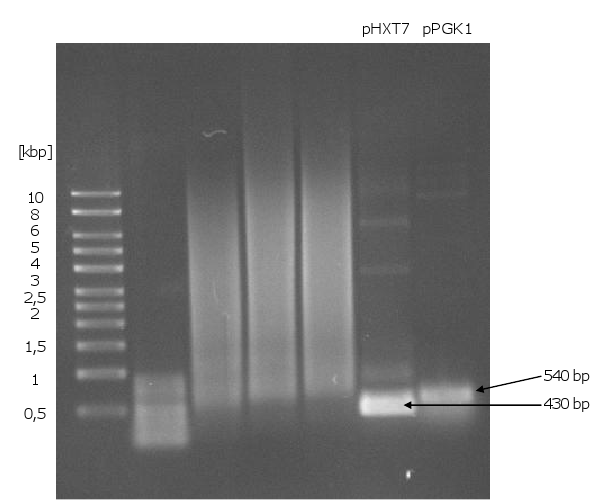
[[Image:Gel10_1.png|400px|thumb|left|'''Gel 10: PCR for biobrick-promoters and –terminators.'''<br> | [[Image:Gel10_1.png|400px|thumb|left|'''Gel 10: PCR for biobrick-promoters and –terminators.'''<br> | ||
| Line 124: | Line 110: | ||
templates were used the plasmids pUD22e (''pPGK1'') and p426 (''pHXT7'').]] | templates were used the plasmids pUD22e (''pPGK1'') and p426 (''pHXT7'').]] | ||
|} | |} | ||
| - | + | 6. Assembly of the ''KO'' and the ''KAH'' fragments | |
| - | + | * trials to assembly the ''KO'' and the ''KAH'' fragments via PCR (the overhangs were used as primer): ''KO'' worked, ''KAH'' not | |
{|width="100%" align="center" | {|width="100%" align="center" | ||
[[Image:Gel8_1.png|400px|thumb|left|'''Gel 8: Assembly of ''KO1/2'', ''KO3/4'', ''KAH1/2'' and ''KAH3/4'' via PCR.'''<br> | [[Image:Gel8_1.png|400px|thumb|left|'''Gel 8: Assembly of ''KO1/2'', ''KO3/4'', ''KAH1/2'' and ''KAH3/4'' via PCR.'''<br> | ||
| Line 132: | Line 118: | ||
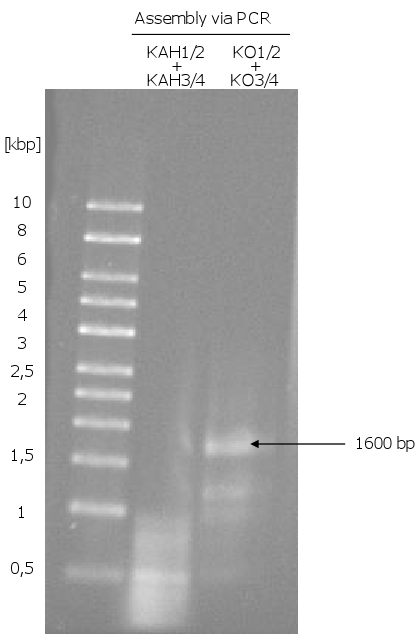
[[Image:Gel9_1.png|350px|thumb|left|'''Gel 9: Assembly of ''KO'' and ''KAH'' via PCR.'''<br>The assembly of ''KAH'' did not work. For the PCR of ''KO'' were used the assembled ''KO1/2'' and ''KO3/4'' with an appropriate annealing temperature (the 30 bp overhang was used as primer). The ''KO'' fragment (1600 bp) was then amplified via a PCR with normal primers.]] | [[Image:Gel9_1.png|350px|thumb|left|'''Gel 9: Assembly of ''KO'' and ''KAH'' via PCR.'''<br>The assembly of ''KAH'' did not work. For the PCR of ''KO'' were used the assembled ''KO1/2'' and ''KO3/4'' with an appropriate annealing temperature (the 30 bp overhang was used as primer). The ''KO'' fragment (1600 bp) was then amplified via a PCR with normal primers.]] | ||
|} | |} | ||
| - | + | * amplification of ''KO'' | |
| - | + | 7. Biobrick production of the genes ''GGPPS'', ''KO'' and the promoters and terminators | |
| - | + | * restriction of 3 µg of the DNA fragments with ''EcoRI'' and ''PstI'' | |
| - | + | * ligation of the promoters, terminators, ''KO'' and the ''GGPPS'' with pSB1C3 | |
| - | + | * transformation of the ligation in ''E.coli'' | |
| - | + | * plasmid isolation of ''E.coli'' clones | |
| - | + | * control restriction of biobrick plasmids with ''EcoRI'' and ''SpeI'' | |
| - | + | 8. Midi-preparation of plasmids for sequencing | |
| - | + | * biobrick plasmids | |
| - | + | * mevalonate overexpression plasmid | |
| - | + | 9. GC | |
| - | + | * GC analysis of the wild type CEN.PK2-1C with p426 and the wild type with the mevalonate overexpression plasmid in SCD-ura | |
| - | + | * standard GGOH | |
| - | + | 10. Formation of the plasmid for steviol synthesis | |
| - | + | * yeast transformation with equimolar quantities of DNA fragments for steviol synthesis (p423 with 7 inserts) | |
| - | + | * problem: could not assemble KAH, assembled ''KAH1/2'' and ''KAH3/4'' were used (''KAH1/2'' and ''KAH3/4'' only got a 30 bp overhang instead of a 45 bp overhang): gap repair did not work | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Latest revision as of 22:13, 26 September 2012

Contents |
Labwork
May and June 2012
1. Arrangements for labwork
- preparation of competent cells (E.coli, S.cerevisiae), agarose plates (LB, YEPD, SCD-ura,…), medium for E.coli and S.cerevisiae
2. Purchasing of the equipment (reaction tubes, glass bottles, pipette tips,..) 3. Primer design
July 2012
1. Plasmid isolation of p426, p423, pUD8e, pUD22e from E.coli
2. Isolation of chromosomal DNA of CEN.PK2-1C
3. Trials to get the genes, promoters and terminators via PCR
August 2012
1. PCR of the genes, promoters and terminators
- all genes (without KO and KAH), promoters and terminators could be amplified
2. Linearization of p426 and p423 with SpeI and XhoI
| Templates | Amplified DNA Fragments |
|---|---|
| synthesized sequence of HMG-CoA | HMG-CoA |
| synthesized sequence of GGPPS | GGPPS |
| synthesized sequence of Cps/Ks | CPS/KS |
| chromosomal DNA of CEN.PK2-1C | ERG20 |
3. Biobrick production of the genes HMG-CoA, ERG20, CPS/KS
- restriction of 3 µg of the genes with EcoRI and PstI
- ligation of biobrick genes with linear pSB1C3
- transformation of the ligation in E.coli
- plasmid isolation of E.coli clones
- control restriction of biobrick plasmids with EcoRI and SpeI
4. Formation of the mevalonate overexpression plasmid via gap repair
- first and second yeast transformation with equimolar quantities of DNA fragments for mevalonate overexpression (p426 with 7 inserts): only very small colonies could grow after the first and second transformation
- using pure GGPPS (purification of a preparative gel) for the third yeast transformation: normal size of the colonies
- inoculation of several clones of the third yeast transformation
- plasmid preparation of the clones
- transformation of the plasmids in E.coli
5. Amplifying pSB1C3 for biobrick production
- trials to amplify pSB1C3, whose blunt ends were ligated and transformed in E.coli
- pSB1C3 should be linearized by EcoRI and PstI : did not work (two fragments instead of one)
- preparative gel of the linear fragment: very low concentration of linear pSB1C3 (was not sufficient for ligation)
6. GC analysis
- GC analysis of the wild type CEN.PK2-1C (standard GGOH): as expected no GGOH could be observed
7. Assembly of the KO and the KAH fragments
- amplification of the fragments via PCR (there are four fragments of each gene with an overhang to the fragment beside of 30 bp)
- Gibson assembly of the fragments of KO and KAH: did not work
September 2012
1. Formation of the mevalonate overexpression plasmid via gap repair
- plasmid isolation (p426 with 7 inserts) from E.coli
- control restriction of the plasmids with EcoRI and SpeI: one clone out of 10 got the right sizes
2. Ergosterol experiment
- idea: maybe the clones of the first and the second yeast transformation grow better after ergosterol addition (0,02 g/l)): wildtype with and without ergosterol and one of the clones with and without ergosterol (no significant difference in growth could be observed)
3. GGPPS-PCR
- PCR of GGPPS with shorter synthesis time in order to get only the correct fragment
4. Amplification of pSB1C3 for biobrick production
- transformation of pSB1C3-RFP in E.coli
- isolation of the plasmid from E.coli
- linearization with EcoRI and PstI: it worked (only the correct fragment was observed)
5. PCR of biobrick-promoters and -terminators
- PCR of biobrick promoters and terminators that were used to build the mevalonate overexpression plasmid
6. Assembly of the KO and the KAH fragments
- trials to assembly the KO and the KAH fragments via PCR (the overhangs were used as primer): KO worked, KAH not
- amplification of KO
7. Biobrick production of the genes GGPPS, KO and the promoters and terminators
- restriction of 3 µg of the DNA fragments with EcoRI and PstI
- ligation of the promoters, terminators, KO and the GGPPS with pSB1C3
- transformation of the ligation in E.coli
- plasmid isolation of E.coli clones
- control restriction of biobrick plasmids with EcoRI and SpeI
8. Midi-preparation of plasmids for sequencing
- biobrick plasmids
- mevalonate overexpression plasmid
9. GC
- GC analysis of the wild type CEN.PK2-1C with p426 and the wild type with the mevalonate overexpression plasmid in SCD-ura
- standard GGOH
10. Formation of the plasmid for steviol synthesis
- yeast transformation with equimolar quantities of DNA fragments for steviol synthesis (p423 with 7 inserts)
- problem: could not assemble KAH, assembled KAH1/2 and KAH3/4 were used (KAH1/2 and KAH3/4 only got a 30 bp overhang instead of a 45 bp overhang): gap repair did not work
 "
"