Team:Cambridge/Lab book/Week 8
From 2012.igem.org
| Week: | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 |
|---|
Contents |
Monday (13/08/12)
Tuesday (14/08/12)
Gibson assembly of positive control
- Fragments used:
- Protocol changed slightly: 0.5 μl of each DNA fragment solution mixed in with master mix to make up 4 μl total volume (1μl DNA and 3μl master mix).
Transformation of e.coli with positive control DNA
- Chemically competent e.coli cells transformed with plasmid DNA produced by Gibson assembly step.
- Transformants plated out on 50 μg/ml kanomycin plates. Incubated overnight at 37 °C.
Wednesday (15/08/12)
PCR of positive control fragments
- Standard PCR settings used
Extraction of positive control DNA
- Positive control DNA from gel excised and purified.
- Additional elution steps used to concentrate resultant solution.
- Verified with nanodropper - final DNA concentrations:
Gibson assembly of positive control DNA
Transformation of positive control DNA into e.coli
- Chemically competent e.coli transformed with positive control Gibson products made previously.
- Transformants plated out onto 50 μg/ml kanomycin plates and incubated at 37 °C overnight.
Characterization of fluoride riboswitch construct
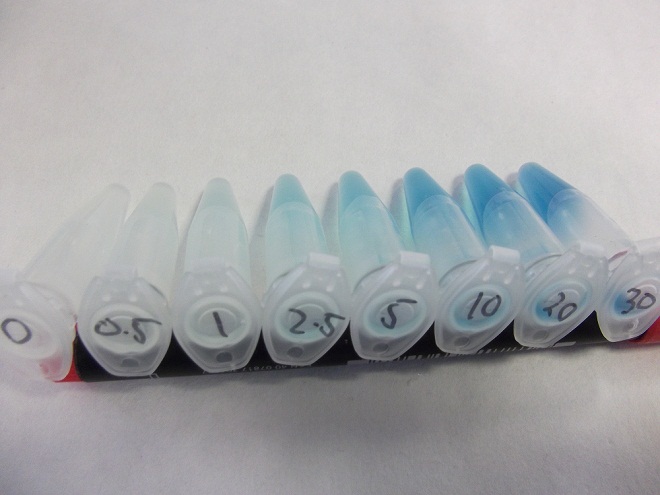
- Strain of bacillus lacking fluoride transporter system tested. Concentrations used:
- 0mM - 0.5mM - 1mM - 2.5mM - 5mM - 10mM - 20mM - 30mM
Thursday (16/08/12)
Characterization of fluoride riboswitch construct
- Eppindorfs containing the bacillus with the fluoride riboswitch removed from incubator and imaged.
- Now this definitely works, we will try quantifying this riboswitch with an ONPG assay and the plate reader.
PCR ran by paul - insert photos/discuss
- Only positive control and two sets of triplicates worked
- YFP and CFP extracted successfully
Friday (17/08/12)
- Team meet up today! No lab work was done.
Saturday (18/08/12)
 "
"