Team:Calgary/Project/OSCAR/Desulfurization
From 2012.igem.org


Hello! iGEM Calgary's wiki functions best with Javascript enabled, especially for mobile devices. We recommend that you enable Javascript on your device for the best wiki-viewing experience. Thanks!
Desulfurization

Why Remove Sulfur?
Sulfur is the third most abundant element in crude oil (Ma, 2010). When sulfur containing hydrocarbons in fuel are burned, S02 and S03 are released into the atmosphere, which are major components in air pollution, acid rain, and dry acid deposition (Reichmuth et al., 2000). Acid rain (rain with a pH less than 5.3) can cause acidification of aquatic and terrestrial environments, causing nutrient leeching and damaging the health and habitats of many species not tolerant to acidification. Sulfur dioxide can also aggravate respiratory health issues. In addition, sulfur in fuels can reduce the effectiveness of vehicular emmision control measures such as catalytic converters, as well as reduce the life and cause damage to valuable catalysts in the oil refining process.
Stringent regulations on sulfur content in fossil fuels have been put into place. As of 2005, low-sulfur gasoline (gasoline with a sulfur content of less than 30 mg/kg) is required across all of Canada (Source: Environment Canada). Naphthenic acid compounds found in the tailings ponds can also be sulfur-containing heterocycles. Because our aim is to convert these compounds to useable fuel, these sulfur atoms must therefore be removed from the naphthenic acids before this goal can be realized. In addition, removal of the sulfur atom in a heterocycle alters the connectivity of the carbon skeleton of the molecule, changing it from a 3-ring compound to a 2-ring compound.
Our Vision
The current process of hydrodesulfurization used to remove sulfur during fuel upgrading is environmentally unsound (requiring high temperature and pressure) as well as being quite costly. This process also releases the sulfur in the form of hydrogen sulfide gas, a toxic compound which is then converted to elemental sulfur or sulfuric acid. During this process temperatures range around 350°C with pressures as high as 130 atm. In contrast to this, microbes have been found capable of harvesting the sulfur out of hydrocarbons, a process which on an industrial scale would be much more environmentally sound and most likely be less costly to carry out. Though a few pathways for biodesulfurization exist in the microbial world, most involve the destruction of part of the carbon skeleton (an example would be the Kodama pathway). This is unwanted in our system, because by removing some of the carbon backbone to be turned into biomass, the cells effectively reduce the fuel quality.
The pathway we have chosen for our system is not destructive in this fashion, and therefore preserves (and improves) fuel quality. The 4S pathway found in Rhodococcus spp. has been characterized and shown to remove sulfur from the model substrate dibenzothiophene (DBT) and convert it to 2-hydroxybiphenyl in a non-destructive manner. DBT and its derivatives make up 60% of the organic sulfur compounds found in crude oil, and are also some of the most difficult to remove. The ability of the 4S pathway to act on derivatives of DBT leads us to hope that it will in addition be capable of acting upon naphthenic acids containing sulfur atoms in their structure.
4S pathway
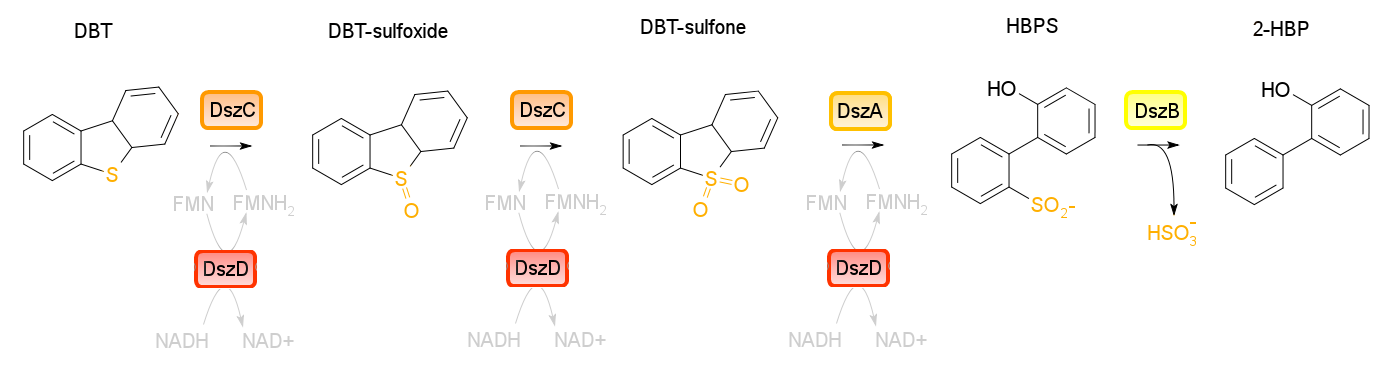
Four enzymes are involved in the 4S pathway, 3 of which are directly involved in the conversion of DBT to 2-HBP. Dibenzothiophene monooxygenase (DszC) is responsible for the first two steps of the pathway, converting DBT to DBT-sulfoxide and finally to DBT-sulfone (DBTO2) through the addition of oxygen to the sulfur atom. DBT-sulfone monooxygenase (DszA) then carries out the next step in the pathway, producing 2-hydroxybiphenyl-2-sulfinic acid (HBPS) through addition of a final oxygen to the heteroatom. This causes cleavage of the chemical bonds at the heteroatom, breaking the ring and converting the compound from a 3-ring structure to a 2-ring structure. HBPS is then converted to the final product of the 4S pathway by HBPS desulfinase (DszB), producing 2-hydroxybiphenyl. At this point, the sulfur has been released from the hydrocarbon in the form of sulfite.
The first three steps of the 4S pathway carried out by the monooxygenase enzymes require FMNH2, and therefore use up the reductive power of the cell. Without a way to recover this, the pathway essentially chokes, desulfurization rates are very low, and cell health suffers. In order to fix this problem, an oxidoreductase (DszD) uses NADH to recycle the FMNH2, allowing the reaction to proceed.
dszA, dszB and dszC form an operon on the pSOX plasmid of R. erythropolis while dszD (a flavin oxidoreductase) is present on the chromosomal DNA of the organism. In natural circumstances the 4S pathway has been found to be quite slow, preventing this method of biodesulfurization from becoming commercially feasable. This is due to a number of limitations found in the native setup of the system. With synthetic biology approaches however, it is possible to optimize this pathway by making some alterations.
Our Approach
1) Find an organism
Rhodococcus spp. have been shown in previous work to be capable of the desulfurization of DBT. An environmental isolate of Rhodococcus shown to be an active desulfurizer was obtained to be the source of the dsz genes we intended to biobrick. The promoter controlling the transcription of this pathway is normally repressed by sulfur containing compounds such as cysteine, methionine, and sulfate (Li et al., 1996). This first problem is easily solved, since our final construct will have a promoter from the registry instead of the native gene promotor, preventing repression from interfering with the expression of the pathway.
In order to get the dsz genes, the desulfurization operon-containing plasmid had to be isolated from the organism. This was performed using a modified version of the miniprep procedure used on Escherichia coli, with the addition of lysozyme so that the cell wall of the gram-positive organism could be broken down. Primers were then designed to amplify the dszA, dszB, and dszC genes from the plasmid DNA, forward primers designed against each gene were designed. These primers contained the biobrick prefix and suffix at their tail ends in order to allow for the construction of the PCR amplicon into a standard biobrick backbone. After fine-tuning PCR conditions using gradient PCR to find conditions that would allow for the amplification of our gene out of the plasmid, each of the dsz genes was constructed into a pSB1C3 backbone. After these parts had been sequence confirmed, the next step would have been to construct these genes into a complete operon for desulfurization. However, through examining the sequences, it was found that before this could be done, steps of mutagenesis would have to be performed in order to remove illegal biobrick cut sites that would interfere with later construction.
2) Mutagenesis: Biobrick Compatability and Increasing DszB Activity
When sequences were examined, the native dszA was found to have four PstI cut sites. Similarily, dszB has a PstI and a NotI cut site, and dszC has two PstI cut sites. In order to be able to use these parts in construction and have them become part of the standard biobrick system, we had to eliminate these cut sites through site-directed mutagenesis. The protocol for Stratagene site directed mutagenesis was used in order to design our mutagenic primers, and for the basics of the entire process, however we chose to use a different (KAPA Biosystems) polymerase because it was available and because of its high fidelity. Because the procedure requires the entire length of the plasmid to be amplified through PCR, we wanted to minimize the risk of any errors being made during this process that would introduce mutations into our genes. During this procedure, two primers are designed for each mutation. Primers are complementary to the DNA strands except for one base pair where the mutation is added, and are complementary to each other so that the mutation site is in the centre of each. The mutation to the DNA was chosen in a way so that a one base pair point mutation is introduced which eliminates the cut site but the codon it belongs to still codes for the same amino acid (a silent mutation). These primers then amplify outward, and therefore after the PCR the whole plasmid is amplified with a point mutation at the desired site.
While we were designing primers to make the dsz genes biobrick compatible, we came across a report in the literature which found a point mutation that was capable of increasing the activity of DszB (Oshiro et al., 2007). This Y63F change to the protein structure changes a tyrosine residue to a phenylalanine. In order to screen for positive colonies resulting from a successful mutation, an HpyAV cut site was also built in without changing the amino acid sequence. The same mutagenesis procedure used for the biobrick sites was used to add this modification to our system, in order to further increase its efficiency.
3) Replacing DszD with HpaC & Introducing Catalase
In the 4S pathway, FMNH2 is consumed by the first three steps in the pathway carried out by DszA and DszC. When this compound is consumed, the speed of oxidation of DBT is reduced as the pathway begins to halt. A high concentration of FMNH2 on the other hand produces hydrogen peroxide (H2O2) that is lethal to cell in high concentrations (Gala´n et al. 2000).
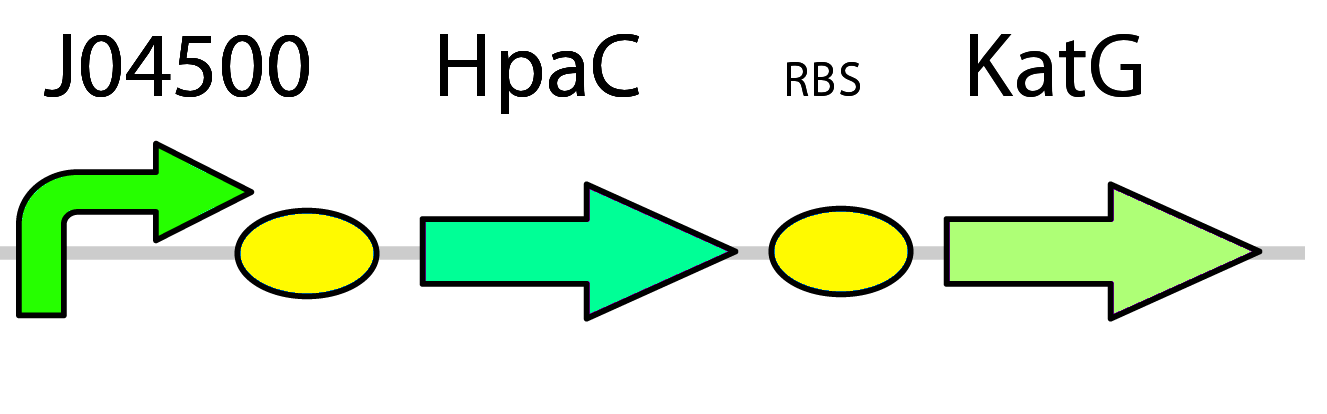
To overcome these problems in our system, two plans of optimization were implemented. First, the hpaC gene coding for another flavin reductase in E. coli W is used in place of the dszD gene which is native to the pathway. It has been shown that HpaC improves the rate of desulfurization by 7-10 times when compared to DszD (Gala´n et al., 2000). Second, a catalase gene was included in the final construct in order to take care of any excess hydrogen peroxide formed during the recycling of FMNH2 and prevent stress on the cell ( pLacI-katG-LAA).
Results
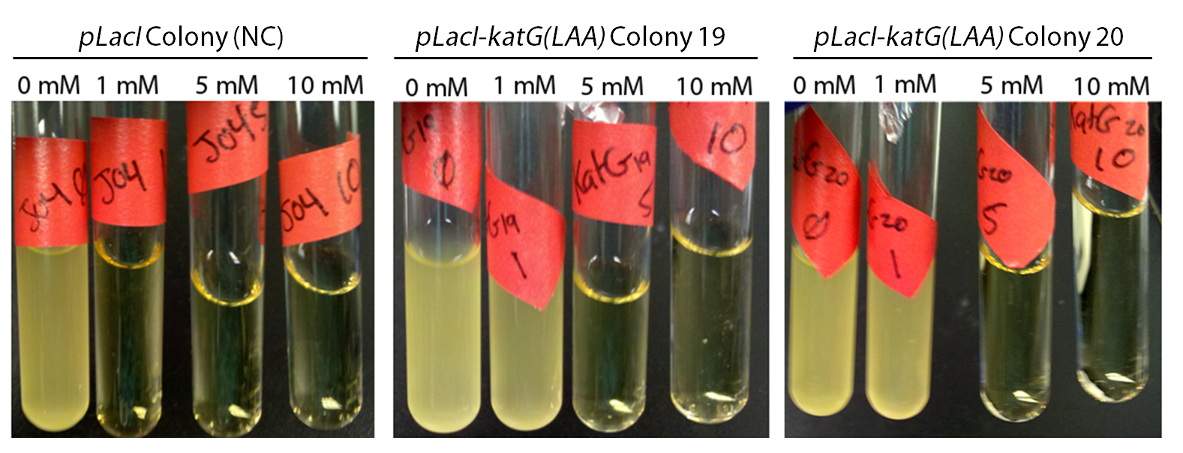
In order to determine if the KatG-LAA protein was functional, catalase activity was tested using a turbitity comparison between E. coli expressing native catalase levels (carrying only the pLacI biobrick) and cultures of pLacI-katG-LAA. Cultures were grown overnight, induced with 100 uM IPTG, and subsequently subcultured into various concentrations of hydrogen peroxide (0 mM, 1 mM, 5 mM, and 10 mM). These cultures were then grown overnight. It was found that the negative control exhibited no growth after 12h at 1 mM peroxide, however cultures with induced expression of catalase were turbid after 12 h of growth at this concentration (Fig. 3). This demonstrated the ability of the catalase to protect the cells from excessive peroxide concentrations.
HpaC activity was tested by comparing cell lysate ability to consume NADH between cultures over-expressing HpaC to a control culture that is not. This is done by monitoring the decrease in absorbance at 340 nm (Kamali et al., 2010). In order to do this, cultures of pLacI-hpaC and pLacI-dszB were grown up overnight in LB with appropriate antibiotics. Following this, protein expression was induced with IPTG, after which the assay was carried out as described in the following figure and on the protocols page.
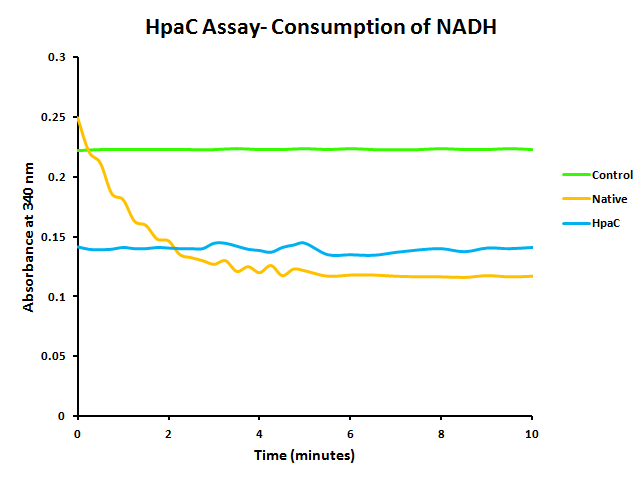
When the assay was run, it was found that NADH does not convert readily to NAD+ on its own. When cell lysate containing the naturally expressed amounts of oxidoreductase was added, a decrease in absorbance could quickly be observed as the NADH was converted to NAD+. When cultures over-expressing HpaC were tested, the absorbance levels were found to start much lower than the control. We believe that this is because with the amount of cell lysate tested, when the HpaC protein is overexpressed the NADH is consumed almost immediately and therefore the data reflecting the drop in absorbance is missed. Further tests will use differing amounts of cell lysate in order to try to capture data that shows the drop in absorbance for HpaC cultures.

4) Optimizing Gene Order
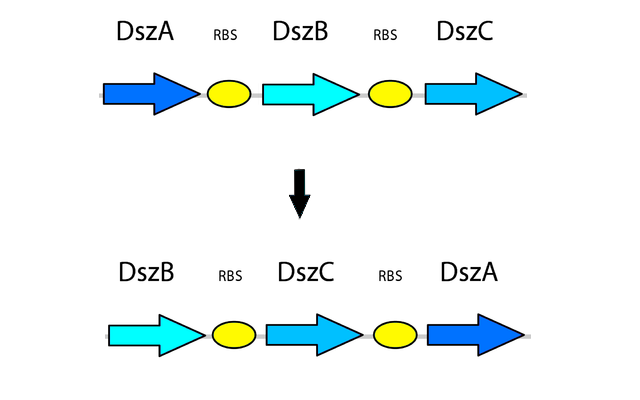
After the genetic elements we intended to use in the pathway were selected, more research was performed into further ways to optimize the system for maximum output and efficiency in order to overcome the issues present in the natural system. Rearranging the order of the genes in the operon has also shown to be beneficial in improving the efficiency of the 4S pathway. The natural order of genes in the operon is dszABC, however the relative activity of the enzymes does not reflect this pattern, as the catalytic activity of DszA:DszB:DszC is 25:1:5 respectively. Because of this, rearranging the operon to dszBCA produces more of the enzyme with lower activity and less of the enzyme with high activity. This is accomplished simply by having more of the less active enzyme transcribed, which translates into more enzyme being present to compensate for its lower activity (Li et al., 2008). In addition, the initiation codon of dszB overlaps with the termination codon of dszA in the natural operon, which leads to translational coupling and further reduces the amount of DszB, leading to a bottleneck in the pathway. By seperately PCR'ing out each of the genes and arranging them in this fashion in a biobrick plasmid, with each separated by an RBS site part (BBa_B0034), this issue is also eliminated.
Final Constructs
After all of the above, we hope to develop four final constructs for our system which will allow us to test the desulfurization under a few different conditions.
The first two constructs have the modified dsz operon (dszB, dszC, dszA) under the control of a constitutive TetR promotor (BBa_J13002) This is to allow for the testing of the optimization circuit, which is under the control of a lacI promotor inducible by IPTG (BBa_J04500). The set-up of these two constructs will therefore allow for the expression of the dsz genes with the ability to test and compare their desulfurization rates
A) On their own
B) With the addition of hpaC
C) With the addition of both hpaC and katG-LAA
This will allow us to determine what the optimal construct and expression levels of the additional genes must be in order to have the most effective sulfur removal system.
Because there are a large number of proteins being expressed in this system, we were concerned about the possibility of inclusion bodies forming which would essentially result in an unusable system, in addition to possible other toxic effects due to protein expression. Because of this a backup system was built, where both the optimization circuit and the dsz operon were under the control of the lacI promoter. This system would allow us to tune the expression of the genes, and determine which expression level is optimal. Once tuning was performed, a constitutive promotor of that strength could then be implemented to maintain optimal gene expression.
Currently, assembly of these final constructs is underway, with only a couple more construction steps before functionality tests can begin.
 "
"